Induction of isoflavonoid pathway in the model legume
Lotus
japonicus
: molecular characterization of enzymes involved in
phytoalexin biosynthesis
Norimoto Shimada, Tomoyoshi Akashi, Toshio Aoki, Shin-ichi Ayabe *
Department of Applied Biological Sciences,Nihon Uni6ersity,Fujisawa,Kanagawa252-8510,Japan
Received 3 May 2000; received in revised form 1 August 2000; accepted 1 August 2000
Abstract
Treatment of the seedlings ofLotus japonicus, a model legume for molecular genetic studies, with reduced glutathione (GSH) resulted in the accumulation of an isoflavan phytoalexin, vestitol. Using PCR strategies based on the conserved amino acid sequences, full length P450 cDNAs were obtained from GSH-treated seedling roots. When the clones,LjCYP-1 (CYP93C family) andLjCYP-2 (CYP81E family), were heterologously expressed in yeast, the proteins exhibited 2-hydroxyisoflavanone synthase (IFS) and isoflavone 2%-hydroxylase (I2%H) activities, respectively. The transcription levels ofLjCYP-1,LjCYP-2 and isoflavone reductase, which are all involved in vestitol biosynthesis, coordinately increased upon elicitation. Genomic Southern blot analysis indicated that the IFS gene forms a small gene family and a single copy of the I2%H gene is present in theL.japonicusgenome.
Molecular biological aspects of P450s involved in the isoflavonoid pathway and the genomic approach to flavonoid metabolism in this unique plant are discussed. © 2000 Elsevier Science Ireland Ltd. All rights reserved.
Keywords:Lotus japonicus; Fabaceae; Cytochrome P450; Isoflavonoid; Vestitol
www.elsevier.com/locate/plantsci
1. Introduction
Flavonoids possess a basic C6C3C6 skeleton
derived from a phenylpropanoid (C6C3) and three
C2 units. Extensive biosynthetic modifications
bring about remarkably diverse structures to which multiple physiological functions are at-tributed. Like the widely occurring anthocyanin pigments in vascular plants, some flavonoids act as screens from harmful UV-light and scavengers of
reactive oxygen species [1]. Major ‘polyphenols’, which are alleged to be beneficial to human health, are flavonoids [2,3].
Leguminous flavonoids are very important in the interactions of the producer plants with envi-ronmental organisms [4,5]. Isoflavonoids, a
dis-tinct class of flavonoid with rearranged C3 parts,
are distributed almost exclusively in this family; they are typical legume phytoalexins active in the defense against phytopathogenic organisms [6]. Flavonoids are also important in the symbiotic relationship of leguminous plants with soil bacte-ria to establish nitrogen-fixing root nodules. In the
initial stage of nodulation, recognition of
flavonoids by NodD proteins of specific Rhizobia
leads to activation of other nod genes [7,8].
Ex-pression of genes encoding enzymes of the
flavonoid/isoflavonoid pathway in developing
al-falfa root nodules has also been reported [9]. The functions of flavonoids in the symbiosis with ar-Abbre6iations: GSH, reduced glutathione; IFS,
2-hydroxyisofla-vanone synthase; I2%H, isoflavone 2%-hydroxylase; IFR, isoflavone reductase; P450, cytochrome P450; RACE, rapid amplification of cDNA ends; RT-PCR, reverse transcription-polymerase chain reac-tion.
The nucleotide sequence data reported in this paper have been deposited in the DDBJ, EMBL and GenBank databases under the following accession numbers: LjCYP-1, AB024931; LjCYP-2, AB025016
* Corresponding author. Tel.: +81-466-843703; fax: + 81-466-801141.
E-mail address:[email protected] (S.-i. Ayabe).
bascular fungi [10] and in the defense against the herbivores have also been suggested [5]. There is, thus, a strong interest in the mechanism of the discrimination between pathogenic and symbiotic interactions with environmental organisms by the host plant cells and their responses, in which flavonoids may play crucial roles.
The biosynthetic scheme of flavonoids as well as their physiological functions described above is as complicated as other pathways of secondary metabolism. A possible regulation network in flavonoid metabolism may exist for the maximal adaptation of the plant to its environment. Thus, there should be organ-specific and temporally reg-ulated expression of the genes required for the construction of specific structural classes of flavonoids in response to pathogen attack and during the establishment of symbiotic relation-ships with microorganisms. Such a total view of secondary metabolism would be explored best in model plants in which genome organization has been clarified and a genetic approach through, for example, positional cloning and functional charac-terization of genes by sense- and antisense-sup-pression is possible. Genetic studies by searching mutants and identifying responsible genes would
also be beneficial. In Arabidopsis thaliana, a
non-legume model plant, several mutants of flavonoid biosynthesis have been described, and genetic analysis of these has greatly contributed to the understanding of flavonoid physiology and
bio-chemistry [11]. Of course, A. thaliana cannot be
the model plant for the investigation of legume-specific phenomena, i.e. symbiotic nitrogen-fixa-tion and isoflavonoid biosynthesis.
In 1992, Lotus japonicus(Regel) Larsen, a
non-agricultural species, was proposed as a model plant that has beneficial traits for classical and molecular genetics of the legume family [12]. It is
a diploid (with the chromosome number n=6)
and has a small genome size and a short genera-tion time. Also, it is self-fertile, and
transforma-tion by Agrobacterium infection is possible. The
utility of this plant for fundamental studies has been demonstrated [13]. So far, for investigation of the mechanisms that underlie the nitrogen fixation and nodulation, analysis of the symbiotic mutants that were induced by the chemical mutagen ethyl-methane sulfonate, T-DNA insertion and transpo-son tagging has been performed [14 – 17]. Genes specifically expressed during nodule development were also identified [18,19]. However, only few studies have been made of flavonoids and their
biosynthesis in L. japonicus.
