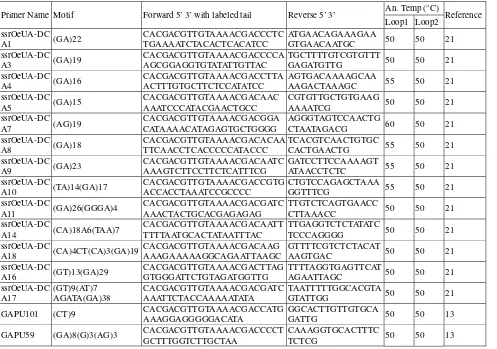
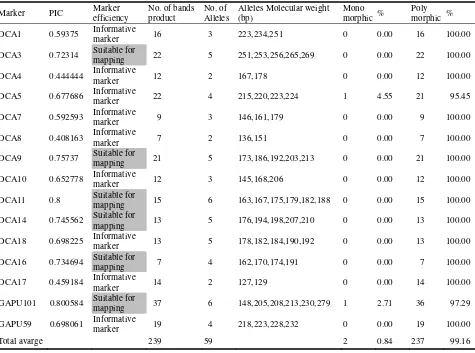
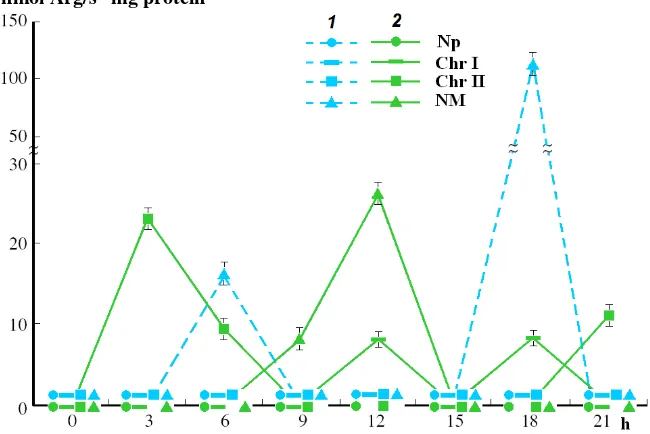
Journal of Life Sciences
Volume 6, Number 12, December 2012 (Serial Number 56)
David Publishing Company www.davidpublishing.com
Publication Information
Journal of Life Sciences is published monthly in hard copy (ISSN 1934-7391) and online (ISSN 1934-7405) by David Publishing Company located at 9460 TELSTAR AVE SUITE 5, EL MONTE, CA 91731, USA.
Aims and Scope
Journal of Life Sciences, a monthly professional academic journal, covers all sorts of researches on molecular biology, microbiology, botany, zoology, genetics, bioengineering, ecology, cytology, biochemistry, and biophysics, as well as other issues related to life sciences.
Editorial Board Members
Dr. Stefan Hershberger (USA), Dr. Suiyun Chen (China), Dr. Farzana Perveen (Pakistan), Dr. Francisco Torrens (Spain), Dr. Filipa João (Portugal), Dr. Masahiro Yoshida (Japan), Dr. Reyhan Erdogan (Turkey), Dr. Grzegorz Żurek (Poland), Dr. Ali Izadpanah (Canada), Dr. Barbara Wiewióra (Poland), Dr. Valery Lyubimov (Russia), Dr. Amanda de Moraes Narcizo (Brasil), Dr. Marinus Frederik Willem te Pas (The Netherlands), Dr. Anthony Luke Byrne (Australia), Dr. Xingjun Li (China), Dr. Stefania Staibano (Italy), Dr. Wenle Xia (USA), Hamed Khalilvandi-Behroozyar (Iran).
Manuscripts and correspondence are invited for publication. You can submit your papers via Web Submission, or E-mail to [email protected] or [email protected]. Submission guidelines and Web Submission system are available at http://www.davidpublishing.com.
Editorial Office
9460 TELSTAR AVE SUITE 5, EL MONTE, CA 91731, USA Tel: 1-323-9847526, Fax: 1-323-9847374
E-mail: [email protected], [email protected]
Copyright©2012 by David Publishing Company and individual contributors. All rights reserved. David Publishing Company holds the exclusive copyright of all the contents of this journal. In accordance with the international convention, no part of this journal may be reproduced or transmitted by any media or publishing organs (including various websites) without the written permission of the copyright holder. Otherwise, any conduct would be considered as the violation of the copyright. The contents of this journal are available for any citation. However, all the citations should be clearly indicated with the title of this journal, serial number and the name of the author.
Abstracted / Indexed in
Database of EBSCO, Massachusetts, USA Chemical Abstracts Service (CAS), USA Cambridge Scientific Abstracts (CSA), USA
Chinese Database of CEPS, American Federal Computer Library center (OCLC), USA Ulrich’s Periodicals Directory, USA
Chinese Scientific Journals Database, VIP Corporation, Chongqing, China Universe Digital Library S/B, Proquest
Subscription Information
Price (per year): Print $520, Online $360, Print and Online $680.
David Publishing Company
9460 TELSTAR AVE SUITE 5, EL MONTE, CA 91731, USA Tel: 1-323-9847526, 323-410-1082; Fax: 1-323-9847374 E-mail: [email protected]
David Publishing Company www.davidpublishing.com
DAV ID P UBL ISH IN G
J LS
Journal of Life Sciences
Volume 6, Number 12, December 2012 (Serial Number 56)
Contents
Molecular Biology
1307 Evaluation of Free Radicals and Antioxidant Properties of Virus Infected Food Crops in
Azerbaijan
Irada Huseynova and Jalal Aliyev
1317 Helicobacter pylori Biofilm Formation and Gene Expression on Abiotic Surfaces Using a
Cyanobacterial Extract
Alba E. Vega, Fabio A. Persia, Gabriel A. Salinas-Ibañez, Teresa Alarcón, Stella M.C. Lucero Estrada, Teresa I. Cortiñas and Humberto J. Silva
1328 RAPD-PCR Based Marker Approach for the Genetic Differentiation of Two Species of Cockroach
(Order-Dictyoptera)
Bharat Neekhra, Divya Pandey and Subodh Kumar Jain
1334 The Determination of Geographical Origin of Foodstuffs by Using Innovative Biological Bar-Code
Aly El Sheikha and Didier Montet
1343 Molecular Characterization of Olive Cultivars in Iraq Using SSR Markers and Compare with
Phenotypic Characterization
Iqbal Harbi, Salwa Jaber Al-Awadi and Ali Imad Mohammad Moner
1351 Arg-X Protease-Sensitive in Supramolecular Structures of Interphase Cell Nucleus during Growth
Morphogenesis Mature Germs of Wheat
Ivanov Ruslan, Vafina Gulnara and Ivanova Evilina
Zoology and Botany
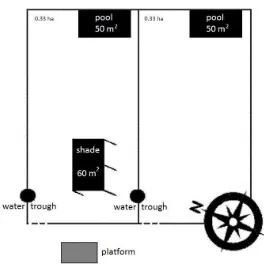
1356 What Do Cattle Prefer in a Tropical Climate: Water Immersion or Artificial Shade?
1363 Genetic Parameters for Udder Traits in Slovak Dairy Sheep and Their Crosses with Specialized Breeds
Milan Margetín, Marta Oravcová1, Dušan Apolen and Michal Milerski
1371 Amino Acid and Fatty Acid Profile in Epidermal Mucus of Bluestreak Cleaner Wrasse (Labroides
dimidiatus): Possible Role as Defense Mechanism against Pathogens
Maziidah Ab Rahman, Roslan Arshad, Faizah Shaharom and Nur Asma Ariffin
1378 Validation of Monomeric Anthocianin Determination Method for Bilberry Juice and Marc
Extracts
Ance Dandena, Ieva Leimane and Uģis Klētnieks
1383 Necrotic Incompatibility in Crosses of Bread Wheats in Argentina
Francisco Javier Di Pane and Gilberto Kraan
1387 Effect of Natural Surface Secretes of Some Common Ornamental Plants Leaves on Pathogenic
Micro-organisms
Fouad H. Kamel, Hero M. Ismael and Akhter A. Ahmed
Interdisciplinary Researches
1391 Enabling Sustainable Management of Non Wood Forest Products in South East Europe–Albania’s
Experience Elvira Bazina
1396 Verbal Communication in HIV-1 Patients: A New Perspective on the Study of Cognitive Disorders
Valeria Abusamra, Lorena Abusamra, Bárbara Sampedro, María Macaya, Mercedes Güemes, Micaela Difalcis and Aldo Ferreres
1408 Cortical Arousal Strategies in Left-Handers during the Aural Perception and Manual Playback of
Mono- and Polyphonic Rhythmical Patterns
Pavlovych Olga Sergiyivna, Morenko Alevtyna Grigorivna, Kotyk Olena Adamivna and Vlasjuk Svitlana Sergiyivna
1414 Impact of Omega-3 Nutritional Protocol on the Occurrence of Preeclampsia among High Risk
Pregnant Women
Abeer Mokhtar Abu Khatwa and Shadia Hassan Abdel Kader
1420 Wildlife Crossing Zones along the Ring Changbai Mountain Scenic Highway, China
Journal of Life Sciences 6 (2012) 1307-1316
Evaluation of Free Radicals and Antioxidant Properties
of Virus Infected Food Crops in Azerbaijan
Irada Huseynova and Jalal Aliyev
Department of Fundamental Problems of Biological Productivity, Institute of Botany, Azerbaijan National Academy of Sciences,
Baku AZ1073, Azerbaijan
Received: July 28, 2012 / Accepted: October 10, 2012 / Published: December 30, 2012.
