R
etrotransposons were first characterized in animal and yeast genomes, but evidence has accumulated in recent years to show that they are present in all plant genomes and can constitute a very large part of some of them. They are one of the two classes of transposable elements, defined according to their mode of propagation: retrotransposons (also termed class I elements) transpose via an RNA intermediate; and class II el-ements only use DNA in movement. Representatives of all types of retrotransposons have been detected in plant genomes1,2. A growing body of evidence shows that the activity of these retro-transposons is, as in animal systems, tightly controlled, and that abiotic and biotic stresses are major factors in their transcriptional and transpositional activation.
Regulation of retrotransposition
Retrotransposons can be separated into two major subclasses that differ in their structure and transposition cycle (Fig. 1). Elements of subclass I are bounded by two long terminal repeats (LTRs) and are termed LTR retrotransposons; elements of subclass II do not possess LTRs and are therefore termed non-LTR retrotrans-posons. Both these subclasses form a DNA daughter copy by reverse transcription of an RNA template, and their replication cycle involves an intermediate cytoplasmic step. This replicative transposition mechanism means that retrotransposons are poten-tially very invasive. To ensure the viability of their host, and hence their own survival, retrotransposition is tightly controlled (Fig. 2). This control involves element-encoded functions and host factors. One of the major control steps is transcription, which determines both the production of the RNA template and the synthesis of mRNAs required for protein synthesis. In LTR retrotransposons, transcriptional control involves cis-regulatory sequences that are usually found in the element’s LTR, in particu-lar the U3 region located upstream of the transcription start site (Fig. 2), or in downstream, untranslated sequences.
A survey of active plant retrotransposons
Transpositional activity has been reported for only a few el-ements, mostly those belonging to the LTR retrotransposon sub-class (Table 1). Most of these were isolated after transposition into or next to a host gene (Table 1). A few mobile elements were first characterized by PCR amplification of genomic DNA or cDNA, but were subsequently shown to transpose. Evidence for transpo-sitional activity can also be inferred from the analysis of LTR sequences, which are identical in newly transposed copies: this is the case of the Osser element, which is probably active, although direct transposition has not been reported3. So far, however, func-tional copies able to transpose in foreign species have only been characterized for Tnt1A (Ref. 4) and Tto1 (Ref. 5).
Homologous transcripts have been reported for a larger number of elements (Table 1). In several cases, however, the nature of the transcript has not been fully established and it could derive from co-transcripts originating from external, upstream promoters (Box 1). Specific transcripts, starting in the element’s LTR or at its 5′ -extremity for non-LTR retrotransposons, were demonstrated for Tnt1A, Tto1, BARE-1, Tos17, Huck (Z. Avramova, unpublished) and possibly SIRE-1, as well as for the short interspersed nuclear elements (SINEs) S1Bnand TS elements (see Table 1). Definitive evidence of LTR transcriptional ability was further obtained for Tnt1A, BARE-1 and Tto1 (Table 1) by using constructs in which re-porter genes were placed under the control of the element’s LTR.
Regulation of the activity of plant retrotransposons Developmental regulation
In animals and yeast, the expression of retrotransposons is under the control of hormonal and developmental factors. A general pic-ture of expression is difficult to establish for many plant retrotrans-posons, because comparative studies in different tissues have not been done. However, the expression of the most well-characterized plant retrotransposons is not constitutive. Developmental regu-lation has been shown for Tnt1, which is only expressed in roots and then only at low levels6
; for Tto1, Tos10 and Tos17, which are not expressed in leaf tissues7,8; and for the mobile B5, Hopscotch,
Stonor and Magellan elements, which are not expressed in most plant tissues (S. Wessler, unpublished). Expression of the maize PREM-2 element was detected only in early microspores9
. How-ever, the expression of Opie, Huck and Cinful (Refs 10 and 11), and of BARE-1 (Ref. 12), was detected in leaf tissues.
Stress activation: the in vitro track
A common feature of most retrotransposons is that they are activated by stress and environmental factors. The most well-characterized plant retrotransposons are particularly affected by protoplast isolation or in vitro cell or tissue culture (Table 1). Insertion of retrotransposons into coding sequences after proto-plast or cell culture was demonstrated in tobacco13
, in Nicotiana plumbaginifolia (C. Meyer, unpublished) and in rice7, indicating that retrotransposition might make a significant contribution to somaclonal variation. The first direct evidence of activation of a plant retrotransposon by stress came from the discovery that the expression of Tnt1A was highly induced in protoplasts isolated from tobacco leaf tissue6
. In accordance with this, Tnt1A transpo-sition into the tobacco nitrate reductase (nia) gene was detected in plants regenerated from protoplast-derived cell cultures13
. How-ever, Tnt1A transcription was not detectable in suspension cell cultures, even though a weak increase in copy number was ob-served8. In contrast, both the expression and the transposition of
Activation of plant retrotransposons
under stress conditions
Marie-Angèle Grandbastien
Tto1, Tos10 and Tos17 is activated in cell
cultures7,8. Tto1, but not Tos17, expression is further increased by protoplast isolation. The barley BARE-1 element is expressed in callus tissues, as well as in leaf-derived protoplasts, although protoplast expression appears to derive from BARE-1 expres-sion in the leaves12. Transcripts of the soy-bean S1BnSINE elements were detected in callus cultures14, and protoplast-specific RNA sequences of LTR retrotransposons were characterized in potato15.