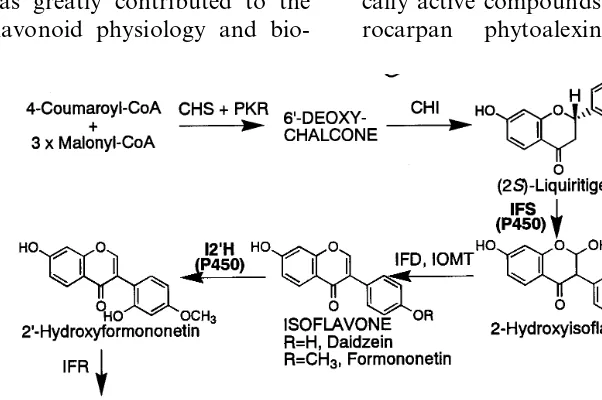
As shown in Fig. 1, a cytochrome P450 (P450), 2-hydroxyisoflavanone synthase (IFS), catalyzes the hydroxylation associated with aryl migration of flavanone to form the isoflavonoid skeleton
[20 – 22]. Another P450, isoflavone 2%-hydroxylase
(I2%H), is essential in the biosynthesis of
biologi-cally active compounds such as isoflavan and
pte-rocarpan phytoalexins. Therefore, for the
investigations of the leguminous plant responses to various environmental factors, focusing attention on the P450s that catalyze critical steps of isoflavonoid biosynthesis would be a promising
avenue. Recently, cDNAs of P450s of the
flavonoid pathway have been cloned, e.g. (2S
)-flavanone 2-hydroxylase (F2H) [23], I2%H [24] and
dihydroxypterocarpan 6a-hydroxylase (D6aH)
[25], in addition to flavonoid 3%- and 3%,5%
-hydroxy-lase (F3%H and F3%5%H) [26,27]. These P450s are
classified as aromatic hydroxylases (F3%H, F3%5%H
and I2%H) and non-aromatic hydroxylases (F2H
and D6aH). F2H and D6aH belong to the CYP93 family. Because IFS catalyzes the hydroxylation of non-aromatic carbon and its substrate is flavanone as well as F2H, the cloning of IFS cDNA by PCR based on amino acid sequences conserved in the CYP93 family was feasible. In addition to P450s, isoflavone reductase (IFR) (a NADPH-dependent reductase) is also critical in isoflavonoid biosyn-thesis, and its cDNA has been cloned from several leguminous plant species. IFR catalyzes the reduc-tion of the heterocyclic ring of isoflavone to form isoflavanone, which is an intermediate of ptero-carpan and isoflavan synthesis.
In this paper, we first describe the examination
of the exudate from L. japonicus seedlings elicited
with reduced glutathione (GSH) and the finding of the induction of isoflavonoid biosynthesis. Then, we present the molecular cloning of cDNAs
in-volved in isoflavonoid biosynthesis inL.japonicus,
their genomic organization, and the effect of GSH on transcription of these genes.
2. Materials and methods
2.1. Elicitation of L. japonicus seedlings
Seeds (30 seeds per culture pot) of L. japonicus
accession B-129 ‘Gifu’ were sown on sterilized filter paper and grown in the dark at 25°C. Four days after sowing, 1 ml of aqueous GSH (10 mM) was added directly to seedlings under sterile condi-tions. After elicitation, seedlings were harvested,
frozen in liquid nitrogen and stored at −80°C
until use for isolation of mRNA. Seedling exu-dates were extracted from the filter paper for 2 – 3 min in 2 ml of 99.5% ethanol at room temperature (three times). The extracts were evaporated under vacuum and dissolved in 1 ml of methanol, passed
through an Ekicrodisc®3 filter (Gelman Sciences,
Japan), dried and stored at −20°C. HPLC
analy-sis was performed using a Shim-pack CLC – ODS
column (6.0×150 mm; Shimadzu) with 55% (v/v)
methanol and 3% (v/v) acetic acid in H2O at a
flow rate 1 ml/min at 40°C. The eluent was
moni-tored at 280 nm.
2.2. Isolation of P450 cDNA fragments by PCR
The template cDNA was prepared from the
roots of GSH-treated seedlings by Dr M.
Kawaguchi (Graduate School of Arts and Sci-ences, The University of Tokyo). Degenerate oligonucleotide primers were designed from highly conserved amino acid regions of CYP75A1,
CYP93A1 and CYP93B1. The primers, 93/S1=5%
-NGCHGGNACNGAYAC-3%, 93/S2=5%
-TGG-GCHHTNGYNGA-3%, 450/AS1=5%
-YCTYTC-NGGRYKRAAYTC-3%and 450/AS2=5%
-TDCC-NGMNCCRAANG-3%, were used to amplify
CYP93 fragments. In addition, the primers, 81/
S2=5%-AARAGRTAYTAYGGNGRNGA-3%and
81/AS2=5%-RAANCKYTCNGGYTTRAA-3%,
were used to amplify CYP81 fragments. For the amplification of CYP93 fragments, the first PCR
was performed with 93/S1 and 450/AS2 primers
(2.5 pmol each) using Taq DNA polymerase
(Takara) in a final volume of 20 ml according to
the manufacturer’s protocol. The PCR was per-formed at 94°C for 3 min followed by 30 cycles of (94°C for 30 s, 37°C for 30 s, 72°C for 1 min) and
final extension at 72°C for 5 min. 93/S2 and
450/AS1 primers and 0.5 ml of the first PCR
product solution as the template were used for the second PCR (nested-PCR) under the same condi-tions except that the annealing temperature was 43°C. The cDNA fragments obtained were named
Lj-1, Lj-3 and Lj-4. For the amplification of
CYP81 fragments, PCR was performed with 81/S2
and 81/AS2 primers under the same conditions as
the second PCR, and Lj-2 was obtained. PCR
products were ligated into a plasmid vector pT7Blue T-Vector (Novagen) and sequenced using a LIC-4000 DNA sequencer (Aloka).