Abstract: Viral diseases are an important limiting factor in many crop production systems in Azerbaijan. Symptomatic plants in main
crop-producing areas were tested by ELISA (enzyme-linked immunosorbent assay) using specific monoclonal and polyclonal antibodies. Then RCA (rolling circle amplification) of circular DNA and PCR using different specific primer pairs have indicated that the tested symptomatic plant samples were completely infected by the following viruses: Luteovirus [BLRV (Bean leaf roll virus)], Potyviruses [BCMV (Bean common mosaic virus), BYMV (Bean yellow mosaic virus)], Bromovirus [(AMV) Alfa-alfa mosaic virus], Geminiviruses [CpCDV (Cickpea chlorotic dwarf virus) and TYLCV (Tomato yellow leaf curl virus)] and Nanoviruses [two different FBNYV (Faba bean necrotic yellow virus) and FBNSV (Faba bean necrotic stunt virus)]. At the same time generation sites of superoxide and hydrogen peroxide radicals and activity of antioxidant enzymes were studied in the naturally infected plants.
Key words: Food crops, virus-like symptoms, viral diseases, molecular methods, reactive oxygen species, antioxidant enzymes.
1. Introduction
The ability to accurately detect and identify a potential plant pathogenic organism is fundamental to plant pathogen diagnostics and plant disease management for food quality. Increasing international travel and trade of plant materials enhances the risk of introducing new viruses and their vectors into production systems. In addition, changing climate conditions can contribute to a successful spread of newly introduced viruses or their vectors and establishment of these organisms in areas that were previously unfavorable [1, 2].
Viral diseases are also an important limiting factor in many crop production systems in our country. It causes extensive leaf yellowing, stem and leaf deformation, reduced fruit quality, substantial crop loss and shortening the life-span of vegetable crops [3].
Corresponding author: Jalal Aliyev, professor,
academician, research field: plant physiology. E-mail: [email protected].
The last four years infection incidences were
determined for nine viruses on major food crops such
as chickpea, lentil, faba bean, tomato, pea and alfa-alfa.
Evaluation of Free Radicals and Antioxidant Properties of Virus Infected Food Crops in Azerbaijan
1308
in peroxisomes. Enzymes and metabolites of the ascorbate-glutathione cycle (APO (ascorbate peroxidase; EC 1.11.1.11); GR (glutathione reductase,
EC 1.8.1.7)) which is important in H2O2 scavenging
are located in organelles and cytoplasm [5, 6]. Antioxidant enzymes were often studied at sites of attempted pathogen attack and in connection with immediate responses of invaded cells [7].
The main aim of this study is focused on detection of viral infections of vegetable crops by different molecular techniques, to find out generation sites of superoxide and hydrogen peroxide radicals in the naturally infected plants and to investigate the possible role of antioxidant systems against stress, in order to deepen our knowledge of the plant-virus interaction.
2. Materials and Methods
2.1 Field Visits and Sample Collections
To determine the presence or absence of virus
infection, the main crop-producing areas of Azerbaijan, fields of faba bean (Vicia faba L.), pea (Pisum sativum
L.), chickpea (Cicer arietinum L.), tomato (Solanum
lycopеrsicum L.) and lentil (Lens culinaris L.) were
surveyed in different regions (including Goychay, Nakhchivan, Masalli and Lerik) during the period of 2009-2011. Samples were collected from plants showing virus-like symptoms, such as leaf rolling, yellowing, mosaic, stunting, wilting, and shortening of the internodes, phloem discoloration, necrosis and stunted growth (Fig. 1).
The number of samples collected in each field depended on the number of symptomatic plants observed. Virus-free plants for negative control were collected under same field conditions. Each field was evaluated using a standard format, recording location, conditions, development stage, virus disease symptoms, and presence or absence of the insect populations. Virus disease incidence in each field
was determined on the basis of visualsymptoms and by
(a) (b)
(c) (d)
Fig. 1 Symptomatic Solanum lycopеrsicum plants collected from Masalli fields associated with virus infestation and showing
Evaluation of Free Radicals and Antioxidant Properties of Virus Infected Food Crops in Azerbaijan 1309
counting the percentage of infected plants at different, randomly selected locations in the field. Collected plant samples with symptoms of potential virus
infection were immediately frozen in liquid N2 and
stored at -20 °C.
2.2 Detection of Viruses Using ELISA
Virus detection was performed with double and triple antibody sandwich ELISA (DAS-ELISA and TAS-ELISA) [8] using polyclonal and monoclonal antibodies (diluted 1:500 and 1:1000) for following
viruses: BLRV (Bean leaf roll virus), BCMV (Bean
common mosaic virus), BYMV (Bean yellow mosaic
virus), AMV (Alfa-alfa mosaic virus), CpCDV
(Cickpea chlorotic dwarf virus) and TYLCV (Tomato
yellow leaf curl virus). All antibodies were kindly
provided by Dr. S. Winter (DSMZ, Braunschweig, Germany). ELISA result was measured by recording its absorbance value using an ELISA plate reader (Stat
Fax) at A405. Samples with absorbance values higher
than the mean value for non-infected control plants plus two or three standard deviations were considered positive.
2.3 DNA Extraction and Molecular Analyses
Plant samples that reacted serologically with the virus antibody were selected for further testing by RCA (rolling circle amplification) and PCR (polymerase chain reaction). Frozen plant samples were ground in liquid nitrogen, and DNA was
prepared using the method of Edwards [9], modified
as described by Grigoras [10]. The circular viral DNA
was amplified by RCA using the TempliPhi
Amplification Kit (GE HealthCare, UK). φ29 DNA
polymerase amplifies single- or double-stranded circular DNA templates by rolling circle amplification. Viral DNA for detection nanovirus infection was amplified by PCR using primer pairs F103/R101 and C5F/C5R, which yielded PCR products of the expected size (770 bp and 660 bp, respectively) [11]. Viral DNA extracted from infected plants was also
amplified by PCR using different specific primer pairs, which yielded PCR products of the expected size. The amplifications were carried out in an Applied Biosystems 2720 Thermal Cycler© Biorad (SingAPOre) using the special program. Amplification products were resolved by electrophoresis in a 1.2% agarose gel with TAE buffer and stained with ethidium
bromide (0.5 μg/mL). The gels were photographed
under UV light by “Gel Documentation System UVITEK” (UK).
2.4 Histochemical Staining of Superoxide Anion Radical
Histochemical staining for ROS accumulation was conducted as described previously [12-14] with some modifications. For superoxide determination Petri dishes were used, the leaf samples were immersed in 6 mM NBT solution containing 50 mM sodium phosphate (pH 7.5) and 10 mM sodium azide for 12 h in the dark. ROS reaction was stopped by soaking the leaves with lacto-glycerol-ethanol (1:1:4 by vol.) and boiling in water for 5 min, and the cleared leaves were preserved in 50% ethanol and photographed.
2.5 Histochemical Staining of H2O2
To detect hydrogen peroxide, traverse sections of the leaf stem were cut by hand, and the detached leaves of the virus infected plants were deposited in Petri dishes containing a solution of 5 mM DAB and 10 mM MES at pH 3.8 for 12 h in darkness. To detect hydrogen peroxide, reaction was stopped by soaking the leaves with lacto-glycerol-ethanol (1:1:4 by vol) and boiling in water for 5 min, and the cleared leaves were preserved in 50% ethanol and photographed [12-14].
2.6 Enzyme Extractions and Determination of the
Activity
Evaluation of Free Radicals and Antioxidant Properties of Virus Infected Food Crops in Azerbaijan
1310
a pestle and ice-cold mortar using different specific enzyme buffers. The homogenates were filtered through four layers of cheesecloth and then centrifuged at 4 °C for 20 min at 15,000× g. The supernatant was collected and used for analyses of enzymatic activities.
2.6.1 CAT
The activity of catalase was determined as a decrease in absorbance at 240 nm for 1 min
following the decomposition of H2O2 as described by
Kumar and Knowles [15]. The reaction mixture contained 50 mM phosphate buffer (pH 7.0) and 15
mM H2O2 and reaction was initiated by adding
enzyme extract. 2.6.2 APO
The activity of ascorbate peroxidase was assayed according to Nakano and Asada [16]. The assay
mixture consisted of 0.05 mM ASA, 0.1 mM H2O2,
0.1 mMEDTA, 50 mMsodium phosphate buffer (pH
7.6), and 0.3 mL enzyme extract. The activity was measured as a decrease in absorbance at 290 nm for 30 s.