A link with defence responses?
Protoplast isolation, as well as cell and cal-lus culture, induces major modifications of cell metabolism and gene expression16,17
. In leaf-derived protoplasts, the former meta-bolic activity of the leaf cell is replaced by a new programme. This is characterized by the activation of growth- and stress-related genes (e.g. defence genes, which are activated after pathogen attack). Growth-related genes are probably involved in the re-initiation of cell division; the activation of stress-related genes might be a conse-quence of the original wounding. Proto-plast isolation also involves enzymatic degradation of the cell wall, using extracts from phytopathogenic fungi. The acti-vation of stress-related genes might thus also result from cell wall hydrolysis or from pathogenic compounds present in fungal extracts. Growth- and defence-related genes are also expressed in callus and cell cultures, suggesting that the pro-grammes of callus tissues are similar to those induced after wounding and during callus formation in the plant, involving both a stress response and cell division16.
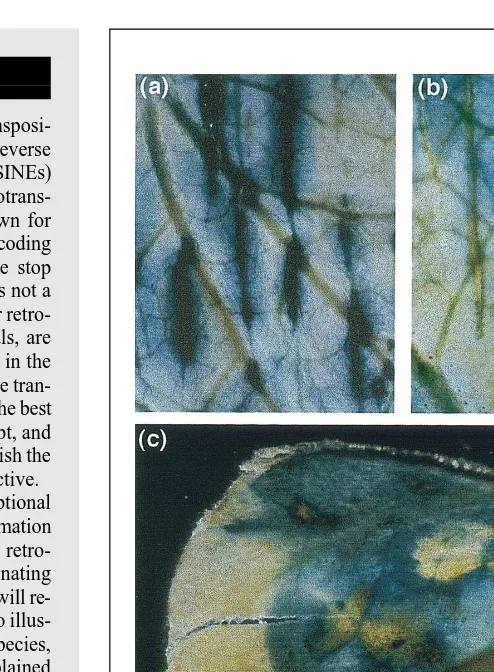
The activation of several plant retrotrans-posons in these particular conditions leads to the question as to whether their expres-sion is linked to the activation of cell di-vision programmes or to the activation of stress responses, or to both. Partial answers were provided by further studies of the expression of the tobacco Tnt1A and Tto1 elements. Tnt1A protoplast-specific ex-pression results mostly from the effect of fungal extracts, and the Tnt1A promoter is also activated by other compounds of mi-crobial origin, salicylic acid, wounding (Fig. 3), and viral, bacterial or fungal at-tacks18
. Similarly, the expression of the
Tto1 element is induced by viral attacks,
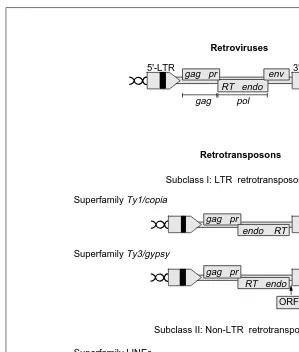
wounding, salicylic acid and jasmonate19,20 . The expression of the two best-character-ized plant retrotransposons is thus induced by different biotic or abiotic factors that can elicit plant defence responses. Further analysis suggested that Tnt1A expression was tightly linked to the early steps of the defence gene activation pathways18. Fig. 1. Overall organization of the different types of retrotransposons in comparison with
retroviruses, with classification according to Ref. 30. Elements of the subclass I have an overall organization similar to retroviruses. They are bounded by long terminal repeats (LTRs) that contain signals for initiation and termination of transcription, and carry one or several open reading frames (ORFs) with coding potential for the structural and enzymatic proteins needed for the retrotransposition cycle: the gag domain, encoding proteins that form the nucleocapsid core; the protease (pr) domain, encoding proteins that are involved in the maturation of the different proteins; the reverse transcriptase (RT) domain, encoding the enzymes responsible for the creation of a DNA copy from the genomic RNA template; and the endonuclease (endo) domain, encoding proteins necessary for the integration of the DNA copy into the host genome. LTR retrotransposons form cytoplasmic, virus-like particles (VLPs) in which the RNA template is reverse transcribed into a DNA daughter copy. LTR retrotransposons are further divided into two superfamilies: the Ty3/gypsy superfamily, in which the organization of the coding domain is the same as that of retroviruses; and the
Ty1/copia superfamily, in which the endo domain is placed upstream of the RT domain. The
major difference between retroviruses and LTR retrotransposons is that the latter do not encode the envelope (env) gene responsible for the formation of the extracellular infectious virion. However, the boundaries between retroviruses and retrotransposons appear increas-ingly blurred, as several members of the Ty3/gypsy superfamily also contain an additional open reading frame (ORF3), sometimes encoding an env-like gene, and might thus represent intermediates between LTR retrotransposons and retroviruses. Non-LTR retrotransposons do not contain LTRs and are terminated by an A-rich tail ([A]n). Long interspersed nuclear ele-ments (LINEs) generally contain two ORFs, the second showing similarities to the RT domain and the first encoding a putative gag-type nucleic-acid-binding protein (‘gag’). Although LINEs do not contain a recognizable endo domain, some elements might contain a putative nuclease domain (N ). The retrotransposition cycle of LINEs is not well understood, but it has been proposed that they might form cytoplasmic particles able to carry the RNA template into the nucleus, where reverse transcription would occur simultaneously with inte-gration31
. Short interspersed nuclear elements (SINEs) have no coding capacity and are thought to use foreign RT domains to achieve their life cycle, through incorporation of their RNA into the cytoplasmic particles of LINEs (Ref. 31).