2.3. Rapid amplification of cDNA ends (3%- and
5%-RACE)
For 3%- and 5%-RACE, an adaptor sequence was
GSH-treated seedling roots using a Marathon™ cDNA Amplification Kit (Clontech). Four specific
primers were designed based on the Lj-1 andLj-2
sequences, 93R/S1=5%
-AGGCTTGTTGACGA-ATCAGATA-3%, 93R/AS1=5%
-TTCCAGTATT-TTGGGTCTCTTTGC-3%, 81rS1=5%
-GCCACA-ACTTTCAAGCCAGAGA-3% and 81rAS1=5%
-ACTGCAGCAACTCGGTCACCAT-3%. Primer
sets (0.5 pmol each) for 3%-RACE were 93R/S1 or
81rS1 and adaptor primer AP1 (5%
-CCATC-CTAATACGACTCACTATAGGGC-3%,
Mara-thon™ cDNA Amplification Kit, Clontech). 93R/
AS1 or 81rAS1 and AP1 were used for 5%-RACE.
Reaction conditions were as described in the man-ufacturer’s manual.
2.4. Cloning of full length P450 cDNAs
Two sets of specific primers containing BamHI,
XhoI, KpnI or XbaI sites (shown in bold type)
were designed to obtain full length DNAs,
93BamHI=5%-AATTCAAGGATCC
AGCAAAC-ACCA-3% and 93Xho=5%-AAACCCTCGAG
CA-CAAAGCAACAT-3% (for Lj-1); 81+Kpn=5%
-AATGGTACC
ATGGATATCATCTCCTTCCTT-3% and 81−Xba=5%-CTCACTCTAGA
AACAT-GTCCCCGATTCAAA-3% (for Lj-2). PCR was
performed using KOD DNA polymerase
(Toy-obo) in a final volume of 20 ml according to the
manufacturer’s protocol. The protocol for the
am-plification of full length Lj-1 was 98°C for 1 min
followed by 25 cycles of (98°C for 15 s, 55°C for 10 s, 74°C for 30 s) and final extension at 74°C for
10 min. The amplification of full length Lj-2 was
performed as described above except that
anneal-ing temperature was altered to 60°C. The 5%-ends
of PCR products (ca.1.5 kbp) were phosphory-lated with T4 polynucleotide kinase (Takara) and
ligated into a dephosphorylated EcoRV site of
pBluescript II SK- (Stratagene). These clones were
named LjCYP-1 and LjCYP-2, respectively.
2.5. Cloning of LjIFR cDNA
Degenerate PCR was carried out with primers designed from known IFR amino acid sequences,
IFR/S1=5%-AARGCHATHAARGARGC-3% and
IFR/AS1=5%- ATCXCCYTTDATYTGYTG - 3%,
and subsequently 3%- and 5%-RACE were
per-formed using IFRR/S1=5%
-TGTTGCCATGC-CTTCACTGGTT-3% and IFRR/AS1=5%
-CCA-ATCTTCTTCTTCCCACAATGC-3%. PCR
condi-tions were the same as described in ‘Isolation of
P450 cDNA fragments by PCR’ and ‘Rapid am
-plification of cDNA ends’. Annealing temperatures were altered to 43 and 68°C, respectively. Based on their sequences, specific primers containing
BamHI or XhoI sites (shown in bold type), were
designed to amplify full length IFR cDNA:
IFRBam2=5%-GTTCTGGATCC
TGATGGCAC-CACAAGACAG-3%, IFRXhoI=5%
-TCACGAG-AGGACAGGCTCGAGTAACAAACA-3%. PCR
was carried out with the same procedure as
de-scribed in ‘Cloning of full length P450 cDNAs’.
Annealing temperature was altered to 57°C for
LjIFR amplification.
2.6. Heterologous expression in yeast cells,
preparation of microsomes and enzyme assay
The P450 cDNAs were digested with BamHI
and XhoI or KpnI and XbaI and ligated into
corresponding sites of pYES2 (Invitrogen) to con-struct the expression plasmid vector pYESLjCYP-1 and pYESLjCYP-2. Introduction of these
plasmids into Saccharomyces cere6isiae strain
BJ2168, selection of transformants, induction of P450 proteins and preparation of microsomes were
performed as described [23]. The assays ofLjCYP
-1 and LjCYP-2 proteins were performed as de-scribed [24,28].
2.7. mRNA preparation and re6erse transcription
(RT)-PCR analysis
mRNA was isolated from elicitor-treated
seedlings (fresh weight 0.2 g) using Straight A’s mRNA isolation system (Novagen), and 500 ng each of mRNAs were used to synthesize single-strand cDNAs. For RT-PCR, the same primers as described in ‘Cloning of full length cDNAs’ were
used for amplification of IFS, I2%H and IFR.