2.6.3 GR
Glutathione reductase activity was determined at 340 nm for 10 min in 1 mL reaction mixture containing 100 mM potassium phosphate buffer (pH 7.8), 1 mM EDTA, 0.2 mM NADPH and 0.5 mM GSSG [17].
2.6.4 SOD
Superoxide dismutase activity was estimated by using SOD Assay Kit-WST (Sigma-Aldrich, USA). The absorbance was recorded at 450 nm and one enzyme unit of SOD activity was defined as the amount of enzime recuired to cause 50% inhibition of the rate of NBT reduction.
3. Results and Discussion
3.1 Virus Infection
All collected plant samples firstly were tested by enzyme-linked immunosorbent assay. Then DNA was extracted from the fresh leaves, amplified by RCA
using φ29 DNA polymerase and restricted by
endonucleases AatII, xBaI, Sau3A, BamHI or HindIII [18-19]. All amplified products were resolved by agarose gel electrophoresis. Detection of virus infection in symptomatic samples was also performed by PCR method using different specific primer pairs [11]. The use of different molecular methods indicated that the tested symptomatic plant samples were completely infected by the following viruses:
Luteovirus [BLRV(Bean leaf roll virus)], Potyviruses
[BCMV (Bean common mosaic virus), BYMV(Bean
yellow mosaic virus)], Bromovirus [AMV (Alfa-alfa
mosaic virus)], Geminiviruses [CpCDV (Cickpea
chlorotic dwarf virus) and TYLCV (Tomato yellow
leaf curl virus)] and Nanoviruses [two different
FBNYV (Faba bean necrotic yellow virus) and
FBNSV (Faba bean necrotic stunt virus)]. It is
important to note that additional pathogens can be expected on these crops.
3.2 Determination of Superoxide Anion
As known, the level of plant resistance to viral diseases provides many physiological and biochemical parameters responsible for maintaining the viability and alterations in plant metabolism under stress conditions. On this basis, histochemical study of the
possible presence of superoxide anion, H2O2 and
activity of antioxidant enzymes in virusinfected plant
leaves were performed in the present work.
Evaluation of Free Radicals and Antioxidant Properties of Virus Infected Food Crops in Azerbaijan 1311
process is initiated by a reaction between NO (nitric oxide) and H2O2, which is formed by O2−(superoxide) dismutation by SOD during the HR.
Plant-virus interaction may result in a host hypersensitive response or in systemic symptoms [22, 23]. One of the earliest responses of plant cells to pathogens is the production of ROS. The typical ROS detected are O2. (superoxide radicals) and H2O2 [24, 25]. ROS play a crucial role during pathogenesis. They are involved in the hypersensitive response typical for plant-pathogen incompatible interactions. They can limit the spread of pathogen by strengthening plant cell walls and/or by killing pathogens directly [26, 27]. However, ROS act as cytotoxic compounds, too.
In this work, the authors detected the presence of
superoxide anion O2– in places of infection with the
use of NBT. Accumulation of insoluble blue-colored formazan complex (reduced NBT) is an indicator of generation of ROS, in particular of superoxide anion. This accumulation was observed in infected leaves after infiltration. Then staining declined rapidly, preceding the apparition of necrosis (Fig. 2). Histochemical staining for superoxide production in
leaves tissues was based on the ability of O2– to
reduce NBT and used to detect in situ the production
of superoxide radicals [28]. Detached leaflets from
plants subjected to the viral diseases above described and their respective controls were immersed in sodium phosphate buffer (pH 7.8) containing 0.1% NBT and 10 mM sodium azide.
Leaflets of healthy plants were also infiltrated with
50 mM sodium phosphate buffer (pH 7.8) containing only 10 mM sodium azide and used as control. Superoxide was visualized as a purple discoloration of NBT. Discoloration of leaf was quantified using a digital imaging system (Fig. 3). In typical incompatible interactions, one of the early events of HR is an oxidative burst with the generation of
superoxide (O2−) and the subsequent accumulation of
H2O2 [21].
(a)
(b)
(c)
Fig. 2 Detection of superoxide anion radical with NBT staining. Leaves were infiltrated after being submerging in a reaction mixture containing 6 mM NBT in 50 mM sodium phosphate buffer (pH 7.6) plus 10 mM sodium azide. (a) Cucumis sativus L.; (b) Zea mays L.; (c) Solanum lycopеrsicum L. Healthy leaves shown on the right panels, infected leaves shown on the left panels. The experiment was repeated two times.
(a)
(b)
(c)
Fig. 3 Detection of H2O2 using DAB (3,3
’
- diaminobenzidine tetrahydrochloride) staining method. The reaction mixture contained 5 mM DAB in 10 mM MES
buffer (pH 3.8). (a) Cucumis sativus L.;(b) Zea mays L.; (c)
Evaluation of Free Radicals and Antioxidant Properties of Virus Infected Food Crops in Azerbaijan
1312
3.3 Determination of H2O2
Accumulation of H2O2 was observed using DAB
staining method. The reaction mixture contained 5 mM DAB in 10 mM MES (pH 3.8). Leaves were incubated overnight at 25 °C, and then cleared twice
in 50% (v⁄v) ethanol for 10 min. DAB polymerizes to
produce a brown precipitate on contact with H2O2 in
the presence of peroxidase, and, thus, provides a useful marker of peroxide accumulation [29]. Superoxide anions are thought to be produced outside the plant cell by a plasma membrane-associated NAD(P)H oxidase, and are usually rapidly converted
to H2O2 by superoxide dismutase. To examine whether
H2O2 is also accumulated at the site of elicited HR,
infected leaves were dipped in a solution of DAB. Fig. 3
clearly shows that H2O2 accumulated during the HR
caused by virus infection. In higher plants, production
of H2O2 is thought to be driven by increases in the
concentrations of superoxide anions. However, a slightly lower level of DAB staining was observed in healthy leaves compared with infected leaves (Fig. 4). Thus, the results indicate that an alteration in the chloroplastic metabolism is produced during the early response to virus infection favoring the accumulation of ROS in the plants.
3.4 Analyses of the Antioxidant Enzymes Activity
Plants have evolved complex antioxidant systems in order to protect cellular membranes and organelles from the damaging effects of ROS [30, 31]. Increase in peroxidase activity is also a response to viral infection, and has been reported in tobacco [32], peaches, apricots [33] and beans [34].
In plant cells, enzymes and redox metabolites act in synergy to carry out ROS detoxification. SOD
catalyses the dismutation of O2– to H2O2, CAT
dismutates H2O2 to oxygen and water, and APO
reduces H2O2 to water by utilising ASC (ascorbate)
as specific electron donor. These are considered the main enzymatic systems for protecting cells against oxidative damage. The balance between SOD and
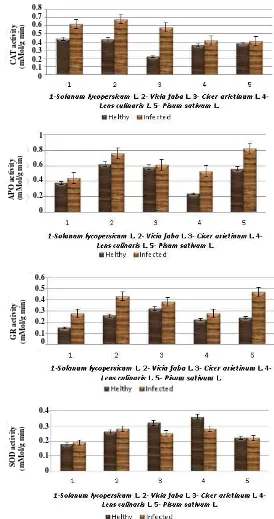
APO or CAT activities in cells is crucial for determining the steady-state level of O2– and H2O2. Redox metabolites, such as ASC and the tripeptide GSH (glutathione), also protect plant cells against ROS-induced damage, either by directly removing reactive chemical species or blocking the oxidative chain reactions triggered by ROS. Results reported in the literature indicate that alteration in the expression/activity of ROS-scavenging enzymes could also be a key step in the activation of phytopathogen defence. On this base, the authors also studied the activities of the antioxidant enzymes, CAT, APO, SOD and GR, in viral infected plant leaves. These enzymes are known to be involved in an immediate plant defense response. Samples for activity measurements of the antioxidant enzymes were collected during the early stage of the infection, when the first visible symptoms of the virus infection appeared on the leaves (in early June). As shown in Fig. 4, the activity of antioxidant enzymes in infected leaves generally was higher than that of comparable healthy leaves.
Analysis of CAT activity in infected leaves showed that this enzyme in all the samples studied had a significant difference compared with the control. CAT activity was 1.4-fold higher (up to 41%)
in infected leaves of Solanum lycopersicum and
1.27-fold higher (up to 32%) in infected leaves of
Vicia faba compared to the healthy plants. The most
significant differences between the values of CAT
activity were observed in infected Cicer arietinum
samples, where the activity was 2.6-fold higher (up to 163%) compared to the control plants. In infected
Lens culinaris and Pisum sativum samples, CAT
activity only was slightly compared with the control.
Analysis of CAT activity in infected Lens culinaris
leaves showed that this enzyme activity did not differ significantly (only up to 17%) compared to the control and accounting 0.42 mmol/mg min, respectively. As shown in Fig. 4, CAT activity in
infected Pisum sativum leaves did not seem to be
Evaluation of Free Radicals and Antioxidant Properties of Virus Infected Food Crops in Azerbaijan 1313
Fig. 4 Activities of catalase, ascorbate peroxidase, glutathione reductase and superoxide dismutase in viral infected plant leaves.