Subclass I: LTR retrotransposons
Subclass II: Non-LTR retrotransposons
Retrotransposons Retroviruses
RT endo env gag pr
5'-LTR 3'-LTR
pol gag
Superfamily Ty1/copia
Superfamily Ty3/gypsy
RT endo gag pr
ORF3 gag pr
endo RT
Superfamily LINEs
Superfamily SINEs
[[A]n gag N RT
However, the expression of Tto1, but not of Tnt1A, in suspension cell cultures shows that the activating pathways differ for each element and that Tto1 expression might be under a dual control such as stress and cell division, as observed for several genes acti-vated after protoplast isolation17
. Interest-ingly, expression of the rice Tos17 element in cell cultures is not enhanced by proto-plast isolation7, suggesting that the control of Tos17 differs from that of Tnt1 and Tto1. Whether the link between retrotranspo-son activation and plant defence responses can be extended to elements other than Tnt1 and Tto1 is not yet known. However, other transcribed retrotransposon sequences have also been detected in stressed tissues, such as Tto5 (detected in tobacco treated with salicylic acid or after viral inoculation19
) and the Tpt sequences (detected in anaerobi-cally stressed seedlings of loblolly pine; C. Kinlaw, unpublished). Transposition of the maize Bs1 element was also detected after viral infection21, although a direct link be-tween Bs1 movement and infection has not been established.
Tnt1A and Tto1 regulatory sequences and defence genes
Promoter structure and function have been studied in detail for Tnt1A, Tto1 and BARE-1. Tandemly repeated cis-regulatory sequences were identified in the U3 region of Tnt1A and Tto1: a 31 bp repeat, the BII box, pres-ent in three or four copies in transcription-ally active elements, is involved in Tnt1A activation by protoplast isolation and fun-gal elicitins18,22; and a repeated 13-bp mo-tif is involved in Tto1 expression in callus and after wounding or jasmonate appli-cation (S. Takeda et al., unpublished). Puta-tive regulatory motifs were also detected in U3 regions of Tos17 (Ref. 7) and BARE-1 (Ref. 12). The specificity of expression of
BARE-1 in leaves or calli also involves the
alternative use of different LTR promoters12.
Interestingly, Tnt1A and Tto1 repeated cis-acting motifs share similarities with a motif involved in the activation of several plant defence genes, the H-box23,24
. In addition, the Tnt1A U3 region contains other sequences highly similar to regulatory motifs of stress-induced plant genes. These similarities provide a plausible explanation for the molecular basis of both Tnt1 and Tto1 acti-vation by stresses and pathogen attacks. A MYB-related factor,
LBM1, involved in Tto1 transcriptional activation in protoplasts
through specific binding to the 13-bp motif, was recently charac-terized (K. Sugimoto, S. Takeda and H. Hirochika, unpublished). However, LBM1 is not expressed in suspension cell cultures, indi-cating that other factors are involved in Tto1 activation in these conditions.
These results illustrate the complexity of retrotransposon tran-scriptional regulation, and indicate that the subtle differences between Tnt1 and Tto1 stress activation features might be linked to the presence in their LTR of several cis-regulatory sequences each capable of responding to a different stimulus. Although some
of the molecular transduction pathways involved in the expression of Tnt1 and Tto1 are different, the overall result appears to be sur-prisingly similar – increased expression in response to a diverse array of stress conditions that activate plant defence responses.
