Journa[
of
Biotogy
Agriculture
and
Heatthcare
Vol
3,
No
13
(2013)
Table
of
Contents
Articles
A Pilot
Studyon
effectsof
vreination
onimmunity of broiler
chickensOjiezeh,
T.I,
Ophori,
E.A., Eghafona, N.
O.,Echeonwlt,
G. O.N,
l-4
Joannis,
T.M., Akele,
R.Y.Adverse
Drug
Reactionsamory
Critically
Ill
Patients atCairo
University Hospital:
Frequency and OutcomesHanan Ahmed
El
Sebaeq YousriaAbd El
SalamSeloma
5-13 Women FarmersOrganisatiors'
PerceivedEffect of Child Labour
Activities in Oyo West Local,Government Area of Oyo
State,Nigeria
Oyeyinka, R.
A., Ayansiru,5.O.,
Adekunmi,A.O., Arowolo.
O.O.
14-21Cerebrovascular Stroke
Recunence
amongCritically
Ill
Patients at aSelected
University Hospital in Egypt
Warda
YoussefMohamncd Morsy, Hanaa
Ali Elfelq,
Rasha
22-33
AlsayedAhmed
Dry Matter Accumulation in Maize
asInfluenced by Row
Arrangement,
Nitrogen
and PhosphorusLevels in Maize
(ZeaMaysL)l
Castor
(Ricinus Commumis
L)
Mixture
Arunah U.L,
E.B.Amans,M. Mahmud,
A. Ahmed, G.L.
Luka,A.
34-41S.Isah, B.A. Babaji, E.COdion
Effect Of Ginger Infusion OnChemotherapy
Induced NauseaAnd
Vomiting In
Breast Cancer PaiientsRahmi
Muthia,lledya
Wahyu,Dachriyanus
.
42-46
Effect of Soil Conservation lnyestment
onEfficiency of
CassavaProduction in
Oyo
Stateof Nigeria
Job
Olatunji
Oladeebo, AdepejuAderinola
Oyeleye,Mutiu
47-52
Oladapo Oladejo
Evaluation
of
aTool for
AssessingClinical
Competenceof Msc
NurseStudents
Margaret
Chege, PeterMwaniki,
TimothyAbuya
53-59Hope
Level
andLife
Satisfaction among Patientswith Colostomy
andtheir Family
caregiversNaglaa Fawzy Hanafy
Taha,Manal
MohamedMoustafa
60-72
Comparison
of lnformational
Needs amongNewly
Diagnosed BreastCancer
Women Undergoing
Different
Surgical TreatmentModalities
The Role
of
Ripe Musa sapientum(Plantain)
Peels Phosphorus andNitrogen from Aqueous Solution
Akpor
OB,Nwonuma
CO,Edewor-Kttponiyi
Genetic
Variation of
CoconutTall
(Cocosnucifera
Bali,lndonesia
Based onMicrosatellite
DNA-in
theRemoval
of
TI,
Amira OJ
L.,
Arecaceae)in
Eniek
Kriswiyanti,
I
Gede RaiMaya
Temaia,I
Made
Sudana,I
Gusti
Ngurah
Alit
SusantalYirya
combining
Ability
andlnheritance of Growth Traits in
RabbitsA.S. Actenaike, T.O. Osisanya, O-D- Ogunsola,
A'O' Asine'
M'
Wheto,
D.O. Ogunlakin,
A.S. Amusan,C.O'N' Ikeobi
Supplementation
with
GoatFollicular Fluid in
theIn
Vitro
Maturation
Medium toward
Cumulus Expansion andNucleus Transformation
Nolasco
Da Costa,
Sri Wahjuningsih,Nurul Isnaini
Reduced
Levels of
SomeIron
Parametersof Protein Energy
Malnourished Children in
Calabar,Nigeria
Naomi
Ernest, Patience Akpan,Emmanuel
UkoStudy
of Nursery
Businessin Harbin Region
Shah Saud, Chen Yaiun,
Shahfahad, Muhammad Abdullah,
MuhammadMaroof Ajmal, Arooi sadiq
The
lncidence of Anemia
and theImpact of Poor Glycemic Control
in
T ype-2
Diabetic
Patientswith
Renallnsuffi ciency
Babatunde
Ishola Adeiumo,
UchechuhuuDimkpa, Chinwe
Obiaruiu Ewenighi,
Tosan AmosErhabor, Uchunor G'A', Odia
5.1., Ukatu Esmond,
Oji
O-J., Besong E.E.Performance
of
Yankasa Rams FedAndropogon
gayanus (GambaGrass)
Hay
Supplementedwith
Faidherbiaalbida (Acacia)
PodsI.
