158
Moehario and Soemanto Med J IndonesStudy of
genetic
diversity
of Salmonella typhi using pulsed-field
gel
electrophoresis
Lucky Hartati Moehario,
RetnoKadarsih
SoemantoAbstrak
Telah dilakukan analisis kekerabatan genetik isolat Salmonella typhi yang
di
isolasi dari pasien demam tifuid sporadik yang dirawatinap
di
RS Persahabatan, Jakarta, pada semesterI
tahun
1998.Dari 25
isolat yang dicerna denganXbal
ditemukan pola elektroforesis medanlistrik
berpulsasi (PFGE)yang
bervariasidan
diidentifikasi18
tipe
PFGE.
Analisissidik
gerombol menunjukkan bahwa kesemua isolat S. typhi berasaldari
dtn
grup utama, dan pada tingkat kesamaan> 84Vo
ditemukan 7 sidikgerombol. Penelitian
ini
menunjukan adanya keragaman genetik yang cukup tinggi pada berbagai isoLat S. typhi, dan tampaknya berbagai isolat fui berasal usul bukan dari klon tunggal. (Med J Indones 2001;I0:
158-63)Abstract
DNA genomes
of
Salmonella typhi (S.typhi), which were isolatedfrom
sporadic typhoid fever cases who were hospitalized in Persahabatan Hospital, Jakarta, during thel"
semesterof
1998, were examinedfor
their genetic diversities. Pulsed-field gel electrophoresis (PFGE) of genomic DNA digested with Xbal was performedfor
25 isolntes. Electrophoresis patterns of most isolates varied and 1 8 PFGE types were identified. Cluster analysis showed that aII isolates originated from two main groups: and at > 84Volevel of similarity, 7 clusters were
found.
Thus, the results showed that genetic diversity of S. typhi was considerably high, and that S.typhi isolated from sporadic typhoid fever cases were derived from muhiple clones. (Med J Indones 2001 ;
I0:
158-63) Keywords: typhoidfever, sporadic cases, Xbal, cluster analysisIntensive
studies
of
genetic diversity and
molecular
epidemiology of
S.typhi, theetiologic
agentof typhoid
fever,
have beencarried out
sinceearly
nineties.Quite
a
range
of
S.typhi isolates
for
various
geographicregions
representing sporadic cases, outbreaks,
andenvironmental
isolates
have been
analysed
usingpulse-field gel
electrophoresis (PFGE). Those
studiesshowed
that there were significant diversity
amongS.typhi isolates,
in which
isolates originated from
outbreak
caseswere more
homogenous compared
tosporadic
isolates. Also,
it
is
suggestedthat
there was a movementof
certain
strainswithin
Southeast Asia.r-sIn
Indonesia,
the
bacteria
causes
a
disease
with
abroader spectrum
of
serious complications,
amongothers,
typhoid hepatitis and typhoid
pancreatitis.6Further,
mortality
is
frequently seen
in
Indonesia,while in
other part
of
Asia, the
disease manifestationsDepartment of Microbiology, Faculty of Medicine, University of Indonesia, Jakarta, Indonesia
are milder. Previous study
by
Thong
et
aI,
19961reported that
certain PFGE
typeswere
associatedwith
the
ability
to cause afatal
disease.In
the
present
study, we
applied
PFGE
technique
toanalyse large fragments
of
chromosomal
DNA
to enable usto
determine thegenetic relationship
amongS.typhi isolates.
The result
obtained can
be
usedfor
epidemiology
purposes,since
it
can contribute
in
theconstruction
of
highly
accurateS. typhi
genetic
mapfor
thewhole region
in
Indonesia.METHODS
All
procedures
below were
performed
in
the Laboratoryof
the Department ofMicrobiology,
Faculty
of
Medicine,
University
of
Indonesia,
during
1999-2000, exceptfor
the electrophoresisusing
CHEF DR
III
electrophoresis
unit,
which
were conducted
in
Microbes and Genetic Engineering
Division,
R&D
Centre
for
Biotechnology, Indonesia
Institute
of
VoI 10, No 3, July
-
September 2001Bacteria
Twenty
five
S.typhi
isolates
were randomly
chosenfrom
the collectionsof Dr. RHH
Nelwan,
Departmentof
Internal Medicine, Faculty
of
Medicine,
University
of
Indonesia,
Jakarta.