Evolution of retrotransposon transcriptional control
Reverse transcription is prone to error. As a consequence, a retro-transposon can generate a population of different but closely re-lated daughter copies, and this variability might play a role in the evolution of the control of retrotransposon activity. For example, the generation of transcriptionally inactive Tnt1A copies, through specific deletions of BII sequences, is a frequent event22. As also demonstrated for SINE S1Bnelements
14
, only a limited number of family members are responsible for the transcripts, possibly in order to limit the hazardous effects of growing populations of retrotransposons22. The variability of retrotransposon populations might also constitute a reservoir of potentially useful genomes, endowing retrotransposon populations with high adaptability22, as
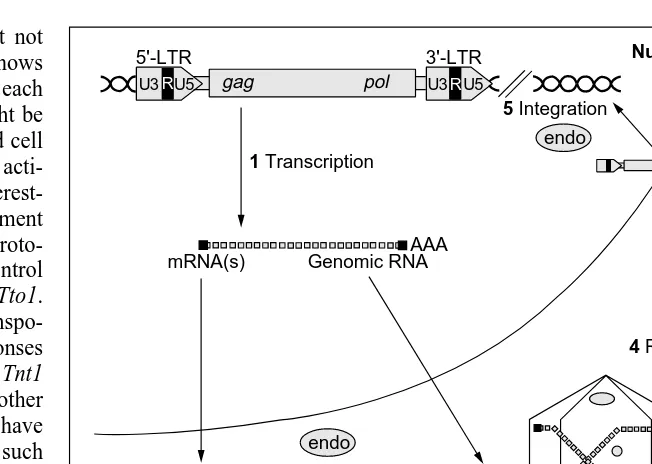
Fig. 2. The major control points of the transposition cycle of long terminal repeat (LTR)
retrotransposons. (1) Transcription begins in the 5′-LTR, at the boundary between the U3 and R domains, and terminates in the 3′-LTR, at the boundary between the R and the U5 domains, to produce a length RNA bounded by the redundant R domain. The full-length RNA will serve as a template for reverse transcription, as well as an mRNA template for the production of the proteins. (2) Control of the gag : pol ratio during protein synthesis is necessary, structural gag protein being required in large quantities for the assembly of virus-like particles (VLPs). The gag and pol domains are usually found in different frames, and the pol products are synthesized as a gag–pol fusion polyprotein resulting from a trans-lational frame shift. Alternative strategies, such as transcript splicing or specific degra-dation of the pol domain of the gag–pol polyprotein, are used by elements in which the gag–pol domains are in the same reading frame. (3) RNA packaging and VLP assembly are tightly dependent on specific interactions between the genomic RNA and gag nucleic-acid-binding domains. A parameter essential to this step is the activation of the protease, which catalyses the maturation of the gag–pol polyprotein during VLP assembly, thus ensuring the incorporation of functional enzymatic proteins into the VLP. (4) Reverse transcription is a very complex process that depends on the availability of a particular host tRNA and leads to the generation of a linear extrachromosomal DNA form bounded by two identical LTRs. (5) Integration involves the processing of the ends of the linear extrachromosomal DNA form and the joining of the retrotransposon DNA to the cleaved host DNA. Both steps are catalysed by the endo-encoded protein, but the mechanism by which the DNA daughter copy is transported to the nucleus is not well understood.
5'-LTR 3'-LTR gag pol
U3RU5 U3RU5
Nucleus Cytoplasm
AAA mRNA(s) Genomic RNA
1 Transcription
2 Protein synthesis
endo
RT VLP gag
RT RNA DNA
5 Integration
endo
4 Reverse transcription
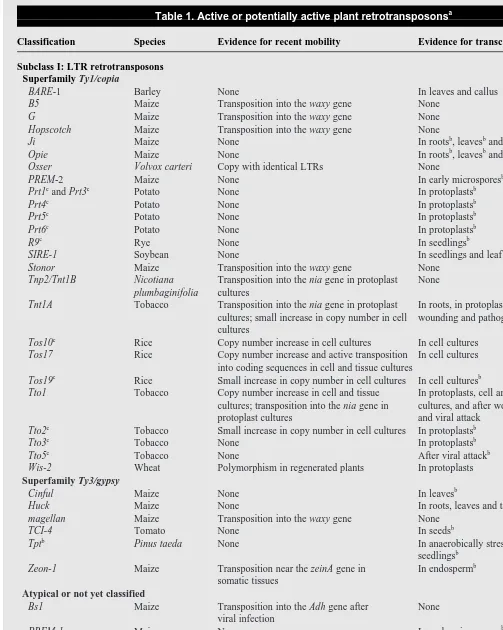
Table 1. Active or potentially active plant retrotransposonsa
Classification Species Evidence for recent mobility Evidence for transcripts Refs
Subclass I: LTR retrotransposons Superfamily Ty1/copia
BARE-1 Barley None In leaves and callus 12
B5 Maize Transposition into the waxy gene None 34
G Maize Transposition into the waxy gene None 34
Hopscotch Maize Transposition into the waxy gene None 25
Ji Maize None In rootsb
, leavesb
and tasselsb
10, 11
Opie Maize None In rootsb
, leavesb
and tasselsb
10, 11
Osser Volvox carteri Copy with identical LTRs None 3
PREM-2 Maize None In early microsporesb 9
Prt1cand Prt3c Potato None In protoplastsb 15
Prt4c Potato None In protoplastsb 15
Prt5c Potato None In protoplastsb 15
Prt6c Potato None In protoplastsb 15
R9c Rye None In seedlingsb 33
SIRE-1 Soybean None In seedlings and leaf tissues 35, d
Stonor Maize Transposition into the waxy gene None 34, e
Tnp2/Tnt1B Nicotiana Transposition into the nia gene in protoplast None f
plumbaginifolia cultures
Tnt1A Tobacco Transposition into the nia gene in protoplast In roots, in protoplasts, and after 18
cultures; small increase in copy number in cell wounding and pathogen attacks cultures
Tos10c
Rice Copy number increase in cell cultures In cell cultures 7
Tos17 Rice Copy number increase and active transposition In cell cultures 7