Aiiii,
H.D. Nyako,
S.A. AshomIdentification
andcharacterization of Actinomycetes for Biological
Control of Bacterial
Scabof
Streptomyces scabiesIsolated from
PotatoHencl Abdulhmeed Hamedo,
Abeer
HarndyMakhlouf
Fluctuation of Daytime
Air
Humidity in
TheMangrove
Forest Edges TinnyKaunang,
Ch.S.Medellu
Hepatoprotective
Activity
Combination
BetweenMorinda
Citrifolia
Linn (Mengkudu) Extract And
Virgin
Coconut
Oil
(Vco)
Rutli
Alexander
Repi,Mokosuli
Yermia Semuel,Jantie Ngangi,
Harry Maurits
SumamPottwSexava
Nubila Behavioral Analysis In Every
ChangesOf Imitation
Sound
From Level Of
IntensitYBahrun
.,Jootie
Warouw, Sartje.J- Rondonuv'u,Mar
Tulung
8s-96
97-l0l
102-107
108-1 13
tt4-t20
12r-t29
130-136
137-t41
t42-153
154-159
r60-166
Joumal of Biology, Aglicultule and Hkalthcare
ISSN 2224-3208 (Paper) ISSN 222593X (Online) Vol.3, No. 13, 20 l3
!\tr!*-ij!!q.oE
t.r
Il
!$r
Genetic
Variation of
Coconut
Tall
(Cocos
nucfura
L.,
Arecaceae)
in Bali,
Indonesia
Based
on
Microsatellite
DNA.
EniekKriswiyanti*, I
Gede Rai Maya Temaja**, I Made Sudana**and I Gusti Ngurah
Alit
SusantaWirya**
t?cCra&rate
Student ofAgirrlturc
Faculty and Departmentof
Biologt
Mathc and Natural Science Faculty ,Ifbys
Uni'gliD.
Cows
Bukit Jimbaran, Kuta Bqli Indonesia**
Facultl, of Agriculture, UdoyanaUirasity
Canpus Sudirman, Denpasar, Bali IndonesiaCorn+odiog
author, Email : [email protected]Ifarud rc ffid
byUodor
General of Higher Education, Ministry of Education and Cultureof
n#
fua
fua dmrae
Sudent) and Udayana University, Bali IndonesiaAbcEett
Ihe
oomr
b ia:r
nles
for
economics,traditional
medicine andculture,
especiallyfor
Hindu's ccremmialpuprc L
Bdi
H
(Indonesia). Each coconutcultivar
has unique characteristics. The aimedof
this rcsearn
ree
n
*.r-rin
gEoetic variation of coconut tall (Cocos nucifera L., Arecaceae) in Bali, based onDNAmftx66aEflh-
SirFlsricrosatellite
markers were used to determine heterozygosity.The
reslle shomd
h El
dEO alleles
were detected by microsatellitewith
an averageof
13.33 alleles perlocus, there
strc
12.lklcs
mmicrosatellite
locusCnCirA3,
12 alleles on locusCnCiC3,
16 alleles on locus CNZ05,l4
afi3lcsc
C}{ZIB,
17 alleles onCNZ21,
and 9alleles
on microsatelliteprimer
pairs CNZSI. The msanvalucsof
gcdivErsity(tle)
and observed heterozygosity(Ho)
were 0.8835 and 0.5421, respectively.The highes
rcdg3rfty
q,i6 on bulantall
coconuts cultivar (0.816), the lowest heterozygosity was on bluluktallcocmrs(03+
Keywords:
rm;rpechracterist,
microsatellite, allele, heterozygosity, coconutl.Introdrcti.r
Cocmut
(Carr;
twifea
L-) is oneof
the primary sources of income for most Indonesian farmers. Coconutfruit
is processodiro
vairxls
prodrrctslike
vegetableoil,
raw rnaterialfor