The
isolates were
originated
from
sporadic
cases
of
typhoid fever,
which
werehospitalized
in
the
first
semester
of
1998,
in
Persahabatan
Hospital, Jakarta.
All
isolates
weretested
for
antibiotic susceptibility
by
standard disk
diffusion
procedures.
No
resistancewas observed to
amoxicillin
(AML),
amoxicillin
clavunamic
acid(AMC),
chloramphenicol
(C),
trimethoprim
sulpha-methoxazole
(SXT), ciproxin (CIP), ofloxacin
(OFP),lefofloxacin (LFX),
streptomycin (S), tetracyclin (T),
cefdinir (CFN), cefprozil (CPR),
ceftriaxone
(CRO), cefèpime(FEP), or amikacin
(AN).
PFGE
and
genomeanalysis
Chromosomal
DNA
was
prepared
as
described by
Suwanto and
Kaplan,
1989,8 andThong
et
al,
1994.sIn brief,
overnight cultures
werecentrituged
at 40009for
5
min. at
4oC and washed
twice
in
I
ml
buffer
containing
l0
mM Tris HCI pH 7.5 and
1
M
NaCl."Pellets were
resuspended
in
the
same
buffer
andmixed
with
lVolow melting
agarosein
1xTE buffer
(10 mM Tris HCI pH
8.0
and
I
mM EDTA pH
8.0).The
agarose
blocks were then
subjected
to
lysing
solution
containing
I
mg/ml
lysozyme and incubated
at
37"C
overnight.
The fbllowing
day,
the
lysing
solution were
replaced
with
solution containing
100pg/ml
proteinase
K,
0.5
M
EDTA pH
8.0
and
lOVo sarcosyltbllowed by
incubation
at55'C for
48
hours.The
XbaI
restriction enzyme (Promega,
Madison,
WI,
USA)
was used, and digestion was performed according to the manut'acturer.The digested chromosomal
DNA
were then
electrophoresed
using
CHEF
DR
III
electrophoresis
unit
(Biorad
Laboratories,
Hercules,CA, USA)
at l4"C
with
a
120"
switch
angle, and
arunning
time
of
20
hours
with
a
linear ramp of
switching
time
fiom
5-50
sec,
at
6
Volt/cm'.
Rhodobacter sphaeroides 2.4.1 digested with Asels was
generously
provided
by DR. Antonius
Suwantofrom
Inter
University
Centerfor
Biotechnology, Institute
of
Agriculture Bogor, Bogor, Indonesia, and used
asmolecular marker. Gels were
stained
with
ethidium
bromide, destained and photographed underUV
light.
S.typhi -
PFGE
159Interpretation
of
PFGE gel patterns was
carried out
asdescribed
by
Bannerman
et
al,
l9g5,e and Zadoks
etal,
2000,10and
assigned
for
arbitrary pattern
rypes,and
compared
by
calculating
Dice
similarity
coefficient
(F, proportion
of
share fragments betweentwo isolates).
F valueof
1.0 indicatesidentical
pattern,and
F
value
of
0
suggests complete dissimilarity.rr'r2Electrophoretic
patternswere analysed
for
relatednessusing
a computer program Numerical Taxonomy andMultivariate
Analysis
(NTSYS-pc)
version
1.80.r3Dendograms
were
constructed
by
using
Dice
coefficient,
and
clustering
by
using
unweighted pair
group arithmetic
means method(UPGMA).
RESULTS
A
total
of
25 human isolates
of
S.typhi were
analysedfollowing
digestion
with
XbaI
(5'-TCTAGA-3')
restriction
endonuclease
and
electrophoresis
usingPFGE.