APO activity was 1.2-fold higher in infected leaves
of Solanum lycopersicum compared to the healthy
plants and accounting 0.44 mmoL/mg min. As shown in Fig. 4, APO activity in infected leaves of Vicia faba
and Cicer arietinum slightly increased (up to 24% and
16%, respectively), although the activity of CAT
significantly differed compared to the control. The most significant differences between the activities of
APO and CAT were observed in infected Lens
culinaris and Pisum sativum samples, where the APO
activity was 2.2 and 1.5-fold higher compared to the control plants. This may indicate the existing
C
A
T
acti
vit
y
(mMol/
g min
)
0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0
A
P
O
activi
ty
(mMol/
g min
)
1
0.8
0.6
0.4
0.2
0
0.6 0.5 0.4
0.3 0.2
0.1 0
GR a
ctivity
(mMol/
g min
)
0.4
0.3
0.2
0.1
0
SO
D
acti
vi
ty
(mMol/
g min
Evaluation of Free Radicals and Antioxidant Properties of Virus Infected Food Crops in Azerbaijan
1314
functional relationship and competition between the studied enzymes under these viral diseases. Other works also suggest that, along with the activation of SOD and APO in the leaves, a sharp decrease in activity of CAT is observed, which may be due to
inhibition of the enzyme substrate—H2O2
Interestingly, it has been reported that in a compatible response between barley and powdery mildew the cytosolic isoenzyme of APO is up-regulated in both epidermal and mesophyll cells. In these cells, that are not able to trigger a response to stop pathogens, the APO increase limits the propagation of oxidative processes allowing cells to maintain their viability, a condition required for the penetration of biotrophic powdery mildew in plant tissues [35]. This up-regulation of APO confirms previous results reporting an increase in APO activity during successful infection of barley leaves by biotrophic compatible pathogens [36-38] and has also been reported to occur in leaves of susceptible apricot infected by plum pox virus [39]. CAT activity has also been reported to decrease in cells undergoing HR.
However, the suppression mechanisms of these two
H2O2 scavenging enzymes are different. CAT is
down-regulated at the transcription level [40], whereas, APO regulation in HR involves both transcription and translation (or posttranslation) processes. In tobacco leaves, inoculated with TMV (tobacco mosaic virus), a rise in APO mRNA occurs [41], probably as an antioxidant response triggered by the increasing
presence of H2O2 within cells and similar to that
activated under abiotic stress [42]. In spite of the increase in its expression, the activity of the enzyme is strongly suppressed in the TMV-infected cells by a mechanism, still not well characterized, that acts at the transcriptional or post-transcriptional level [43]. In this case, the high activity of antioxidant enzymes can probably be one of the markers of resistance to the pathogen.
The effects of a viral infection caused an increase in GR activity about 1.86-fold higher (up to 72%) and
accounting 0.28 mmoL/mg min in infected leaves of
Solanum lycopersicum compared to the healthy plants.
GR activity in infected samples of Vicia faba increased up to 17% compared to the control. In contrast, the GR
activity in infected samples of Cicer arietinum and
Lens culinaris did not significantly differ from the
control plants. As shown in Fig. 4, the most significant difference between the GR activities were observed in
infected Pisum sativum samples, where activity
increased approximately 2-fold higher, i.e. up to 96% compared to the control plants.
As shown in Fig. 4, the change of SOD activity in infected plant leaves was different, the actives of Cu/Zn-SOD decreased (up to 46% and 22%) in infected samples of Cicer arietinum and Lens culinaris compared to the healthy plants. The most interesting
value of SOD activity observed in infected Pisum
sativum samples, where the activity did not differ from
the control and accounted 0.22 unit/mg proteins.
Obviously, the activation of antioxidant defence systems in plants by abiotic and biotic stresses is a general phenomenon and probably contributes to increased resistance against a subsequent stress.
4. Conclusion
The following viruses on major food crops collected from different regions of Azerbaijan:
Luteovirus [BLRV(Bean leaf roll virus)], Potyviruses
[BCMV (Bean common mosaic virus), BYMV(Bean
yellow mosaic virus)], Bromovirus [AMV (Alfa-alfa
mosaic virus)], Geminiviruses [CpCDV (Cickpea
chlorotic dwarf virus) and TYLCV (Tomato yellow
leaf curl virus)] and Nanoviruses [two different
FBNYV (Faba bean necrotic yellow virus) and
FBNSV (Faba bean necrotic stunt virus)] were
detected in this study. ROS generation is a common feature in both incompatible and compatible plant-pathogen interactions. On this basis, histochemical evaluation of the possible presence of
superoxide anion, H2O2 and activity of antioxidant
Evaluation of Free Radicals and Antioxidant Properties of Virus Infected Food Crops in Azerbaijan 1315
in the present work. Results reported in this study also indicate that the alteration in the expression/activity of ROS-scavenging antioxidant enzymes could also be a key step in the activation of phytopathogen defence.
The ability to maintain high physiological function in the presence of viruses may be a more important resistance or tolerance mechanism than actual avoidance of infection. The information presented in the current study is important because the better understanding of the mechanisms behind the viral impact on host plant physiology can lead to the development of improved cultivars that either resist viral infection or can better tolerate infection by experiencing less severe symptoms.
References
[1] V. Ortiz, E. Navarro, S. Castro, G. Carazo, J. Romero, Incidence and transmission of faba bean necrotic yellows virus (FBNYV) in Spain, Spanish Journal of Agricultural Research 4 (2006) 255-260.
[2] H.J. Vetten, P.W.G. Chu, J.L. Dale, R. Harding, J. Hu, L. Katul, et al., Nanoviridae, in: C.M. Fauquet, M.A. Mayo, J. Manoloff, U. Desselberger, L.A. Ball (Eds.), Virus Taxonomy, Eighth Report of the International Committee on Taxonomy of Viruses, Elsevier, London, 2005, pp. 343-452.
[3] A.F. Bent, Plant disease resistance genes: Function meets structure, Plant Cell 8 (1996) 1757-1771.
[4] J. Stanley, D.M. Bisaro, R.W. Briddon, J.K. Brown, C.M. Fauquet, B.D. Harrison, et al., Family Geminiviridae, in: C.M. Fauquet, M.A. Mayo, J. Manoloff, U. Desselberger, L.A. Ball (Eds.), Virus Taxonomy, Eighth Report of the International Committee on Taxonomy of Viruses, Elsevier, London, 2005, pp. 301-326.
[5] S.H. Lee, N. Ahsan, K.W. Lee, D.H. Kim, D.G. Lee, S.S. Kwak, et al., Simultaneous overexpression of both CuZn superoxide dismutase and ascorbate peroxidase in transgenic tall fescue plants confers increased tolerance to a wide range of abiotic stresses, Journal of Plant Physiology 164 (2007) 1626-1638.
[6] J.L. Dangl, J.D.G. Jones, Plant pathogens and integrated defence responses to infection, Nature 411 (2001) 826-833.
[7] R. Goldbach, E. Bucher, M. Prins, Resistance mechanisms to plant viruses: An overview, Virus Research 92 (2003) 207-212.
[8] M.F. Clark, A.N. Adams, Characteristics of the microplate method of enzyme-linked immunosorbent
assay for the detection of plant viruses, Journal of General Virology 34 (1977) 475-483.
[9] K. Edwards, C. Johnstone, C. Thompson, A simple and rapid method for the preparation of plant genomic DNA for PCR analysis, Nucleic Acids Research 19 (1991) 13-49.
[10] Grigoras, T. Timchenko, L. Katul, A. Grande-Pérez, H.J. Vetten, B. Gronenborn, Reconstitution of authentic nanovirus from multiple cloned DNAs, Journal of Virology 83 (2009) 10778-10787.
[11] I.M. Huseynova, N.F. Sultanova, A.Ch. Mammadov, Detection of single-stranded DNA plant viruses in vegetable plants using PCR method, Proceedings of ANAS 66 (2) (2011) 5-12.
[12] M.J. Fryer, L. Ball, K. Oxborough, S. Karpinski, P.M. Mullineaux, N.R. Baker, Control of ascorbate peroxidase 2 expression by hydrogen peroxide and leaf water status during excess light stress reveals a functional organisation of Arabidopsis leaves, Plant Journal 33 (2003) 691-705.
[13] T. Kariola, G. Brader, E. Helenius, J. Li, P. Heino, E.T. Palva, Early responsive to dehydration 15—A negative regulator of ABA-responses in Arabidopsis, Plant Physiology 142 (2006) 1559-1573.
[14] R. Mahalingam, N. Shah, A. Scrymgeour, N. Fedoroff, Temporal evolution of the Arabidopsis oxidative stress response, Plant Molecular Biology 57 (2005) 709-730. [15] C.N. Kumar, N. Knowles, Changes in lipid peroxidation
and lipolytic and free-radical scavenging enzyme during aging and sprouting of potato (Solanum tuberosum L.) seed-tubers, Plant Physiology 102 (1993) 115-124. [16] Y. Nakano, K. Asada, Hydrogen peroxide is scavenged
by ascorbate specific peroxidase in spinach chloroplasts, Plant Cell Physiology 22 (1981) 867-880.