food, industry, medicine, cosmetic andoleo
&mitals
Totat land
area plantationof
coconutis
about3.7 million
hectares,with
97o/oof
overallpro&rlin
ftom fumcrs
(Novarianto, et al.,2005). In Bali,
besideof
its economic value, partsof
coconut tree(leaveg
ftrig yong
flower)
also usedfor
ceremonial and medicinal purposes, which called"nyuh
madan"Morphologitztly,
some coconut plantsdiffer
in
some characters,which
is
very
unique and specificto
itsindiyi&lal
plam-Bced
on the unique characters, there were more than twenty unique coconuts identified thatdiftr
iodivienty-
Someof
them are ancak which has brancheson its trunk;
the beiulit
has plicated laminateaves,
fu
bingin 's root growfrom
nodes of stem, theBojog's fruit
husk is colored like the hairof
long tailedmookey- Differences
in fruit
color such as white, green,yellow
and red were used to distinguish between thefuloa
gadang" CadinC and surya coconuts respectively. Inflorenscentia spicata was characteristicof
bluluk,urhih
fu
udang and mulung were characterized by red mesocarpium. The Rangda coconut has petiole and theryor ofthe
stem were twisted. The sudamala was characterized by many kinds 0f unique characters, which someofthem are doubled spatha, and flat apex of male spikelet (Kriswiyanti,2012).
To study its genetic diversity, simple sequence repeats (SSR) microsatellite markers were used. This technique had been used successfully by many scientists
to
characterize the genetic diversityof
the coconut population(Perera et al. 2001; Manimekalai and Nagarajan 2006; Kumar et al., 201 I ). The microsatellite analysis
of
the 9coconut populations
with
8 primers revealed a totalof
37 alleles (Devakumar et.al.,2010). From the used 26 SSR markers, detecteda
totalof
188 alleleswith
anaverage of
7 .23 per locus(Liu,
et.al., 201l).
In
thisresearch, genetic variation
of
tall
coconut (Cocosnucifera
L.,
Arecaceae) was determined basedon DNA
microsatellite, using alleles size on each individual
of tall
coconut. The alleles data from each accession can be applied to predict natural breeding for unique coconut conservation purpose.Materials and Methods
Plant materiul used:
Leaf
samples were collectedfrom
Klungkung,Bangli,
Karangasem, Gianyar, Tabanan, Badung, Jembrana residence and Denpasar,Bali
province, Indonesia.Of
the 58 samples from twenty cultivarsJoumal of Biology. Agliculturc and llcatrhcare
ISSN 2224-3208 (Paper) ISSN 2225-09_iX 1C)nlincr
Vol.3, No.l3.2013
www.iiste.org
IfrUI
Ni
were
'tall coconut'
category(ntuh
ancak, biasa, bulttn. bingin, rangda, bluluk, sudamala, surya,udang,
kopas,be
1utir,bojig,
detection
of
microsatellite
polymorphismswere
analysed ai University, Bali Indonesia.gadang, goding, kebo, srogsogan; mui:lung,
mocqn, naga,
pudak
).