Digested chromosomal
DNAs
produced
l0
to20 fragments
with
molecular
weight
ranging
from
30kb
to
650
kb
and showed various
PFGE
patterns
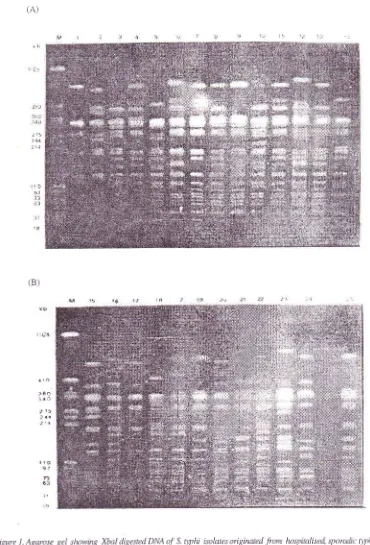
as shown inFig.
lA
andlB.
Close examination
to
thesebands showed
that
DNA
fragments
with
molecular
weight
between
31 kb
to
97 kb,
and around 340 kb
were relatively
conserved
in
most
of
the
isolates.However,
bands between97
kb to
340
kb,
and above340
kb
were more
various.
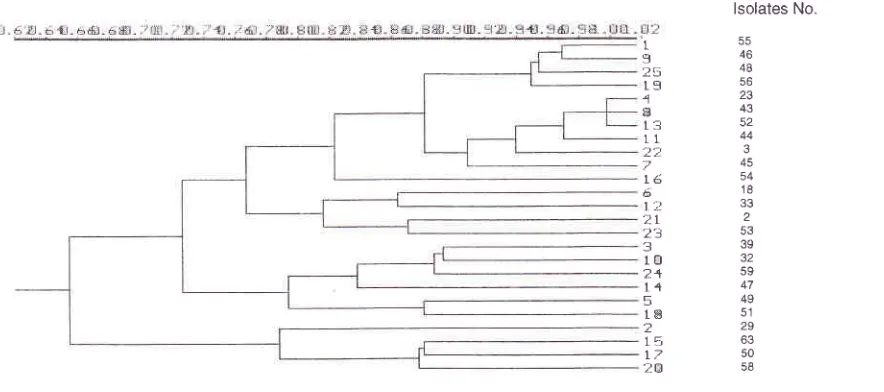
Analysis
of
geneticrelatedness
among S.typhi isolates showed
a
quite
significant
diversity among them as
represented by
their
F
values
(Fig.
2A),
and
the
presence
of
18different
PFGE
types
(pulsotypes) designated
Xl
toXl8,
and4
subtypes asfollows: X1.1, X1.2,
X2J
andX2.2
(see
Table
l).
Despite
the
divergence
of
F
values, certain pulsotypes were eitheridentical
or verysimilar,
in
that they were different
in
only
I
or
2bands. There were 5 S.typhi isolates
(isolate
no.3,23,
43,44
and 52)with
F value between0.923-1.000,
andtherefore
they were grouped
together
in
PFGE cluster
I[.
Three
of
these isolates,i.e. isolate
no.23,43
and52
were
identical
with
F
value
1.000.
Moreover,
4other isolates (isolate no.
46, 48, 55
and
56)
showedhigh
degreeof similarity with
F between0.941-0.971,
160
Moehario and Soemanto(A)
? 4.1
:rJ
(B)
a ro
]êo
: r.Q
[image:3.612.98.468.89.634.2]I rt lct
Figure 1 . Agarase gel showing Xbal digested DNA of S. tythi isolates oigirnted from hnspinlised sporadic 4,photlfever paients.
(A)
Lanesl-l4representrestricfbnenqnædi7estedDNApattemsofisolnte number: 55,29,39,23,49, 18,45,43,16,32,44,33,52, and 47.(B)lntes15-25 represent isolate runber: 63,54,50,51,45,56,58,2,3,53,59, anà 48.lnteM:PFGE
marker (Rhodo-bacter sphaeroides 2.4.1. digested with Asel) in L<b pairs.Vol 10. No 3, Jug
-
September 200)(A).