[17] G.G. Yannarelli, A.J. Fernandez-Alvarez, Glutatione reductase activity and isoforms in leaves and roots of wheat plants subjected to cadmium stress, Phytochemistry 68 (2007) 505-512.
[18] N.F. Sultanova, I.M. Huseynova, A.Ch. Mammadov, Detection and identification of main virus infection in vegetable plants by sandwich enzyme-linked immunosorbent assay (DAS-ELISA), Reports of ANAS LXVII (5) (2011) 113-122.
[19] I.M. Huseynova, A.Ch. Mammadov, N.F. Sultanova, J.A. Aliyev, First report of circular single-stranded DNA viruses of vegetables in Azerbaijan: Fast definition by RCA (rolling circle amplification) method and molecular characterization, Reports of ANAS LXV (6) (2009) 106-114.
Evaluation of Free Radicals and Antioxidant Properties of Virus Infected Food Crops in Azerbaijan
1316
[21] P. Wojtaszek, Oxidative burst: An early plant response to pathogen infection, Biochemistry Journal 322 (1997) 681-692.
[22] K.E. Hammond-Kosack, J.D.G. Jones, Responses to plant pathogens, in: B.B. Buchanan, W. Gruissem, R.L. Jones (Eds.), Biochemistry & Molecular Biology of Plants, American Society of Plant Physiologists, Rockville, Maryland, 2000, pp. 1102-1156.
[23] J.A. Hernández, J.M. Talavera, P. Martínez-Gómez, F. Dicenta, F. Sevilla, Response of antioxidative enzymes to plum pox virus in two apricot cultivars, Physiologia Plantarum 111 (2001) 313-321.
[24] I.E. Johansen, O.S. Lund, C.K. Hjulsager, J. Laursen, Recessive resistance in Pisum sativum and potyvirus pathotype resolved in gene-for-cistron correspondence between host and virus, Journal of Virology 75 (2001) 6609-6614.
[25] E. Kombrink, E. Schmelzer, The hypersensitive response and its role in local and systemic disease resistance, European Journal of Plant Pathology 107 (2001) 69-78. [26] M.R. Sahoo, M. DasGupta, P.C. Kole, J.S. Bhat, A.
Mukherjee, Antioxidative enzymes and isozymes analysis of taro genotypes and their implications in Phytophthora blight disease resistance, Mycopathologia 163 (2007) 241-248.
[27] S.M. Salazar, A.P. Castagnaro, M.E. Arias, N. Chalfoun, U. Tonello, J.C. Dнaz Ricci, Induction of a defense response in strawberry mediated by an avirulent strain of Colletotrichum, European Journal of Plant Pathology 117 (2006) 109-122.
[28] K.T. Leath, L.J. Rowell, Histological study of resistance
of Zea mays to Puccinia graminis, Phytopathology 56
(1966) 305-1309.
[29] C. Rusterucci, D.H. Aviv, B.F. Holt, J.L. Dangl, J.E. Parker, The disease resistance signaling components EDS1 and PAD4 are essential regulators of the cell death pathway controlled by LSD1 in Arabidopsis, Plant Cell 13 (2001) 2211-2224.
[30] V. Paranidharan, A. Palaniswami, P. Vidhyasekaran, R. Velazhahan, Induction of enzymatic scavengers of active oxygen species in rice in response to infection by Rhizoctonia solani, Acta Physiologia Plantarum 25 (2003) 91-96.
[31] Z. Šubr, S. Nováková, H. Drahovská, Detection of transgene copy number by analysis of the T1 generation of tobacco plants with introduced P3 gene of Potato virus A, Acta Virology 50 (2006) 135-138.
[32] M.L. Lagrimini, S. Rothstein, Tissue specificity of tobacco peroxidase isozymes and their induction by wounding and tobacco mosaic virus infection, Plant
Physiology 84 (1987) 438-442.
[33] P. Dıaz-Vivancos, M. Rubio, V. Mesonero, P.M. Periago, A. Ros Barcelo, P. Martınez-Gomez, et al., The
APOplastic antioxidant system in Prunus: Response to long-term plum pox virus infection, Journal of Experimental Botany 57 (2006) 3813-3824.
[34] D.E.M. Radwan, K.A. Fayez, S.Y. Mahmoud, G. Lu, Modifications of antioxidant activity and protein composition of bean leaf due to bean yellow mosaic virus infection and salicylic acid treatments, Acta Physiologia Plantarum 32 (2010) 891-904.
[35] K. Burhenne, L. Gregersen, Up-regulation of the ascorbate-dependent antioxidative system in barley leaves during powdery mildew infection, Molecular Plant Pathology 1 (2001) 303-314.
[36] H.M. El-Zahaby, G. Gullner, Z. Kiraly, Effects of powdery mildew infection of barley on the ascorbate-glutathione cycle and other antioxidant in different host-pathogen interactions, Phytopathology 85 (1995) 1225-1230.
[37] E. Kuzniak, M. Sklodowska, The effect of Botrytis cinerea infection on ascorbate glutathione cycle in tomato leaves, Plant Science 148 (1999) 69-76.
[38] H.Vanacker, T.L.W. Carver, C.H. Foyer, Pathogen-induced changes in the antioxidant status of the apoplast in barley leaves, Plant Physiology 117 (1998) 1103-1114.
[39] J.A. Hernández, J.M. Talavera, P. Martinez-Gomez, F. Dicenta, F. Sevilla, Response of antioxidative enzymes to plum pox virus in two apricot cultivars, Physiologia Plantarum 111 (2001) 313-321.
[40] S. Dorey, F. Bailleul, P. Saindrenan, B. Fritig, S. Kauffmann, Tobacco class I and II catalases are differentially expressed during elicitorinduced hypersensitive cell death and localized acquired resistance, Molecular Plant-Microbe Interactions 11 (1998) 1102-1109.
[41] R. Mittler, E. Lam, V. Shulaev, M. Cohen, Signal controlling the expression of cytosolic ascorbate peroxidase during pathogeninduced programmed cell death in tobacco, Plant Molecular Biology 39 (1999) 1025-1035.
[42] Kubo, H. Saji, K. Tanaka, N. Kondo, Expression of Arabidopsis cytosolic ascorbate peroxidase gene in response to ozone of sulfur dioxide, Plant Molecular Biology 29 (1995) 479-489.
Journal of Life Sciences 6 (2012) 1317-1327
Helicobacter pylori
Biofilm Formation and Gene
Expression on Abiotic Surfaces Using
a Cyanobacterial Extract
Alba E. Vega1, Fabio A. Persia1, Gabriel A. Salinas-Ibañez1, Teresa Alarcón2, Stella M.C. Lucero Estrada1, Teresa I. Cortiñas1 and Humberto J. Silva1
1. Microbiology Laboratory, Department of Biochemistry and Biological Sciences, Faculty of Chemistry, Biochemistry and Pharmacy, National University of San Luis, San Luis 5700, Argentina
2. Department of Microbiology, University Hospital La Princesa, Princess Health Research Institute (IP), Madrid 28006, Spain
Received: July 10, 2012 / Accepted: September 17, 2012 / Published: December 30, 2012.
Abstract: The effects of a cyanobacterial extract (CE) on Helicobacter pylori biofilm formation onto hydrophobic and hydrophilic
abiotic surfaces and the expression of luxS, flaA, omp18, lpxD and ureA genes associated to biofilm were studied. NCTC11638 reference strain and HP796, a resistant clinical isolate, were grown in Mueller-Hinton broth supplemented with 5% fetal calf serum (FCS) or 1% CE. The ability to form biofilm, viability, morphological changes and gene expression of adhered H. pylori cells were determined. The strains were able to form biofilm on both surfaces with the nutritional supplements analyzed. H. pylori conserved a characteristic bacillary morphology and viability with CE. Cells attachment was higher with CE than FCS regardless of strains and surfaces. The most remarkable increase in gene expression was observed with the omp18 gene using the CE supplement, indicating the important participation of outer membrane proteins in biofilm establishment. The clinical isolate showed similar and even greater gene expression than the reference strain. The results obtained indicated that the nutrients provided by CE favored biofilm formation with retained pathogenicity that under certain conditions can occur in natural aquatic environments.
Key words: Helicobacter pylori, biofilm, cyanobacterial extract, gene expression.
1. Introduction
Helicobacter pylori is a human pathogen that colonizes the gastric mucosa, resulting in an acute inflammatory response and damage to epithelial cells, progressing to a number of disease states, including gastritis, peptic ulceration, and gastric cancer [1].
Recent studies indicate that H. pylori can exist both in human gastric mucosa and on abiotic surface forming biofilms, explaining the ability of the
organism to survive within and outside the host. H.
pylori biofilm survival in drinking water is discussed as a possible waterborne route of transmission [2, 3].
Corresponding author: Alba E. Vega, Ph.D., professor,
research field: microbiology. E-mail: [email protected].
The biofilm mode of growth confers a protective advantage to the bacteria which are physiologically distinct from the free-swimming counterpart of the same species, becoming more resistant to host defense and adverse environmental conditions [3], and up to 1,000 times more resistant to antibiotics [4, 5].