DNA
extraction andForensics
and
Primata Laboratory UduyuouDNA extraction: DNA
was extracted from fresh coconut young leaves using a CTAB based protocol modified from Doyle and Doyle (1987). The primer sequences and associated information are givenin
TableI
(pandin, et.aI.,2008).SSR analysis
Polymerase
chain Reactions
(PCR) assay and gel analysis :DNA
was amplifiedin l3
pL reactions containing 2 pL sample, 3'5pL
H2o, 6.5pL
Mastennix (Qiagen), andI
pL primer. ThepcR
programmed for 30 cyclesof
45 seconds each at94oc,
I
min
at the difrerent annealing temieratures standardizedfor
theindividual
sSRlocus, and extension
for
I
min 30 secondsat
72oC. The first cycle was preceded by a 5 min denatuption at 94oC and the last cycle ended with 5 min extension at ?2"c.
Reaction products were separate don
6yopolyacrylamide(denatured) and visualized
with
silver nikatestaining
(Tegelstorm, 1984).The alleles were scored based on the size of each PCR amplified fragment by electrophorisis all samplesin
a single gel.Allele
sizewere determined by semilog plotting of distance migration of amplicon on
racE
luutchinsoi,
z6or). oir"rsity
values based on allele frequencies were calculated for each microsatelite locus and coconut cultivar using Nei,s methods (19g7).!!le.l
Details of microsatelites mer used in the [image:5.612.63.517.317.610.2]From 58 extracted
DNA
samples, some sample were successfully amplified on each primer used. For examples locus CNZ09 on figurel,
C and D were not successfullyamplifiei.
Figure l
'
An
exampleof allelic
polymorphisme at microsatellite locuscNZ0g
in
12individuals
coconut tallfrom some village in
Bali
island . A.BeJulit
tall
Babung, B BeJulit
tall
Buruan,c
BeJutit
tall
ruwed
D. BeJulit
toll
ramblang,
E.Biosatall
Pejeng,F.Bingin
tall
lelektngkang,G.Bingin
tall
Babwg,
H.
Biasq tall
Babung, r.Ancak
tall
Plkat, J .strrya tctll Jelekungkang, K.Gadangiail
Babung, L.Rangdatall
BabungGenetic
Variation
(Heterozigosity)Total number
of
investigated alleles, number alleles, heterozygosity, and variation alleles lengthof
twentytall
coconut cultivars for each SSR locus were showed in
Tablel. the
clmbination of six sSR locigenerated a totalof
80 alleles,with a
meanof
13-33 alleleper
locus and rangingfrom
nine(cNzsl)
to
seventeen alleles(cNZ2l).
Gene diversity range
from
0.8561for
theloci
cNZ5l
to
0.92040.8835. The loci
CnCiC3
and CNZ05 presented the lowest (0.45) for observed heterozygosity, with a mean value of 0.542 I .for
theloci CNZ2I, with
an overall meanand the highest value (0.7619) respectively,
Forward
(5'-3')
Riverse(5'
-
3')
CTTATCC AUMTCGTCACAGAG AGGAGAAGCCAGG AA
AGATTT
ATCTACCAGTGTGGTCCTCTC ACCAGGA A AA AGAGCGGAGAA
ATGTTTTAGCTTCACCATGAA
TCAAGTTCAAGAAGAC CTTTGCTTTAGGG AJqu{AAGGACTGAG ATCCATGAGCTGAGCTTGAAC
AATCTAAATCTACGfuMGCA
AG AJqu{GCTGAGAGGGAGGATT GTGGGGC ATGAJqJMGTAu{C
Jounral of Biology, Agriculture and Healthcare ISSN 2224-3208 (Paper) ISSN 2225-091X (Online) Vol.3. No.l3,2013
rvrvw.iiste.org
d{ilt
il$r
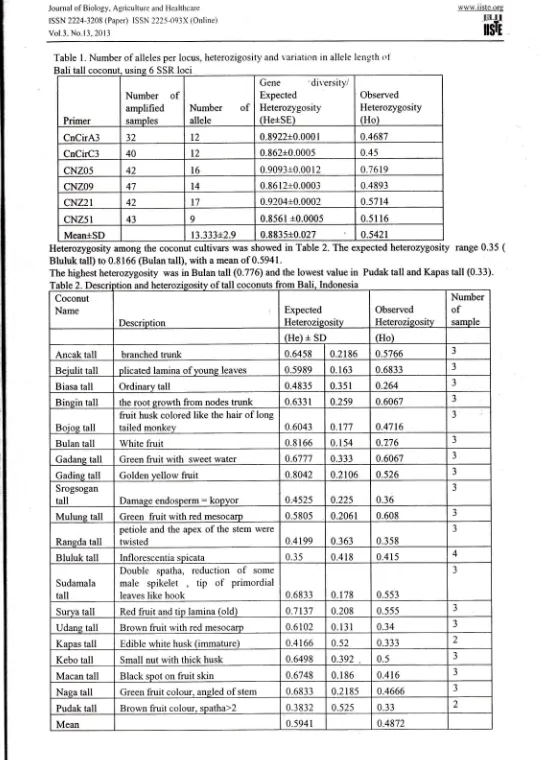
Table
l.