S.q'phi - PFGE 161
5-s I 000
29
0.5r4 t000_r9 0667070.1 t 000
23
09r405000757 I 00019
06290'7220865 0667 I 00018 07650-571 0778080006E6 l 000
4-5 0 882 0-5140 72209140 6860 76-5 I 000
,{l
0 914 0 500 0 757 1.000 0 667 0 800 0 914 I 000,16 097r 05s6070308890667080008s70889 r 000
32
012206490895 0157 0151 0 833 0667 075707-s7 I 00044
08890486 0 717 0e7t 0 6490 8_13 09,+40 973 0 865 0 717 r 00013
07s70.526082t 0842078908650757084207890 82t 0821 I 00052
09r40s000757 r.0000667080009r41.0000889 0,1510973 0842 r 00011
07220 649089s075708r r 07?80833 0't57 0757 078907890769075? i.00063
0 686 0 778 0 1 57 0 661 0 722 0 1 43 0 629 0 661 0 122 0 1 51 0 649 0 684 0 667 0 649 I 00054
0788 0.s29 0 8000882 01060121 0848 0 8820765 0 7.+3 0 857 0778 0 882074-r 0706 I 00050
0667 0765 0 741 0647 0 -s88 0121 06060 647 0 7060 14306290661 0641 0686 0 882 0625 I 000-5r 066705880857070608820727 07210706 07060743068608-31070608s7064?0688062s I 000
56
094r 0571 0667085706E6076s082408570971 0722081-r07570857072206860127A667 0667 l 000s8
06060 8240 80006.17 01060121 0606 0 647 0 647 0 80006290 661 0611 06860 8820 688 0 87-s 062-5 0 606 l 00C2
080005560703 077806t I 08s706860778 0813 0151 01s1 08420778064907780 7060765064708000706 r 000-r
0865052607690947 0684081 I 086-s 0947 0895 076909210E5009,+707690684083306670722086506670789 r 00053 0 68406t5 06500718 06t5 0789063207r807r80 7000 700078007J8 060001r8 064906490 59s 0 6840703 08720780 l 000
59 0765 062e 0 889 08000741 0 765 0765 0 800074.r 0 889 0778 0 7-s7 0 8000 8-]] 0741 0 8480727 0727 07060788 06860757 0632 r.000
48 0941 0571O'12209t4 0686082,+0882091409710778088908110914077806860.7880661 0121 0941 0667080009190737076-5 1000
Figure 2. (A). Matrix of F values of 25 S. D1rhi isolates. The
far
left ordinate reyesents the isolate's number.(B).
lsolates No.
46
4B
56
43 52 44
3
45 54
18
2 53 39
59
47 49
51
29
50 58
L I
?5
ts
{
B
t5
L1 't'
tét Ê;
t;i
2L10 2+
l4
5l-E
[image:4.612.37.510.80.427.2] [image:4.612.55.497.478.674.2]t5
I7
2EVoL 10, No 3, Juty
-
September 2001REFERENCES
l.
Wain
J, Hien TT, Connerton P,Ali
T, Parry CM, ChinhNTT, et
al.
Moleculartyping
of
multiple antibioticresistant SalmonelLa enterica serovar typhi from Vietnam:
Application to acute and relapse cases of typhoid fever. J
Clin Microbiol 1999:3'l: 2466-72.
2.
ThongKL,
Nair S,
SubramaniamG,
Puthucheary S,Yassin
R,
CheongYM, et al.
Genetic dynamics andmolecular epidemiology
of
Salmonellatyphi.
Med
I
Indones, 1998; 7 Suppl
l:
147-50.3.
Koay
AS,
JegathesanM,
RohaniMY,
Cheong YM.Pulsed-field gel electrophoresis as an epidemiologic tool
in
the
investigationof
laboratory acquired Salmonellaryphi
infection
Southeast Asia J Trop Med Public Health1997:28:82-4.
4.
ThongKL,
Puthucheary S, YassinRM,
Sudarmono P,Padmidewi M, Soewandojo E, et al. Analysis of Salmonella
ryphi isolates
from
Southeast Asiaby
pulsed-field gelelectrophoresis. J Clin Microbiol 1995; 33: 1938-41.