Helicobacter pylori Biofilm Formation and Gene Expression on Abiotic Surfaces Using aCyanobacterial Extract
1318
regulates a variety of physiological functions including the generation of bioluminescence, sporulation and the expression of virulence factors [8].
The luxS gene codifies for the autoinducer 2 (AI-2)
of QS system that is functional in stimulating the H.
pylori lux operon present in several gram-positive and gram-negative bacteria [9, 10].
The expression of luxS gene is essential for H.
pylori colonization of the human stomach and represents a significant indicator of biofilm production in which bacteria migrate and adhere forming microcolonies [9, 11].
Apart fromthe luxS gene, other genes are involved
in biofilm formation including genes encoding for
flagella (flaA), type I and type IV pili and surface
adhesins [12].
The flagella play important roles in biofilm formation in several gram-negative bacteria, both as surface adhesins and as providers of force-generating
motility [13]. The expression of H. pylori flaA is
dependent on luxS and the flaA transcription increases with culture density [14].
Bacterial outer membrane proteins (OMPs) are important for ion transport, osmotic stability, bacterial
virulence and adherence. Omp18 is a
peptidoglycan-associated lipoprotein precursor,
present in H. pylori, which is involved in adhesion to
gastric cell [15]. The cell envelope gene (lpxD)
encoding the UDP-3-0-(3-hydroxymyristoyl) glucosamine N-acyltransferase is up-regulated after adhesion to gastric cell in vitro [16]. Therefore, omp18
and lpxD genes could be involved in biofilm
formation.
H. pylori urease enzyme (ureA) is essential for pH regulation. The loss of urease activity acidifies the biofilm, decreasing the stability of the biofilm community [17].
Epidemiological data suggest that contaminated water is a potential reservoir for this microorganism
[18], even specific H. pylori DNA fragments have
been detected in river water [19, 20].
The presence of cyanobacteria is common in natural aquatic environments where they can generate blooms generally associated with a diverse community of heterothophic cultivable bacteria, some of them considered putative pathogens [21-23]. Aditionally a cyanobacterial extract (CE) obtained from the
cyanobacterium Nostoc sp. have nutrients, including
amino acid, soluble and crude proteins, carbohydrates, and different minerals, useful in the culture of fastidious bacteria [24].
The replacement of fetal calf serum (FCS) by CE in
liquid culture media of H. pylori increased growth and
cell viability with delays in the appearance of coccoid forms considered viable but nonculturable cells
(VBNC) [25, 26]. H. pylori adhesion and biofilm
formation is strongly affected by the presence of certain nutrients [27]. In this work the effects of CE on H. pylori biofilm formation and the expression of
luxS, flaA, omp18, lpxD and ureA genes associated to biofilm is presented.
2. Material and Methods
2.1 Strains and Media
H. pylori NCTC11638 (reference strain) a kind gift for Dr. Manuel López-Brea, Microbiology Service of Hospital Universitario de la Princesa, Madrid, Spain and HP796, characterized as clarithromicyn (CLA) and
metronidazole (MTZ) resistant strain and cagA + and
vacA s1m1, were used for this study. The bacterial
strains were routinely grown on Mueller-Hinton agar (MHA), supplemented with 7% horse blood (MHA-HB), and incubated in a microaerobic
atmosphere for 48 h at 37 °C. The identity of H. pylori
was confirmed by the following criteria: microaerophilic growth requirement, morphology, Gram’s stain, oxidase, catalase and urease reactions. Strains were stored in trypticase soy broth supplemented with 20% glycerol at -80 ºC until use.
2.2 Cyanobacterial Extract
Helicobacter pylori Biofilm Formation and Gene Expression on Abiotic Surfaces Using aCyanobacterial Extract
1319
Silva et al. [24].
2.3 Biofilm Assays
The ability of H. pylori to form biofilm on abiotic
surfaces was determined by total bacterial count and
viable cell count methods using 2 cm2 coverslips of
glass and polypropylene (PP) placed on 90 mm Petri dishes added with 12 mL of 0.3% glucose Mueller-Hinton Broth (MHB) supplemented with 5% FCS (MHB-FCS) or 1% CE (MHB-CE). The biofilm
formation was initiated by inoculating H. pylori cells
at a final concentration of 1 × 106 colony forming
units (CFUs)/mL in each plate. Cultures were incubated under microaerophilic conditions for 196 h at 37 ºC without shaking. In order to analyze biofilm evolution, coverslips were sampled at different times, rinsed three times with phosphate-buffered saline (PBS) to remove planktonic cells and biofilm debris and stained with 0.1% crystal violet (CV). Also rinsed coverslips were vortexed for 3 min in PBS to allow cell detachment from biofilm. Total bacterial count was performed using the Breed Counting Method by spreading 0.01 mL of resuspended biofilm cells over 1
cm2 microscope slide, the smear was dried and stained
with 0.1% CV for 30 min and the cells counted with an optical microscope. For viable cell counts, undiluted and 1:10 dilution resuspended biofilm cells were plated onto MHA-HB by duplicate. CFUs were counted after incubation for three days at 37 ºC.
2.4 Fluorescence Microscopy
To assess the membrane integrity of sessile bacteria, the coverslips were washed with 0.9% saline and stained with fluorescent dyes of the Live/Dead BacLight kit (Molecular Probes, Invitrogen Corporation) and incubated for 20 min in dark. Coverlips were observed with a Zeiss Axioplan 2 fluorescent microscope. Images were acquired by a camera using Axiovision 3.0 software.
2.5 Scanning Electron Microscopy
Biofilms were analyzed by scanning electron
microscopy (SEM) using a Zeiss LEO 1450VP microscope. Biofilms formed on abiotic surfaces were washed with 0.9% saline solution, dried and dehydrated using a graded ethanol series (70%, 95%, and 100% three times for 10 min each). Samples were mounted on aluminium stubs and coated with a gold layer and processed in a standard sputter. Observations were made at 20 KV.
2.6 Gene Expression
Cells coming from biofilms developed on the abiotic surfaces and planktonic cells were treated with TRIzol reagent (Invitrogen) for total RNA extraction.
The QS luxS, virulence ureA, flaA, adhesion lpxD,
omp18 and housekeeping 16S rRNA genes were
analyzed. cDNA was performed with random hexamer and 200 U Moloney murine leukaemia virus reverse transcriptase (Invitrogen). The identification of amplified fragments of 465-, 411-, 111-, 1001- 165- and 390- bp for, luxS, ureA, flaA, lpxD, omp18 and 16S rRNA genes respectively, was performed with 1.8% agarose gel electrophoresis at a constant voltage of 80 V/cm for 45 min. The gels were stained with GelRed nucleic acid gel stain (Biotium, Inc.), visualized under UV light and photographed. The DNA fragment size was determined by comparison with molecular weight markers with a range of 50 bp to 1,000 bp.
2.7 Statistical Analysis
The statistical analysis was determined by Tuckey-Kramer multiple comparisons tests. Differences were considered statistically significant at
P≤ 0.05.
Semi-quantification of the bands was performed with an image analyzer (ImageJ WCIF) against the constitutive gene 16S rRNA.
3. Results
3.1 Biofilm Formation
Helicobacter pylori Biofilm Formation and Gene Expression on Abiotic Surfaces Using aCyanobacterial Extract
1320
surfaces with the nutritional supplements analyzed (FCS or CE) (Table 1). The attachment of cells increased up to 96 h and in all the cases was higher
with CE (P ≤ 0.05) and with the HP796 strain (P ≤
0.05).
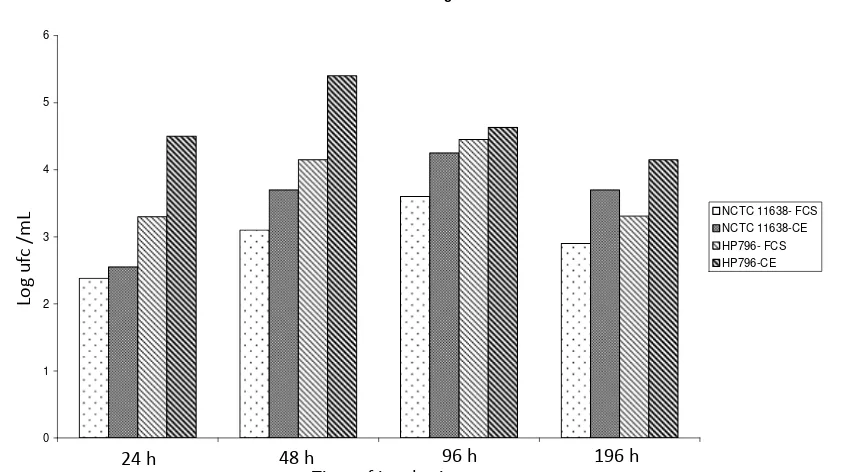
The viable counts for the strains attached to the hydrophilic surface (glass) at several times of incubation showed a similar pattern (Fig. 1). The same results were obtained with the hydrophobic surface PP (data not shown). NCTC11638 and HP796 strains biofilm on glass surface showed a major prevalence of bacillary forms at 196 h of incubation when culture was supplemented with CE (Fig. 2). The evolution of HP796 strain biofilms on PP surface at 48, 96 and
196 h, with CE or FCS supplements by optical microscopy is showed in Fig. 3.