into coding sequences in cell and tissue cultures
Tos19c
Rice Small increase in copy number in cell cultures In cell culturesb
7
Tto1 Tobacco Copy number increase in cell and tissue In protoplasts, cell and tissue 8, 20, g cultures; transposition into the nia gene in cultures, and after wounding
protoplast cultures and viral attack
Tto2c
Tobacco Small increase in copy number in cell cultures In protoplastsb
8
Tto3c
Tobacco None In protoplastsb
8
Tto5c
Tobacco None After viral attackb
19
Wis-2 Wheat Polymorphism in regenerated plants In protoplasts 32, h
Superfamily Ty3/gypsy
Cinful Maize None In leavesb
10, 11
Huck Maize None In roots, leaves and tassels 10, 11
magellan Maize Transposition into the waxy gene None 37
TCI-4 Tomato None In seedsb
38
Tptb
Pinus taeda None In anaerobically stressed i
seedlingsb
Zeon-1 Maize Transposition near the zeinA gene in In endospermb
36 somatic tissues
Atypical or not yet classified
Bs1 Maize Transposition into the Adh gene after None 21
viral infection
PREM-1 Maize None In early microsporesb
39
TOC1 Chlamydomonas Transposition into the OEE1 gene and Yes 40
reinhardtii increase in copy number during mitotic growth
Subclass II: Non-LTR retrotransposons Superfamily SINEs
S1Bn Rapeseed None In shoots, roots and callus 14
TS Tobacco None By in vitro transcription 41
aThe table includes all those plant retrotransposons for which transcripts or mobility have been reported. However, any transcripts only shown to be expressed
from foreign promoters are excluded; elements showing intervarietal or interallelic polymorphisms, indicative of fairly recent activity, are also excluded, because this is a poor criterion for present activity. bThe possibility that transcripts are initiated from external upstream promoters has not been ruled out. cPartial
sequences isolated by PCR methods. dH. Laten, unpublished. eS. Marillonnet and S. Wessler, unpublished. fC. Meyer, unpublished. gP. Grappin and M-A.
proposed for retrovirus quasispecies. For instance, Tnt1A and S1Bn transcript populations vary between different cellular contexts14,22, and this variability affects mostly regulatory sequences in the case of Tnt1A. Retrotransposon populations might thus evolve differ-ently, in particular in their expression features, if maintained under different environmental conditions.
Rapid evolution of retrotransposon regulatory sequences is also illustrated by the observation that the Tnt1 family is composed of three different subfamilies, Tnt1A, Tnt1B and Tnt1C, highly similar in their coding domains, but completely different in their regulatory
U3 sequences22
. The three subfamilies appeared after a period of evolution in which regulatory sequences diverged widely while flanking regions remained more constant. The stress-activated
Tnt1 elements cloned after transposition in tobacco are members
of the Tnt1A subfamily, which are predominant in tobacco. How-ever, whether divergences in the regulatory sequences between the three subfamilies are linked to differences in their expression features remains to be determined. Interestingly, the three sub-families were present before Nicotiana speciation, but are differ-ently distributed in each Nicotiana species.
Box 1. What is an active retrotransposon?
Retrotransposons encode for proteins involved in retrotransposi-tion, and produce RNA both for protein production and for reverse transcription [although short interspersed nuclear elements (SINEs) are an exception]. As for class II elements, a defective retrotrans-poson can be trans-activated. Transposition has been shown for
Bs1, Zeon-1, G, Stonor and TOC1, which all lack important coding
regions or have coding domains interrupted by unsuitable stop codons. The presence of functional gag–pol domains is thus not a prerequisite for transposition, provided signals important for retro-transposition, such as priming sites or encapsidation signals, are still present. However, a retrotransposon will not transpose in the absence of the genomic RNA used as the template for reverse tran-scription. In terms of mutagenic impact on the host genome, the best criterion for activity is thus the ability to produce a transcript, and this criterion has been used, in addition to mobility, to establish the list of plant retrotransposons that are active or potentially active.