RT-PCR was carried out with 0.5 pmol each of specific primers using Taq DNA polymerase in a
final volume of 20 ml according to manufacturer’s
protocol. The reaction was performed at 95°C for 1 min followed by 24 cycles of (95°C for 30 s, 57°C for 30 s, 72°C for 1 min) and a final
exten-sion at 72°C for 5 min. The products (5 ml) were
2.8. Southern blot analysis
Genome DNAs were isolated from seedlings of
L. japonicus accession B-129 ‘Gifu’ using the
cetyltrimethylammonium bromide (CTAB)
method. Genome DNAs (10 mg each) were
di-gested with DraI, EcoRI, HindIII or XbaI. The
restricted fragments were separated by elec-trophoresis on 0.7% agarose gel and transferred to Nylon membranes, positively charged (Roche). Hybridization probes were prepared by a random priming method using DIG-11-dUTP (Roche). Hybridization and wash were done according to the DIG hybridization protocol (Roche).
3. Results
3.1. Accumulation of 6estitol in elicited L.
japonicus seedlings
GSH is the most abundant thiol compound in living cells. In higher plants, GSH is assumed to play a role in protection against oxidative damage. GSH stimulates the activation of defense re-sponses and the production of phytoalexins [29].
In birdsfoot trefoil (Lotus corniculatus), a
te-traploid relative of L. japonicus, isoflavan
phy-toalexins, vestitol and sativan, are induced to accumulate in response to inoculations with fungi [30]. After treatment with GSH, vestitol and
sati-van also accumulated in Agrobacterium rhizoge
-nes-transformed L. corniculatus root cultures
[31,32]. Thus, in this study, GSH was used as an elicitor.
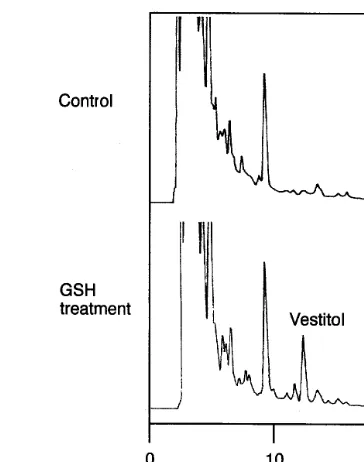
Fig. 2 shows the effect of GSH treatment on the
composition of the materials released from L.
japonicus seedlings. The accumulation of vestitol was clearly observed in the exudate of seedlings 10 h after the GSH treatment. The amount of vestitol exuded 10 h after GSH-treatment was estimated to
be 37.7 ng per seedling (5.14 mg/g fresh weight).
Levels of vestitol accumulation 10 and 20 h after GSH-treatment were constant (data not shown).
Other (iso)flavonoid compounds were not
identified.
3.2. Cloning of cDNAs encoding isofla6onoid
biosynthesis
To isolate P450 cDNAs, the PCR strategy was
employed using GSH-treatedL. japonicusseedling
Fig. 2. HPLC profile of exudates of L. japonicus seedlings. After 10 h elicitation with 10 mM GSH, an isoflavan vestitol, which was not detected in the control (H2O treatment), had accumulated.
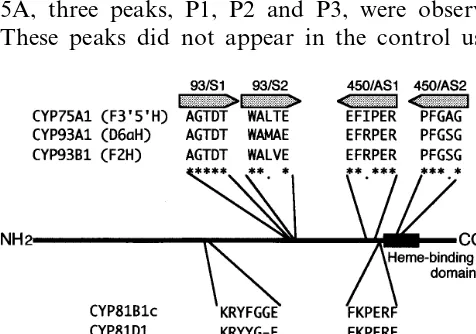
roots. Six degenerate oligonucleotide primers were designed from highly conserved regions of known plant P450s of the flavonoid pathway (Fig. 3).
Nested-PCR with 93/S2 and 450/AS1 primers
us-ing the product of the first PCR with 93/S1 and
450/AS2 primers as the template gave
primer-spe-cific fragments. Sequence analysis revealed that three clones were similar to known plant P450s.
These clones were named Lj-1,Lj-3 andLj-4, and
their sequences were about 78, 80 and 85% identi-cal at the nucleotide level to CYP93C1, CYP73A (trans-cinnamate 4-hydroxylase) and CYP98A2,
respectively. PCR with 81/S2 and 81/AS2 primers
amplified a 700 bp fragment of expected size. Sequence analysis revealed that this fragment showed 73% identity to CYP81E1 at nucleotide
level. This fragment was named Lj-2.
Two sets of specific primers were designed based
on the Lj-1 and Lj-2 sequences, and 3%- and
5%-RACE were performed. 3%-RACE withLj-1 and
Lj-2 yielded fragments of about 830 and 700 bp,
respectively, each containing the stop codons and
3%-non-coding sequences. Similarly, 5%-RACE with
Lj-1 andLj-2 yielded fragments of about 1000 and
730 bp containing initiation codons.
Using two sets of re-designed specific primers and the cDNA of the roots of GSH-treated seedlings as the template, two full length P450
LjCYP-1 contains an ORF consisting of 1554 bp encoding 518 amino acids, and showed 83.6%
identity at the amino acid level to Glycyrrhiza
echinataCYP93C2 [28] (see Fig. 4A). The ORF of
LjCYP-2 consists of 1497 bp (499 amino acids)
with 82.2% identity at the amino acid level to G.
echinataCYP81E1 [24] (see Fig. 4B). The deduced
amino acid sequences of LjCYP-1 and LjCYP-2
displayed the consensus sequence (F/W)(S/G/N/
H)X(G/D)X(R/H/P/T)XC(L/I/V/M/F/A/P)(G/A/ D) of P450s for the heme-binding region. LjCYP-1 and LjCYP-2 proteins were named CYP93C17 and CYP81E6, respectively, by P450 Nomencla-ture Committee.