The membrane integrity ofbiofilm cells developed
on glass surface with MHB-FCS and MHB-CE media after 96 h of incubation, was assessed by fluorescence microscopy using the Bacto Ligth Live/Dead kit, with green labeled cells (viable) and red cells (nonviable).
The biofilms of HP796 produced with MHB-FCS presented a higher proportion of nonviable coccoid forms than MHB-CE, additional, with MHB-CE the biofilm showed a great amount of spiral, viable, aggregated bacteria (Fig. 4). Viability staining of the biofilms showed a loss of viability after long term incubation.
Table 1 Attachment of H. pylori strains to abiotic surfaces in MHB supplemented with FCS or CE. Data are represented as means
log10 ± S.D. of total cells counts of three independent experiments.
Time (h)
Glass PP
NCTC11638 HP796 NCTC11638 HP796
FCS CE FCS CE FCS CE FCS CE
24 3.9 ± 0.2 4.3 ± 0.2 5.3 ± 0.1 5.9 ± 0.1 3.3 ± 0,2 3.5 ± 0.2 4.6 ± 0.1 4.9 ± 0.1 48 5.3 ± 0.1 5.6 ± 0.1 6.1 ± 0.2 6.7 ± 0.2 4.7 ± 0.1 5.1 ± 0.2 5.7 ± 0.2 6.7 ± 0.2 96 5.5 ± 0.2 5.8 ± 0.2* 6.3 ± 0.2 6.9 ± 0.2* 4.9 ± 0.1 5.3 ± 0.1* 5.9 ± 0.2 6.9 ± 0.2* 196 4.5 ± 0.1 5.1 ± 0.2 5.1 ± 0.2 5.7 ± 0.1 3.9 ± 0.2 4.9 ± 0.2 5.3 ± 0.1 5.5 ± 0.1
*: P≤ 0.05.
Biofilm formation on glass surface
0 1 2 3 4 5 6
24h 48h 96h 196h
time of incubation
log uf
c
/m
l NCTC 11638- FCS NCTC 11638-CE HP796- FCS HP796-CE
Fig. 1 Viable counts of H. pylori strains obtained on glass surface at specific time points in MHB added of FCS or CE. Results are
the average of three independent experiments.
24 h 48 h 96 h 196 h
Time of incubation
Log
uf
c
/m
L
Helicobacter pylori Biofilm Formation and Gene Expression on Abiotic Surfaces Using aCyanobacterial Extract
1321
Fig. 2 Biofilms of H. pylori strains on glass surface. Bacteria
were grown in (1) MHB-FCS and (2) MHB-CE. (a) H. pylori
NCTC11638 (1), (b) H. pylori NCTC11638 (2), (c) H. pylori
HP796 (1), (d) H. pylori HP796 (2). Biofilms were stained with
CV after 196 h of incubation. White arrows indicate bacillary forms and black coccoids forms. Results are representative of three independent experiments.
Similarly, H. pylori HP796 biofilm developed for
196 h on glass surface, examined by SEM, showed predominant bacteria with coccoid shapes and U forms using MHB-FCS and filamentous aggregates that conserved the characteristic spiral morphology with MHB-CE (Fig. 5).
3.2 Gene Expression
The expression of luxS, flaA, omp18, ureA, lpxD
and 16S rRNA genes of H. pylori strains in planktonic
and sessile cells on glass surface, at specific time points (48, 96 and 196 h) using MHB-FCS and MHB-CE media are shown in Fig. 6. The expression of the same genes in cells attached to polypropilene surface at 196 h is also shown in Fig. 6.
Fig. 3 Biofilm evolution of HP796 strain on PP surface. Bacteria were grown in MHB-FCS and MHB-CE media and stained with CV. (a), (b) and (c): MHB-FCS; D, (e) and (f): MHB-CE after 48, 96 and 196 h of incubation respectively. White arrows indicate bacillary forms and black arrows indicate coccoid forms. Results are representative of three independent experiments.
Fig. 4 Images of H. pylori NCTC11638 biofilm formed with (a): MHB-FCS and (b): MHB-CE. Bacteria were labeled with the
Bacto Live/Dead viability stain after 96 h of incubation, wherein live bacteria fluoresce green and dead bacteria fluoresce red. Results are representative of three independent experiments.
(a) (b)
(c) (d)
(a) (b) (c)
(d) (e) (f)
Helicobacter pylori Biofilm Formation and Gene Expression on Abiotic Surfaces Using aCyanobacterial Extract
1322
Fig. 5 Scanning electronic micrographs of H. pylori HP796 cells attached to coverslips after growth for 196 h with (a): MHB-FCS
and (b): MHB-CE, showing aggregated bacteria with coccoid shape and U forms in (a) and aggregated bacteria with filamentous characteristic morphology in (b). Scale bar represents 2 µm. Results are representative of three independent experiments.
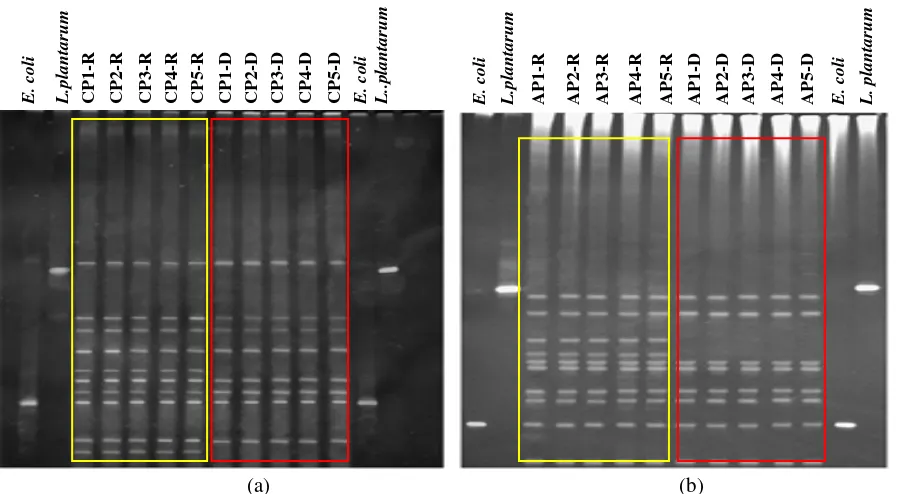
Fig. 6 Gene expression of H. pylori strains in MHB-FCS and MHB-CE media at specific time points. Planktonic (p) and sessile (s)
cells on glass (g) and PP surfaces were assessed by PCR for 16S rRNA, luxS, ureA, flaA, omp18, lpxD genes. (a) Reference strain; (b)
HP796 H. pylori strain. Lines 1, 5 and 9: p cells in MHB-FCS; lines 2, 6 and 10: p cells in MHB-CE; lines 3, 7, 11 s cells in
MHB- ;FCS; lines 4, 8, 12 s cells in MHB-CE; lines 13, 14 s cells in MHB-FCS and MHB-CE respectively; lines 1-12: g surface; lines 13, 14: PP surface; lines 1-4: 48 h; lines 5-8: 96 h; lines 9-12: 196 h; lines 13 and 14 s cells attached on PP at 196 h. Results are representative of three independent experiments.
The relative increase in gene expression of H.
pylori strains forming biofilm on glass surface in relation to that of planktonic cells using MHB-FCS and MHB-CE media at specific time points is shown in Table 2.
The luxS gene expression of the HP796 strain
supplemented with CE was 1.7 fold higher while the reference strain 0.9 fold inferior after 48 h of biofilm evolution.
The expression of H. pylori HP796 flaA gene
increased 1.5 fold using MHB-CE meanwhile with MHB-FCS the expression was 0.7 fold inferior than
planktonic cells at initial stage of biofilm evolution
(48 h). At this stage H. pylori HP796 strain showed
higher flaA expression (p ≤ 0.05) than the reference
strain. No significant differences were observed in the expression of this gene between strains and nutritional
supplements at other biofilm stages. The ureA gene
expression increased 1.2 fold and 1.5 fold for reference and HP796 strains respectively at initial stage of biofilm evolution with MHB-FCS and then diminished at later stages. While, with MHB-CE
ureA gene expression remained constant along
biofilm evolution (Table 2). (b)
(a)
Helicobacter pylori Biofilm Formation and Gene Expression on Abiotic Surfaces Using aCyanobacterial Extract
1323
No significant differences were observed in lpxD
expression during biofilm evolution with respect to planktonic cells, using both supplements. The most
remarkable increase in gene expression (P≤ 0.05) was
observed with the omp18 gene at 196 h. For the HP
796 and reference strain, 4.6 fold and 2.4 fold increases resulted using the MHB-CE medium, values that were higher than that obtained in the biofilm developed with the MHB-FCS medium with 3.1 fold and 1.9 fold increases respectively.