Number of alleles per locus, heterozigosity and variation in allele length olusl
Primer
Number of
amplified
samnles
Number
of
allele
Gene
-diversity/Expected Heterozygosity (He+SE) Observed Heterozygosity (Ho)
CnCirA3 32
t2
0.8922+0.000 t 0.4687CnCirC3 40 t2 0.862+0.0005 0.45
CNZO5 42
l6
0.9093*0.0012 0.76t9
CNZO9 47
t4
0.861210.0003 0.4893CNZ2I
42 t1 0.9204+0.0002 0.5714CNZ5I
43 9 0.8561+0.0005 0.5116Mean+SD 13.333*2.9 0.8835+0.027 0.5421
Bali tall ins 6 SSR loci
Heterozygosity among the coconut cultivars was showed
in
Table 2. The expected heterozygosity range 0.35 (Bluluk tall) to 0.8 166 (Bulan tall), with a mean of 0.5941 .
The highest heterozygosity was in Bulan tall (0.77 6\ and the lowest value
in
Pudak tall and Kapas tall (0.33).Table2. Description and heterozigosity of tall coconuts from Bali, lndonesia Coconut Name Description Expected Heterozigositv Observed Heterozisositv Number
of
samole(He)
+
SD (Ho)Ancak tall branched trunk 0.6458 0.2186 0.5766 3
Beiulit tall olicated lamina of vouns leaves 0.s989 0.1 63 0.6833 3
Biasa tall Ordinarv tall 0.4835 0.3s1 0.264 J
Binsin tall the root growth from nodes trunk 0.6331 0.259 0.6067 3
Boios tall
fruit husk colored like the hair
of
longtailed monkey 0.5043
0.t77
0.47t6
J
Bulan tall White fruit 0.8166 0.1 54 0.776 3
Gadans tall Green fruit
with
sweet water 0.6777 0.333 0.6067 3Gadine tall Golden yellow fruit 0.8042 0.2106 0.526 3
Srogsogan
tall Damase endosoerm
:
kopvor 0.4525 0.225 0.36J
Muluns tall Green fruit with red mesocarp 0.5805 0.2061 0.608 3 Raneda tall
petiole and the apex
of
the stem weretwisted 0.4199 0.363 0.358
3
Bluluk tall Inflorescentia spicata 0.3s 0.418 0.415 4
Sudamala
tall
Double
spatha,reduction
of
somemale
spikelet
, tip of
primordialleaves like hook 0.6833 0.178 0.553
3
Surva tall Red fruit and
tip
lamina (old) 0.7t37 0.208 0.555 JUdane tall Brown fruit with red mesocarp 0.6102 0.131 0.34 J
Kaoas tall Edible white husk (immature) 0.4166 0.52 0.333 2
Kebo tall Small nut with thick husk 0.6498
0.392
. 0.5 3Macan tall Black spot on fruit skin 0.6748 0.1 86 0.416 3
Naea tall Green fruit colour. angled of stem 0.6833 0.218s 0.4666 J
Pudak tall Brown fruit colour, spatha>2 0.3832 0.525 0.33 2
Mean 0.5941 0.4872
[image:6.612.31.570.30.790.2]Journal of Biologv. Aericulture and Htalthe :rrc
ISSN 222.1-3208 (Paper) ISS\
llli-ttel\
rt)rrirnc Vol.3. No l-1. 20ll,,vww.iiste.ort
Hrr
ffi
Discussion
The alleles size
of
several locus fromtall
coconut that explored during this research differed to that found in other studies of coconut tree population using same SSR markers. Alleles size on locus CnCirA3 range 210-280 bp, with highest frequency on 255bp allele(0,203).