Full length cDNA of putative IFR was also obtained from GSH-treated seedlings and named
LjIFR. The deduced amino acid sequence of
LjIFR is 77.4, 79.6, 81.1 and 83.0% identical to soybean, pea [33], chickpea [34] and alfalfa [35]
IFR cDNAs, respectively.
3.3. Catalytic functions of LjCYP-1 and LjCYP-2
proteins
To examine the catalytic functions of LjCYP-1 and LjCYP-2 proteins, the cDNAs were expressed in yeast cells.
After induction, the microsome of the recombi-nant yeast expressing LjCYP-1 was incubated with
(RS)-liquiritigenin and NADPH, and the ethyl
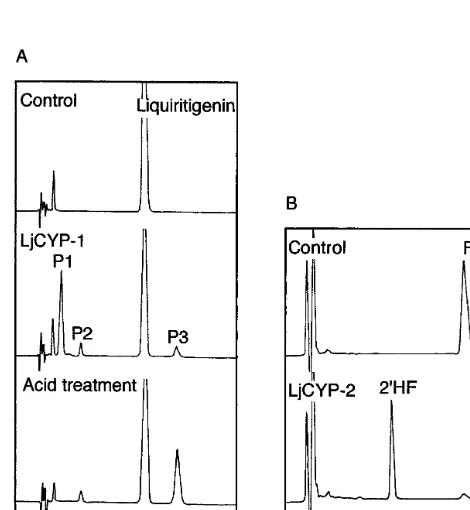
acetate extract of the reaction mixture was ana-lyzed by reverse-phase HPLC. As shown in Fig. 5A, three peaks, P1, P2 and P3, were observed. These peaks did not appear in the control using
the vector without the insert. The Rt’s of P1 and
P3 were identical to those of 2,7,4%
-trihydroxyisofla-vanone and 7,4%-dihydroxyisoflavone (daidzein),
respectively, and P2 to putative 3-hydroxyliquiriti-genin. These compounds have recently been iden-tified in the reaction with CYP93C2 protein (IFS of
G. echinata) [28]. On acid treatment of the ethyl acetate extract, the P1 peak decreased whereas the P3 peak increased. Furthermore, the chemical struc-ture of P3 was confirmed to be daidzein by electron impact mass spectrometry (data not shown).
There-fore, it was concluded that LjCYP-1 encodes IFS.
The microsome of yeast cells transformed with LjCYP-2 was incubated with formononetin and NADPH, and the reaction product was analyzed by HPLC. As a result, a new peak that was not observed in the control reaction appeared, and its
retention time was the same to that of 2%
-hydroxy-formononetin (Fig. 5B). The identity of the
reac-tion product was further confirmed by
co-chromatography. Therefore, LjCYP-2 protein
was shown to be I2%H.
3.4. Expression of genes encoding enzymes of the isofla6onoid pathway
mRNA was isolated from elicitor-treated
seedlings harvested at different times after elicita-tion. For RT-PCR, the quantity of each template was adjusted to give roughly equal amplification of actin cDNA. As shown in Fig. 6, the transcript
levels of IFS, I2%H and IFR all increased 10 h after
elicitation. Only a low level of transcription was observed in untreated seedlings. The transcripts had decreased to the control level by 20 h. The transient accumulation of mRNAs of these P450s and IFR in the GSH-treated seedlings indicated a coordinated expression of essential enzymes of vestitol biosynthesis upon elicitation.
3.5. Genomic organization of isofla6onoid
biosynthetic genes
Genomic DNA was digested with several re-striction enzymes, and Southern blot analysis was performed using the P450 ORFs as probes (Fig. 7).LjCYP-1 has one restriction site forEcoRI and
no restriction sites for DraI, HindIII or XbaI.
With this probe, one strong band from each
EcoRI or XbaI-digested DNA, two bands from
Fig. 4. Amino acid sequence alignments of LjCYP-1 and LjCYP-2. Gaps (-) are inserted to optimize the alignments. Positions with identical amino acid residues in both sequences are in reverse type. (A) Comparison of the LjCYP-1 deduced amino acid sequence with that of theG.echinata(CYP93C2, AB023636) [28],G.max(CYP93C1v2, AF135484) [37], mung bean (AF195806) [39], pea (AF195812) [39] and red clover (AF195810) [39] IFS sequences. LjCYP-1 exhibited 80.9 and 83.6% identity to CYP93C1v2 and CYP93C2, respectively. (B) Comparison of the LjCYP-2 deduced amino acid sequence with that of theG.echinata(CYP81E1, P93147) I2%H sequence [24]. 410 of the 499 amino acids are identical (82.2%).
HindIII-digested DNA, respectively, were
ob-served. In addition, 2 weak bands appeared with
EcoRI-digested DNA, and 1 weak band was
ap-parent with XbaI-digested DNA.
On the other hand, using LjCYP-2 as a probe,
only one strong band was observed in each lane
except for DraI-digested DNA. Because LjCYP
-2 has no restriction sites for these four enzymes,
two bands in DraI lane may be due to the
pres-ence of a restriction site in the intron sequpres-ence.