5.
Thong KL, Cheong YM, Puthucheary S, Koh CL, Pang T.Epidemiologic analysis
of
sporadic SalmonelLa typhiisolates and those
from
outbreaksby
pulsed-field gelelectrophoresis. J Clin Microbiol 1994; 32: I 135-41 .
6.
Nelwan RHH. Changing patternof
typhoid fevercom-plications in Indonesia. Med J Indon, 1998; 7 Suppl
1:
105.7.
ThongKL,
PasseyM,
Clegg A, Combs BG, Yassin RM,Pang T. Molecular analysis of isolates of Salmonella ryphi
obtained
from
patientswith
fatal and nonfatal typhoidfever. J Clin
Microbiol
1996; 34:1029-33.8.
SuwantoA,
Kaplan S. Physical and genetic mapping oflhe
Rhodobacter sphaeroides 2.4.7. genome: Genomesize, fragment identification, and gene localization. J Bact 1989; 171(11):5840-9.
9.
BannermanTL,
Hancock GA, Tenover FC,Miller
JM.Pulsed-field
gel
electrophoresisas
a
replacement forbacteriophage typing
of
Staphylococcus aureus.J
ClinMicrobiol 1995; 33: 551-5.
10.
ZadoksR,
LeeuwenW,
BarkemaH,
Sampimon O,Verbrugh H, Schukken YH, et al. Application of
pulsed-field
gel
electrophoresis and binarytyping
as tools inS.typhi -
PFGE
163veterinary clinical microbiology and molecular epidemiology
analysis
of
bovineand
human StaphyLococcus aureusisolates. J Clin Microbiol 2000; 38: 1931-9.
Romling
U,
GrothuesD,
Heuer T, Tummler B. Physicalgenome analysis
of
bacteria. Electrophoresis 1992; 13:626-31.
El-Adharni W, Roberts
L,
Vickery A, Inglis B, Gibbs A,Stewart
P.
Epidemiological analysisof a
methicillin-resistant Staphylococcus aureus outbreak using restriction
length polymorphisms of genomic
DNA.
J Gen Microbiol1991; 137:2713-20.
Rohlf
FJ.
NTSYS-pc: Numerical Taxonomy
andMultivariate Analysis System Version 1.80. New York:
Exeter Software; 1993.
Prevost G, Jaulhac B, Piemont Y. DNA fingerprinting by
pulsed-freld
gel
electrophoresisis
more effective thanribotyping
in
distinguishing among methicillin-resistantStaphylococcus aureus isolates. J
Clin
Microbiol 1992;39:967-73.
Lefevre
JC,
FauconG,
SicardAM,
GascM.
DNA fingerprintingof
Streptococcus pneumoniae strains bypulsed-field gel electrophoresis. J
Clin
Microbiol 1993;3l:2724-8.
Anderson DJ. Kuhns JS, Vasil
ML,
Gerding DN, JanoffEN.
DNA
fingerprintingby
pulsed-fieldgel
electro-phoresis
and
ribotyping
to
distinguish Pseudomonascepacia isolates
from
a
nosocomial outbreak.J
ClinMicrobiol
l99l;
29: 648-9.Hector JSR, Pang YJ, Mazurek GH, Zhang YS, Brown
BA,
WallaceRJ.
Large restriction fragment patterns ofgenomic My c o b acteri um fo rtuitum DNA as s train- speci fi c
markers and their use
in
epidemiologic investigationof
four
nosocomial outbreaks.J Clin Microbiol
1992; 30:I 250-5.
Chee
CS,
NoordinN,
IbrahimL.
Epidemiology andcontrol of typhoid fever in Malaysia. In: Pang T, Koh CL,
Puthucheary SD, editors. Typhoid fever: Strategies for the
90's. Singapore:
World
Scientific Publishing;1992.p.3-10.
Liu
SL,
SandersonKE.
Genomic cleavagemap
ofSalmonella typhi.
I
Bacteriol 1995; 17 7 : 5099- I 07.11
12
13
t4
15
l6
t7
l8