For some elements, preliminary information on transcriptional activation is provided by partial cDNA sequences. This information should be taken cautiously, because transcripts containing retro-transposon sequences can derive from cotranscripts originating from external, upstream promoters. Their expression pattern will re-flect the activation of the foreign promoters only. This is also illus-trated by the detection, in several cases, of multiple RNA species, or even smears or RNA of variable sizes, which are best explained by simultaneous cotranscription from different cellular promoters. However, this does not exclude the possibility that a specific el-ement transcript could also be expressed, as demonstrated for Tnt1,
TOC1, Tos10 and Tos17, and S1Bn. Different transcript sizes can
also be produced internally from modified members of a given fam-ily, such as deleted copies or copies carrying additional sequences. For instance, the mobile G element of maize is a deleted derivative of the B5 element. Similarly, the maize Cinful, Zeon-1 and Cin1 elements are very closely related, Zeon-1 being a copy in which most coding domains, except gag, are deleted and replaced by se-quences of unknown origin; Cin1 represents a single long terminal repeat (LTR) of the same family. The transposition of G and B5, in particular, has required two mRNA species of different sizes. Cotranscripts originating from the transcriptional start site of the element and proceeding into downstream cellular sequences also cannot be excluded. It thus appears that, if retrotransposon se-quences isolated from cDNA clones are by themselves not con-vincing evidence for the transcriptional activation of the element, then transcripts of abberrant or multiple sizes could still provide information on the expression conditions of a given element. Never-theless, active retrotransposons have been successfully isolated after preliminary characterization of partial sequences through reverse transcription PCR (Tto1 and Tos17) or even genomic PCR (Tos10). Tto1, Tos10 and Tos17 PCR sequences were also shown to hybridize to an RNA of adequate size, thus increasing the prob-ability that this transcript was specific. It thus appears that, pro-vided some precautions are taken, active retrotransposons can successfully be isolated via their transcription products.
Fig. 3. Transcriptional activation of the Tnt1A tobacco
Why do retrotransposons contain regulatory motifs similar to those of cellular genes?
As with plant retrotransposons, many animal or yeast retroviral-like elements contain motifs similar to regulatory sequences of cellular genes, and several hypotheses have been advanced to explain this: • Ancestral captures of cellular sequences by mechanisms
simi-lar to retroviral transduction.
• Retrotransposon-mediated transfers of regulatory sequences to cellular genes; data have accumulated on the presence of trans-posable elements in the regulatory regions of genes25and on their involvement in changes in gene regulation, and it has been pro-posed that new regulatory features could be acquired by cellular genes after nearby transposable element insertions and subse-quent disappearance of the insertion through random drift26. • Appearance of retrotransposon regulatory sequences by
con-vergent evolution; for instance, the high U3 plasticity of the
Tnt1 family would by itself explain the appearance of
stress-related regulatory sequences in the Tnt1A subfamily.
None of the above hypotheses is completely satisfactory: the first implies that evolutionary patterns should be different between regulatory and coding sequences, which is not the case for Tnt1 (Ref. 22), and the presence in Tnt1A of several regulatory motifs found associated with a diversity of defence genes in many plant species appears incompatible with the two others. However, the possibility that these and other mechanisms might have combined with those of their host during retrotransposon co-evolution, cre-ating the present situation, cannot be excluded. Characterization of LTR evolutionary features in other retrotransposon families would provide further information on the origin and fate of retrotranspo-son regulatory regions.
The putative biological impact of retrotransposon activation by stress
Transposable elements are a major source of genetic variation that ranges from gross chromosomal alterations up to very fine tuning of the expression of cellular genes2. This, together with the obser-vation that transposons are activated by stress and environmental changes, led to the hypothesis that transposable elements are involved in host adaptation to environmental changes27,28
. In par-ticular, through modifications of gene regulation, transposable el-ements have been proposed as major factors in macroevolution26
. However, no clear example of an important biological role for such a modification in a natural population has yet been provided, and the question as to whether transposable elements are selfish para-sitic sequences or pacemakers of evolution is still controversial2
. In the light of this debate, the maintenance of retrotransposon stress activation might be purely fortuitous, and could have been driven by random choice within highly variable element popu-lations. However, the transcriptional features and regulatory se-quences of elements such as Tnt1A or Tto1 appear too specific to endorse this hypothesis in full. The situation is perhaps best sum-marized by Freeling29:
‘For those of us who still trust our senses, the design and be-havior of every living thing evidences an adaptedness born of exploitation. I cannot imagine a living system not exploiting transposons for something once they existed.’
Whatever the origin of Tnt1 and Tto1 regulatory sequences, they might have been maintained because they confer a selective ad-vantage, either to the host plant, such as through stress adaptation, or to the element itself. In particular, stress activation might allow elements to move only in rare situations, without major effects on host viability, a prerequisite for their own survival. In addition, the activation of retrotransposons during pathogen attack could increase the possibility of horizontal transmission between plants
and pathogens, allowing the element to colonize new hosts. The true answer might lie between these extremes, and different elements could have been maintained for very different reasons. Further re-search, in particular a demonstration that stress-induced transposi-tion events can be transmitted to the progeny in natural situatransposi-tions, will be necessary. One argument against any positive impact of stress-activated retrotransposons on evolution derives from the observation that the stress-activated Tnt1A subfamily is not the predominant Tnt1 subfamily in all Nicotiana species22
: the Tnt1B and Tnt1C subfamilies are predominant in the related species
Nicotiana plumbaginifolia, for instance. However, although Tnt1B
and Tnt1C are apparently not expressed in tobacco protoplasts, members of the Tnt1B subfamily were isolated after their trans-position in protoplast-derived cell cultures of N. plumbaginifolia (C. Meyer, unpublished). This suggests that other Tnt1 subfami-lies might also be activated by stress, through pathways that might differ and could be more precisely adapted to the particular spe-cies in which they predominate. All Tnt1 subfamilies might thus have been maintained as a consequence of their similar overall biological impact, each subfamily being preferentially amplified in a different Nicotiana genome to enable a better adaptation to its host and to its particular environmental history.