The gene expression of biofilms evolving on the PP surface for the strains showed similar results to those
obtained on glass surface. Apart from omp18 gene
there were no statistical differences in gene expression between both supplements and strains (Table 3). However, for this gene, a clear tendency of higher expression was observed for the HP796 strain using
the MHB-CE medium (P = 0.3).
4. Discussion
Epidemiological studies performed in a wide variety of natural habits show that bacteria live in complex communities usually attached within a structured biofilm ecosystem and not as planktonic cells [28].
4.1 Biofilm Formation
H. pylori has the ability to form biofilms in vitro on different surfaces and on the human gastric mucosa [2, 29, 30].
H. pylori strains were able to form biofilms on glass and polypropilene surfaces growing in MHB-FCS or MHB-CE media. Large aggregates of predominant filamentous forms were characterized in a mature biofilm of 48 h, independently of the surface and nutritional supplement used. However, for the same
Table 2 Relative gene expression of H. pylori strains forming biofilm on glass surface in relation to planktonic cells, using
MHB-FCS and MHB-CE media at specific time points.
NCTC11638 HP796
Time (h) 48 96 196 48 96 196
(a) Using FCS as supplement
luxS 0.9 1.2 1.0 1.0 1.0 1.5
flaA 1.1 1.1 0.7 0.7 0.8 0.9
ureA 1.2 0.9 0.7 1.5 0.8 1.1
lpxD 0.8 0.8 0.9 0.7 0.8 1.4
omp18 1.0 1.6 1.9 1.1 1.8 3.1*
(b) Using CE as supplement
luxS 0.9 1.4 1.3 1.7 1.1 1.5
flaA 0.7 0.9 0.8 1.5 1.0 1.1
ureA 0.7 0.9 1.0 1.0 1.1 1.2
lpxD 0.7 1.0 0.9 1.1 1.5 1.2
omp18 0.8 2.3 2.4 1.3 3.7 4.6*
*: P≤ 0.05.
Table 3 Relative gene expression of H. pylori strains forming biofilm on polypropylene surface in relation to planktonic cells using
MHB-FCS and MHB-CE media at 196 h.
Genes NCTC11638 HP796
FCS CE FCS CE
luxS 1.0 1.9 1.3 1.4
flaA 0.7 1.1 1.2 1.6
ureA 0.6 1.0 0.8 0.9
lpxD 0.8 1.2 1.0 1.1
omp18 2.0 3.1 3.1 3.8*
Helicobacter pylori Biofilm Formation and Gene Expression on Abiotic Surfaces Using aCyanobacterial Extract
1324
time of incubation H. pylori biofilm developed with
Brucella broth supplemented with 2% FCS presented cells with a predominant coccoid morphology [11].
During the process of biofilm formation the proportion of coccoid to spiral forms clearly varied
with the supplement used. H. pylori conserved the
characteristic bacillary morphology and viability with CE which correlated with a major attachment rate in relation to that with FCS regardless of strains or surface. These results can be attributed to the different nutrient potential provided by the two supplements.
The virulent strain HP796, Cla- and Mtz-resistant, had higher ability to form biofilm in vitro compared with the susceptible reference strain. For this strain the
viable cell count significantly (p ≤ 0.05) increased
with both surfaces and nutrition supplements. Relation not found between resistant and susceptible virulent genotypes and the ability to form biofilm by Cellini et al. [11]. In this sense, the study of a greater number of
H. pylori strains is required to improve the knowledge of biofilm formation by resistant and virulent strains of epidemiological importance.
While the attachment of H. pylori cells increased
with incubation time, a difference of two logarithmic units was observed between total and viable counts at 196 h of biofilm formation, indicating a loss in the culturability status of the bacteria in the biofilm wich was associated with morphological changes.
The increase of coccoid cells considered as VBNC (viable but nonculturable state) has been observed during dynamic process of biofilm formation [31, 32]. The results obtained showed that the morphological
condition of H. pylori did not affect its ability to
surface attachment. In fact H. pylori may present a
patchy distribution forming cluster of bacillary and coccoid cells as a strategy of preservation during colonization in the host [2]. The presence of these dormant cells in the bacterial biofilm can be considered one cause of treatment failure by antibiotics.
The fluorescent dyes of the Live/Dead kit with the
Syto9/PI reagents [32-34] allowed the characterization of cell viability and the distinction of different morphotypes present in the biofilms. The use of this technique demonstrated that CE was superior to FCS as nutrient supplement in maintaining the viability of
H. pylori within the biofilm structure, according to the biofilm viable counts obtained. A higher proportion on nonviable coccoid forms was present with FCS in glass biofilms of 196 h evolution. Observations of biofilm with SEM confirmed that the agglomerates were mostly constituted by coccoid or U-shaped bacteria in contrast to the spiral morphology predominant in biofilms developed with CE. As far as the bacillary form is associated with a more infective state [35], the maintenance of spiral morphology for longer periods of time indicates that CE can provide both essential nutrient and protection against adverse environmental conditions outside the human host.
4.2 Gene Expression
The development of a mature biofilm is achieved through a number of sequential steps, each of which is marked by changes in gene expression in response to environmental cues and cell-cell signaling [36], indicating that a biofilm is a highly regulated developmental biological system.
Depending on the bacterium, the autoinductor AI-2
encoded by the luxS gene, plays a role in motility,
pathogenicity and biofilm formation [37].
The QS system, normally associated with the regulation of virulence factors, could also regulate the various phases of biofilm development from the initial adhesion to the final detachment of cells [38, 39].
H. pylori upon adhesion to gastric epithelial cells change only a short proportion of 1,542 genes analyzed with 22 up and 21 down regulated genes [16]. As adhesion is the first step in biofilm formation it was of interest to analyze the gene expression of QS
luxS, virulence ureA, flaA, adhesion lpxD, omp18 and
housekeeping 16S rRNA genes of H. pylori in the
Helicobacter pylori Biofilm Formation and Gene Expression on Abiotic Surfaces Using aCyanobacterial Extract
1325
The QS luxS gene expression of the HP796 strain
supplemented with CE was 1.7 fold up regulated while the reference strain 0.9 fold down regulated
after 48 h of biofilm evolution. The virulence ureA
gene, responsible for the ureasa enzyme required for colonization and maintenance of the organism in hostile environments, showed similar levels of expression in biofilm and planktonic cells. The HP796
strain increased 50% the ureA gene expression at the
initial state of the biofilm using both nutrient supplements.
Flagella are necessary for colonization of the gastric mucosa and play an important role in biofilm formation suppressing repulsive forces of hydrophobic surfaces [3, 13]. The expression of flaA gen was strain
and supplement dependent. The flaA gene of the
HP796 strain, with higher ability to form biofilm, was 1.5 fold up regulated in comparison to planktonic cells while the reference strain was 0.7 fold down regulated at the early stage of biofilm formation using the
MHB-CE medium. The lpxD gene with MHB-CE was
up regulated with both strains and surfaces in biofilms of 196 h evolution.
The most remarkable increase in gene expression
was observed with the omp18 gene with the CE
supplement producing the most effective induction, indicating the important participation of outer membrane proteins in cell aggegation and biofilm establishment. Both strains up regulated the
expression of omp18 gene, however the strain HP796,
ClaR and MtzR, and genotype virulent cagA+, vacA+ and iceA1, showed similar, and even greater expression than the reference strain.
The soluble metabolites present in the CE increased
biofilm formation, viability and gene expression of H.
pylori. The CE was produced as a dry powder using concentrated biomass of the filamentous
cyanobacterium Nostoc sp., heat treated with
subsequent lyophilization of the soluble material released. The high protein content of CE [24] could
stimulate the initial attachment of H. pylori cells to
abiotic surfaces as far as its adherence is a protein mediated process [27]. Although quite different in procedure a similar release of soluble and non-soluble matter into the surrounding medium can be produced growing dense populations of filamentous cyanobacteria in enriched water lake, followed by cells collapse and liberation of fresh organic material, with an expected positive response of heterotrophic bacteria, ciliates and nanoflagellates [21].
5. Conclusion
The main contribution of this work was to
demostrate the effect of a cyanobacterial extract on H.
pylori biofilm formation. CE markadely increased the
survival, gene expression and maintenance of H.
pylori cells bacillary forms in the biofilm. The effect was superior to that obtained using FCS. The effect of the cyanobacterial extract favouring biofilm formation of H. pylori with retained pathogenicity can be considered a fact of possible occurence in water bodies where cyanobacteria are present that might constitute a novel source of transmission.
Acknowledgments
The authors thank Patricia Gomez, Patricia Vallejos and Ruben Majul for providing the gastric biopsy specimens. The authors have no conflict of interest to declare. This work was supported by funds of Science and Technology Project 9303 and Project 0310 from the National University of San Luis.
References
[1] H.L. Mobley, G.L. Mendz, S. Hazel, Helicobacter pylori: Physiology and Genetics, American Society for Microbiology Press, Washington, DC, 2001, Chapter 2, pp. 7-18.
[2] M.A. Carron, V.R. Tran, C. Sugawa, J.M. Coticchia, Identification of Helicobacter pylori biofilms in human gastric mucosa, Journal of Gastrointestinal Surgery 10 (2006) 712-717.