These results indicate that, in the L.
japonicus genome, two or three copies of the IFS
gene are present while only one copy of I2%H
4. Discussion
Changes in flavonoid metabolism in leguminous plants are very important in successful defense responses and in symbiotic nitrogen fixation.
Isoflavonoid phytoalexins ofL. japonicushave not
been studied yet. In this study, induction of vesti-tol synthesis was demonstrated, for the first time, in the seedlings after GSH treatment. Full length
cDNA clones, LjCYP-1 and LjCYP-2 encoding
IFS and I2%H, respectively, andLjIFR conceivably
encoding IFR, were also isolated from GSH-treated seedling roots.
GSH is involved in the oxidation – reduction cycle in living cells. The phytoalexin production induced by the exogenously added GSH is likely to be a consequence of the perturbation of the oxidative state in the cells that mimics some stages of the signal transduction pathway during
elicita-Fig. 6. RT-PCR analysis of IFS, I2%H and IFR expression in
GSH-elicited or H2O-treated (20 h) seedlings. Reverse tran-scription was performed using 500 ng each of mRNAs. The concentration of each template for PCR was adjusted to give roughly equal amplification of actin. Amplification efficiency was considered to be linear during the PCR (30 cycles for IFS; 24 cycles for others).
tion [29]. In L. corniculatus, treatment with GSH
as well as with a fungal elicitor induced a transient increase of phenylalanine ammonia-lyase activity [31,32] and phytoalexin synthesis. In bean suspen-sion cultures, the selective induction of defense
Fig. 5. HPLC profiles of the reaction products with the microsomes of yeast cells expressing LjCYP-1 or LjCYP-2. (A). The microsome of yeast cells expressing LjCYP-1 cata-lyzed the formation of 2-hydroxyisoflavanone (P1), small amounts of 3-hydroxyflavanone (P2; byproduct of IFS reac-tion) and the isoflavone, daidzein (P3), from a 5-deoxyfla-vanone, liquiritigenin. On acid treatment, P1 was readily converted to P3. (B) HPLC of the reaction product from an isoflavone, formononetin, with LjCYP-2 protein expressed in the recombinant yeast microsome. F, formononetin; 2%HF,
2%-hydroxyformononetin. No reaction was observed with the microsome of control yeast cells harboring the vector only.
genes by GSH treatment has been reported [36]. In
L. japonicus, too, the level of mRNAs for IFS,
I2%H and IFR increased coordinately after GSH
treatment as shown by RT-PCR analysis (Fig. 6), suggesting that the biosynthesis of phytoalexin
vestitol in L. japonicus is regulated in the same
manner as in bean and in L. corniculatusby GSH
at the transcriptional level. The result obtained here with the model plant having a small genome size should be useful in the elucidation of the intracellular signaling pathway, possibly in the near future, leading not only to flavonoid gene expression but also to total changes of the cell metabolism after oxidative stress using the DNA array systems and through the analysis of artifi-cially tagged mutants in GSH perception.
IFS is the central enzyme of isoflavonoid metabolism that is present almost exclusively in legumes [6]. However, in spite of the very intense interest in its physiological roles and unique reac-tion scheme, IFS has long been uncharacterized because of the difficulty in enzyme purification and in vitro reconstitution. In 1999, IFS cDNAs of soybean and licorice were first cloned based on amino acid sequence information combined with functional analysis of heterologously expressed re-combinant proteins [28,37]. These were shown to be the P450s of CYP93C family, CYP93C1v2 of soybean and CYP93C2 of licorice. Very recently, more soybean IFS cDNAs having very high ho-mology (ca. 96% identity at the amino acid level) to CYP93C1v2 were reported [38] together with many cDNAs of surprisingly high homology in various leguminous plants (red clover, white clover, hairy vetch, mung bean, alfalfa, lentil, snow pea and lupine) and even in a non-legume, the sugarbeet [39]. The present study adds another IFS of the legume to the CYP93C array, and the functional identification of its enzymatic activity confirms that the P450 of CYP93C family cata-lyzes the unique aryl migration accompanied by hydroxylation. It is noteworthy that the homology
of IFS cDNAs between L. japonicus, licorice and
soybean is 80 – 83%, a typical value for the similar-ity of the DNA sequences of genes with the same function among the different plant species, while that of soybean and other plant species including the sugarbeet reported by Jung et al. [39] is excep-tionally high (95 – 99% similar to each other).
In the assay with recombinant licorice [28] and
L. japonicus CYP93C P450s, the native reaction
product, 2-hydroxyisoflavanone, was unambigu-ously characterized (Fig. 5A). In the assay with soybean CYP93C P450s, dehydrated isoflavones were reported to be the sole or the major products [37,38]. Sequence analysis showed that IFSs of
licorice and L. japonicus share several common
features (e.g. V3, V45, D52, P54, Y67, I131, S153, E155, R261 and N262) that soybean IFSs and others most recently reported lack (Fig. 4). There may be two or more types of IFS in the plant genome, and the reaction characteristics catalyzed by those proteins may be slightly different.