Conclusions and future prospects
A variety of retrotransposons has now been characterized in higher plants, and a clearer picture of their genomic organization, as well as of their evolution, is emerging. However, the control of their transposition cycle, from transcription up to the integration of the daughter copy, is not well understood. Further studies, in par-ticular characterization of additional active plant retrotransposons, will be necessary in order better to understand their impact on plant genomes. The activity of class II elements has been easily traced in plants partly because of their conservative mechanism of transposition, which results in frequent somatic instabilities. As retrotransposon insertions are stable, many plant retrotransposons were obtained by chance as inactivated insertions. A fruitful strat-egy based on the high conservation of some retrotransposon coding domains, originally developed in tobacco8, has allowed the iso-lation of retrotransposon sequences from their transcription prod-ucts, thus increasing the probability that they represent sequences from active copies (Box 1). Several studies have now been started in different species to isolate retrotransposon sequences expressed in specific stress conditions. This strategy could be applied to any plant species and will undoubtedly allow the characterization of new active elements, as well as provide important information on their conditions of expression and mobility.
Many years have passed since Barbara McClintock’s seminal analysis of variegated maize kernels and her description of regu-lated mobile DNA sequences. Transposon research has largely been fuelled by the promise of new applications, such as gene-tagging and the creation of transformation vectors, but the natural behaviour of these elements remains an intrinsically exciting research theme. It has implications for a better understanding of various areas: the control of gene expression; genome evolution and speciation; the idea of the environment influencing genome structure; and the possibility that the activity of plant retrotrans-posons is directly linked to defence responses.
Acknowledgements
We are very grateful to Z. Avramova, T. Brown, H. Hirochika, C. Kinlaw, A. Kumar, H. Laten, H. Lucas, C. Meyer and S. Wessler for preprints and unpublished information, and to H. Lucas for critically reviewing the manuscript.
References
01 Bennetzen, J.L. (1996) The contributions of retroelements to plant genome
organization, function and evolution, Trends Microbiol. 4, 347–353
02 Kunze, R., Saedler, H. and Lönnig, W-E. (1997) Plant transposable elements,
Adv. Bot. Res. 27, 332–470
03 Lindauer, A. et al. (1993) Reverse transcriptase families and a copia-like
retrotransposon, Osser, in the green alga Volvox carteri, FEBS Lett. 319, 261–266
04 Lucas, H. et al. (1995) RNA-mediated transposition of the tobacco
retrotransposon Tnt1 in Arabidopsis thaliana, EMBO J. 14, 2364–2373
05 Hirochika, H. et al. (1996) Autonomous transposition of the tobacco
retrotransposon Tto1 in rice, Plant Cell 8, 725–734
06 Pouteau, S. et al. (1991) Specific expression of the tobacco Tnt1
retrotransposon in protoplasts, EMBO J. 10, 1911–1918
07 Hirochika, H. et al. (1996) Retrotransposons of rice involved in mutations
induced by tissue culture, Proc. Natl. Acad. Sci. U. S. A. 93, 7783–7788
08 Hirochika, H. (1993) Activation of tobacco retrotransposons during tissue
culture, EMBO J. 12, 2521–2528
09 Turcich, M.P. et al. (1996) PREM-2, a copia-type retroelement in maize
is expressed preferentially in early microspores, Sex. Plant Reprod. 9, 65–74
10 Avramova, Z. et al. (1995) Matrix attachment regions and transcribed
sequences within a long chromosomal continuum containing maize Adh1,
Plant Cell 7, 1667–1680
11 SanMiguel, P. et al. (1996) Nested retrotransposons in the intergenic regions
of the maize genome, Science 274, 765–768
12 Suoniemi, A., Narvanto, A. and Schulman, A.H. (1996) The BARE-1
retrotransposon is transcribed in barley from an LTR promoter active in transient assays, Plant Mol. Biol. 31, 295–306
13 Grandbastien, M-A., Spielmann, A. and Caboche, C. (1989) Tnt1, a mobile
retroviral-like transposable element of tobacco isolated by plant cell genetics,
Nature 337, 376–380
14 Deragon, J-M. et al. (1996) A transcriptional analysis of the S1Bn(Brassica