Rajesh et.al.,(2008)and
Ribeiro et al (2010) found 228-240bp alleles sizes. The alleles sizes on the CNZ21 locus that range 180-300bpwith
highest frequency on 240bp (0,207), on locus
CNZ5I
range I 10-230 bp with the highest frequency 0,221 on 140 and 160 bp. Pandin et al (2009) found 270bp on lociCNZ2I
and I l0bp on loci CNZ5 .Total alleles were found in this research were 80 elleles ,
with
range 9-17alleles
and meanvalues
was 13,3.This result had more variation than others. Perera et al. (2001) found 56 alleles sizes range 3-10 on 33 samples
coconut
in
Srilangka used 8 loci.Thirty
seven alleles were found ontall
and dwarf from nine population fromAgatti and Kavaratti Island.
Lakshadweep,India
(Devakumaret.al.,
2010).
Other
researchthat
used 8microatellites loci from
l4
accession from Kidu India coconut population, found 28 alleles with mean two alleles per locus (Kumar et al.,201l).
Overall, genetic diversity in tall coconuts in Baliwas
veryhigh
(Table 1). Mean valuesof
gene diversity(He)
and observed heterozygosity(Ho)
were 0.8835 and 0.5421, respectively. The highest expected heterozygosity was on lociCNZ?I
(0.9204), and the lowest gene diversity onCNZ5I
(0.8561). Genetic variation foundin
this result higher than others research, e.g research coconut populationin
Brazil, Phillipina, India, Sri Langka (Perera et aI,2003; Devakumar et a1.,2010; Liu, at a1.,2011; Xiao, et al., 2013).Genetic variation
on
thetall
coconut cultivars range value 0.35-0.8166(Table
2),
high
expectedheterozygosity on bulan
tall
(0.816), and loweston
bluluktall (0.35).
The estimate of heterozygosity was high (0.590)for
LaccadiveMicro
Tall
and Laccadive SmallTall
was lowest (0,240) (Devakumar,et.al
2010). Genetic diversityof
10 coconut accessions from six location in Hainan province, China; expected heterozygosity of Haikou Green Tall accession was significantly higher (0.4753) than other tall type(Liu
et al.,201l)
Higher heterozygosity was determined by number of alleles variation and frequency. Higher diversity in tall coconuts in Bali, Indonesia was caused by germplasm variation or natural mutation.
References
Devakumar,
K;
V.Niral,
B.A
Jerard,C
Jayabose,R
Chandramohanan and PM
Jacob. (2010). MicrosatelliteAnalysis of Distint Coconut Accsseions from Agatti and Kavaratti Island Lakshadweep, India. Scientia
Hort
125(2010):309-315
Doyle
J.J. andDoyle J.L.
(1987).A
Rapid
DNA
Isolation Procedurefor
Small Amount
of
Leaf
Tissue. Phytochem.Bull.
l9
:ll-15.
Hutchinson, F.(2001).
DNA
Band Size Semi-Log Plotting: Estimation Size ofDNA
Band by Semi-Log. Cancer Research Center : Science Education Patnership.