The structures of major isoflavonoid-derived phytoalexins of soybean and plants like licorice and Lotus are characteristically different. In soy-bean, daidzein is the basic structure and further modifications of the pterocarpan skeleton, includ-ing 6a-hydroxylation and prenylation, lead to
glyceollins [6]. In contrast, in licorice and Lotus,
formononetin, a 4%-methylated isoflavone, is the
precursor of the phytoalexins, and no further
hy-droxylation (except for 2%-hydroxylation) and
prenylation are involved in their biosynthesis. The
similarity of L. japonicus and licorice IFSs may
reflect these structural features of phytoalexins. The methylation step of formononetin biosynthe-sis has not been elucidated: daidzein cannot be
4%-O-transmethylated fromS-adenosyl-L
-methion-ine in vitro by the extracts of the plants producing
formononetin but only 7-O-transmethylated to
give isoformononetin, an unnatural product
[40,41]. There have been concepts of enzyme inter-action of IFS and methyltransferase proteins to
perform correct O-methylation in formononetin
biosynthesis [42]. We have proposed a new biosyn-thetic scheme involving 2-hydroxyisoflavanone as the methyl acceptor [43], and the possible involve-ment of protein interactions in formononetin
biosynthesis would be studied best in L. japonicus
taking advantage of the relative simplicity of its genome organization.
Genomic Southern analysis (Fig. 7) indicated that two or three copies of the IFS gene exists in
L. japonicus. Because IFS is present at the central branching point in the flavonoid biosynthesis and the isoflavonoids exhibit various biological activi-ties, it may be presumed that the expression of different IFS genes is regulated by different signals during development, in different organs and in response to different environmental stimuli.
The functional cloning of I2%H cDNA confirms
CYP81E family. In addition to our identification
of licorice [24] and L. japonicus I2%H cDNAs,
other CYP81E P450s have been identified [44] in
chickpea in which I2%H enzyme activity was earlier
reported [45]. It is very interesting that the I2%H
gene was found to be present as a single copy inL.
japonicus while at least two copies exist in
chick-pea [44]. The regulation mechanism of I2%H
ex-pression in response to infection by pathogens or symbionts like Rhizobia should be studied in this and other leguminous species in the future.
IFS catalyzes the first reaction in the
isoflavonoid pathway, and I2%H is essential for
isoflavan and pterocarpan phytoalexins biosynthe-sis. Thus, the cloning of these cDNAs should provide a valuable tool in analyzing the role of isoflavonoid compounds in plant-pathogen inter-actions and in nodulation. To clarify the physio-logical role of isoflavonoid compounds in legumes,
introduction of sense or antisense IFS and I2%H
cDNA constructs into L. japonicus is now in
pro-gress in this laboratory.
Acknowledgements
We thank Dr M. Kawaguchi (The University of
Tokyo) for the gift of the cDNA of L. japonicus
and Dr David R. Nelson (University of Tennessee)
for the assignment of P450 sequences to
CYP93C17 and CYP81E6. This work was sup-ported by a Grant-in-Aid (No. 09640782) from the Ministry of Education, Sports, Science and Cul-ture of Japan.
References
[1] R.E. Koes, F. Quattrocchio, J.N.M. Mol, The flavonoid biosynthetic pathway in plants: function and evolution, BioEssays 16 (1994) 123 – 132.
[2] R.A. Dixon, C.L. Steele, Flavonoids and isoflavonoids — a gold mine for metabolic engineering, Trends Plant Sci. 4 (1999) 394 – 400.
[3] B. Stavric, in: T. Johns, J.T. Romeo (Eds.), Recent Advances in Phytochemistry, Functionality of Food Phy-tochemicals, vol. 31, Plenum Press, New York, 1997, pp. 53 – 87.
[4] J.B. Harborne, in: F.A. Bisby, J. Buckingham, J.B. Harborne (Eds.), Phytochemical Dictionary of the Legu-minosae, vol. 1, Chapman & Hall, London, 1994, pp. XX – XXII.
[5] H.A. Stafford, Role of flavonoids in symbiotic and defense functions in legume roots, Bot. Rev. 63 (1997)
27 – 39.
[6] P.M. Dewick, in: J.B. Harborne (Ed.), The Flavonoids. Advances in Research Since 1980, Chapman & Hall, London, 1986, pp. 125 – 209.
[7] N.K. Peters, J.W. Frost, S.R. Long, A plant flavone, luteolin, induces expression ofRhizobium meliloti nodula-tion genes, Science 233 (1986) 977 – 980.
[8] J.W. Redmond, M. Batley, M.A. Djordjeic, R.W. Innes, P.L. Kuempel, B.G. Rolfe, Flavones induce expression of nodulation genes inRhizobium, Nature 323 (1986) 632 – 635.
[9] H.I. McKhann, N.L. Paiva, R.A. Dixon, A.M. Hirsch, Chalcone synthase transcripts are detected in alfalfa root hairs following inoculation with wild-type Rhizobium meliloti, Mol. Plant-Microbe Interact. 10 (1997) 50 – 58. [10] M.J. Harrison, The arbuscular mycorrhizal symbiosis: an underground association, Trends Plant Sci. 2 (1997) 54 – 60.
[11] B.W. Shirley, Flavonoid biosynthesis: ‘new’ functions for an ‘old’ pathway, Trends Plant Sci. 1 (1996) 377 – 382. [12] K. Handberg, J. Stougaard, Lotus japonicus, an
autoga-mous, diploid legume species for classical and molecular genetics, Plant J. 2 (1992) 487 – 496.
[13] Q. Jiang, P.M. Gresshoff, Classical and molecular genetics of the model legumeLotus japonicus, Mol. Plant-Microbe Interact. 10 (1997) 59 – 68.
[14] H. Imaizumi-Anraku, M. Kawaguchi, H. Koiwa, S. Akao, K. Syono, Two ineffective-nodulating mutants of Lotus japonicus — different phenotypes caused by the blockage of endocytotic bacterial release and nodule maturation, Plant Cell Physiol. 38 (1997) 871 – 881. [15] L. Schauser, K. Handberg, N. Sandal, J. Stiller, T.