napus) family of SINE retroposons, Plant Mol. Biol. 32, 869–878
15 Pearce, S.R., Kumar, A. and Flavell, A.J. (1996) Activation of the Ty1-copia
group retrotransposons of potato (Solanum tuberosum) during protoplast isolation, Plant Cell Rep. 15, 949–953
16 Meyer, Y. et al. (1993) Gene expression in mesophyll protoplasts, in
Morphogenesis in Plants (Roubelakis-Angelakis, K.A. and Tran Thanh Van,
K., eds), pp. 221–236, Plenum Press
17 Durr, A. et al. (1993) Why are quiescent mesophyll protoplasts from
Nicotiana sylvestris able to re-enter into the cell cycle and re-initiate a mitotic
activity? Biochimie 75, 539–545
18 Grandbastien, M-A. et al. (1997) The expression of the tobacco Tnt1
retrotransposon is linked to the plant defense responses, Genetica 100, 241–252
19 Hirochika, H. (1995) Regulation of plant retrotransposons and their use for
genome analysis (Gamma Field Symposia No. 34), Institute of Radiation Breeding, NIAR, MAFF
20 Takeda, S. et al. (1998) Transcriptional activation of the tobacco
retrotransposon Tto1 by wounding and methyljasmonate, Plant Mol. Biol. 36, 365–376
21 Johns, M.A., Mottinger, J. and Freeling, M. (1985) A low copy number,
copia-like transposon in maize, EMBO J. 4, 1093–1102
22 Casacuberta, J.M. et al. (1997) Quasispecies in retrotransposons: a role for
sequence variability in Tnt1 evolution, Genetica 100, 109–117
23 Mhiri, C. et al. (1997) The promoter of the tobacco Tnt1 retrotransposon is
induced by wounding and by abiotic stress, Plant Mol. Biol. 33, 257–266
24 Hirochika, H. (1997) Retrotransposons of rice: their regulation and use for
genome analysis, Plant Mol. Biol. 35, 231–240
25 White, S.E., Habera, L.F. and Wessler, S.R. (1994) Retrotransposons in the
flanking regions of normal plant genes: a role for copia-like elements in the evolution of gene structure and expression, Proc. Natl. Acad. Sci. U. S. A. 91, 11792–11796
26 McDonald, J.F. (1990) Macroevolution and retroviral elements, BioScience
40, 183–191
27 McClintock, B. (1984) The significance of responses of the genome to
challenge, Science 226, 792–801
28 Wessler, S. (1996) Plant retrotransposons: turned on by stress, Curr. Biol. 6,
959–961
29 Freeling, M. (1984) Plant transposable elements and insertion sequences,
Annu. Rev. Plant Physiol. 35, 277–298
30 Capy, P. (1998) Classification of transposable elements, in Dynamics and
Evolution of Transposable Elements (Capy, P. et al., eds), pp. 37–52, Landes
Bioscience
31 Finnegan, D.J. (1997) How non-LTR retrotransposons do it, Curr. Biol. 7,
R245–R248
32 Moore, G. et al. (1991) A family of retrotransposons and associated genomic
variation in wheat, Genomics 10, 461–468
33 Pearce, S.R. et al. (1997) Characterisation and genomic organization of
Ty1-copia group retrotransposons in rye (Secale cereale), Genome 40, 617–625
34 Varagona, M.J., Purugganan, M. and Wessler, S.R. (1992) Alternative splicing
induced by insertion of retrotransposons into the maize waxy gene, Plant Cell 4, 811–820
35 Bi, Y-A. and Laten, H.M. (1996) Sequence analysis of a cDNA containing the
gag and prot regions of the soybean retrovirus-like element, SIRE-1, Plant Mol. Biol. 30, 1315–1319
36 Hu, W., Das, O.P. and Messing, J. (1995) Zeon-1, a member of a new maize
retrotransposon family, Mol. Gen. Genet. 248, 471–480
37 Purugganan, M.D. and Wessler, S.R. (1994) Molecular evolution of magellan,
a maize Ty3/gypsy-like retrotransposon, Proc. Natl. Acad. Sci. U. S. A. 91, 11674–11678
38 Su, P-Y. and Brown, T.A. (1997) Ty3/gypsy-like retrotransposon sequences in
tomato, Plasmid 38, 148–157
39 Turcich, M.P. and Mascarenhas, J.P. (1994) PREM-1, a putative maize
retroelement has LTR (long terminal repeat) sequences that are preferentially transcribed in pollen, Sex. Plant Reprod. 7, 2–11
40 Day, A. and Rochaix, J-D. (1991) Structure and inheritance of sense and
anti-sense transcripts from a transposon in the green alga Chlamydomonas
reinhardtii, J. Mol. Biol. 218, 273–291
41 Yoshioka, Y. et al. (1993) Molecular characterization of a short interspersed
repetitive element from tobacco that exhibits sequence homology to specific tRNAs, Proc. Natl. Acad. Sci. U. S. A. 90, 6562–6566
Marie-Angèle Grandbastien is at the Laboratoire de Biologie Cellulaire, Institut National de la Recherche Agronomique (INRA), Centre de Versailles, 78026 Versailles cedex, France