Kriswiyanti,
E.(2012). Diversityof
Coconut Cultivar (Cocos nuciferaL.,
Arecaceae)in Bali
Island, Indonesia. Research report, Department of Biology Udayana UniversityKumar, S. P; Manimekalai, R. and Kumari B.D.R. 201
l.
Microsatellite Marker Based Characterizationof
SouthPasific Coconut (Cocos nucifera L.) Accessions. Int. J. Plant Breed. Genet.,
5:34-43.
Liu,
X;
Tang,H;Li, D
and Hou,L.
(2011). Genetic Diversityof
Coconut Cultivarsin
Chinaby
Microsatellite(SSR).Moleculer Plant Breeding 2 (2):
Manimekalai, R and Nagarajan, P. (2010) : SSR and ISSR Markers Based Population
on Genetic Structure of Coconut (Cocos nucifera
L.)
Germplasm Accssion Indian J. of Plant Genetic Resources, 23(t)
: 77 -82.Nei, M.(1987). Moleculer Evolusionary Genetic. Colurhbia University Press, New York, USA
Noel,
K K
J., Edmond,K.K.,
Konan,K
J
L
and
Eugene,K.K.
(2011). Microsatellite genediversity within
Philippines dwarf coconutpalm (Cocos nucifera L.) resources at Port-Bout
d'Ivoire.
Scientific research Essays 6(28):5986-5992.
Novarianto,
H.,
Akuba,R.H.,
Mashud,N.,
Tenda,E
and Kumaunang, J.(2005). Statusof
Coconut GeneticResoures Research
in
Indonesia.In
Coconut Genetic Resources. Batugal, P., Ramanatha Rao,V.
and Oliver, J.100
.:
*
Joumal ol Biology, Agriculture and Healthcare ISSN 2224-1208 (Paper) ISSN 2225-093X (Onlinr') Vol.3, No.l3.2013
(eds). tnternational Plant
Genetic
Resources Institute-
Regionai Offrcefor
Asia, the Pacific and OceaniaIPGRI-APO), Serdang, Selangor D.E, lvlalaysia. 608-617.
Pandin, D.S., Hartana,
A.,
Aswidinnoor,H.
and Setiawan,A.
(2008). Parentage analysisof
Mapanget TallCoconut No.32
(DMT-32)
population via geneflow
based on Micro-satellite Markers(SSR). Littri
Journal, 14(4):
l3l-140.
Pandiq D.S.(2010).
DNA
Marker in Coconut (Cocosnuciferal.)
Breeding Programm. Perspektif 9(l):
2l-35.
Perera, L; Russel, R., Provan, J.and Powell, W. (2001). Levels and Distribution of Genetic
Diversitytof
Coconut(Cocos nucifera L., var Typica from typica) from Sri Lanka by Microsatellite Markers. Euphytica
122:381-389.
Perera
L.,
RussellJ.R.,
ProvanJ., and Powell
W.,
2003,
Studlng
genetic relationships among coconut varieties/populations using microsatellitemarkers, Euphytica,132(l):
l2l-128.
Sambrook, J., Fritsclu and Maniatis,
T.
(1989). Moleculer Cloning.A
Laboratory Manual Second Edition. ColdSpring Harbor [,aboratory Press.
Tegelstom H. (1986). Mitochondrial
DNA
in Natural Population : an Improved Routine for Screening of Genetic Variation Base on Sensitive Silver Staining. Electrophoresis. 7: 226-229Xiai,Y.,
Lou, Y.,Yang,Y-,Fan, H.,Xia, W.,Mason,AS., Zhao, S., Sager,R., and Qiao,F. (2013). Developmentof
Microsatellite Markers
in
Cocos nucifera and their applicationin
evaluating the levelof
GeneticDiversity
of
Cocos nucifera POJ: 6(3):193-200.
lvww.llste.org
tMJT
NT