Thykjaer, E. Pajuelo, A. Nielsen, J. Stougaard, Symbiotic mutants deficient in nodule establishment identified after
T-DNA transformation of Lotus japonicus, Mol. Gen.
Genet. 259 (1998) 414 – 423.
[16] L. Schauser, A. Roussis, J. Stiller, J. Stougaard, A plant regulator controlling development of symbiotic root nod-ules, Nature 402 (1999) 191 – 195.
[17] K. Szczyglowski, R.S. Shaw, J. Wopereis, S. Copeland, D. Hamburger, B. Kasiborski, F.B. Dazzo, F.J.D. Bruijn, Nodule organogenesis and symbiotic mutants of the model legumeLotus japonicus, Mol. Plant-Microbe Inter-act. 11 (1998) 684 – 697.
[18] P. Kapranov, T.J. Jensen, C. Poulsen, F.J.D. Bruijn, K.
Szczyglowski, A protein phosphatase 2C gene,
LjNPP2C1, from Lotus japonicus induced during root nodule development, Proc. Natl. Acad. Sci. USA 96 (1999) 1738 – 1743.
[19] K. Szczyglowski, D. Hamburger, P. Kapranov, F.J.D. Bruijn, Construction of a Lotus japonicus late nodulin expressed sequence tag library and identification of novel nodule-specific genes, Plant Physiol. 114 (1997) 1335 – 1346.
[21] H. Kessmann, A.D. Choudhary, R.A. Dixon, Stress responses in alfalfa (Medicago sati6aL.) III. Induction of medicarpin and cytochrome P450 enzyme activities in elicitor-treated cell suspension cultures and protoplasts, Plant Cell Rep. 9 (1990) 38 – 41.
[22] G. Kochs, H. Grisebach, Enzymic synthesis of
isoflavones, Eur. J. Biochem. 155 (1986) 311 – 318. [23] T. Akashi, T. Aoki, S. Ayabe, Identification of a
cy-tochrome P450 cDNA encoding (2S)-flavanone 2-hy-droxylase of licorice (Glycyrrhiza echinataL.; Fabaceae) which represents licodione synthase and flavone synthase II, FEBS Lett. 431 (1998) 287 – 290.
[24] T. Akashi, T. Aoki, S. Ayabe, CYP81E1, a cytochrome P450 cDNA of licorice (Glycyrrhiza echinata L.), en-codes isoflavone 2%-hydroxylase, Biochem. Biophys. Res.
Commun. 251 (1998) 67 – 70.
[25] C.R. Schopfer, G. Kochs, F. Lottspeich, J. Ebel, Molec-ular characterization and functional expression of dihy-droxypterocarpan 6a-hydroxylase, an enzyme specific for pterocarpanoid phytoalexin biosynthesis in soybean (Glycine max L.), FEBS Lett. 432 (1998) 182 – 186. [26] T.A. Holton, F. Brugliera, D.R. Lester, Y. Tanaka,
C.D. Hyland, J.G.T. Menting, C.-Y. Lu, E. Farcy, T.W. Stevenson, E.C. Cornish, Cloning and expression of cytochrome P450 genes controlling flower colour, Na-ture 366 (1993) 276 – 279.
[27] F. Brugliera, G. Barri-Rewell, T.A. Holton, J.G. Mason, Isolation and characterization of a flavonoid 3%
-hydroxy-lase cDNA clone corresponding to the Ht1 locus of
Petunia hybrida, Plant J. 19 (1999) 441 – 451.
[28] T. Akashi, T. Aoki, S. Ayabe, Cloning and functional expression of a cytochrome P450 cDNA encoding 2-hy-droxyisoflavanone synthase involved in biosynthesis of the isoflavonoid skeleton in licorice, Plant Physiol. 121 (1999) 821 – 828.
[29] C. Lamb, R.A. Dixon, The oxidative burst in plant disease resistance, Annu. Rev. Plant Physiol. Plant Mol. Biol. 48 (1997) 251 – 275.
[30] M.R. Bonde, R.L. Millar, J.L. Ingham, Induction and identification of sativan and vestitol as two phytoalexins from Lotus corniculatus, Phytochemistry 12 (1973) 2957 – 2959.
[31] M.P. Robbins, J. Hartnoll, P. Morris, Phenylpropanoid defence responses in transgenic Lotus corniculatus. 1. Glutathione elicitation of isoflavan phytoalexins in transformed root cultures, Plant Cell Rep. 10 (1991) 59 – 62.
[32] M.P. Robbins, B. Thomas, P. Morris, Phenylpropanoid defence responses in transgenic Lotus corniculatus. II. Modelling plant defense responses in transgenic root cultures using thiol and carbohydrate elicitors, J. Exp. Bot. 46 (1995) 513 – 524.
[33] N.L. Paiva, Y. Sun, R.A. Dixon, H.D. VanEtten, G. Hrazdina, Molecular cloning of isoflavone reductase from pea (Pisum sati6um L.): evidence for a
3R-isofla-vanone intermediate in (+)-pisatin biosynthesis, Arch. Biochem. Biophys. 312 (1994) 501 – 510.
[34] K. Tiemann, D. Inze, M. Van Montagu, W. Barz, Pterocarpan phytoalexin biosynthesis in elicitor-chal-lenged chickpea (Cicer arietinumL.) cell cultures. Purifi-cation, characterization and cDNA cloning of NADPH: isoflavone oxidoreductase, Eur. J. Biochem. 200 (1991) 751 – 757.
[35] N.L. Paiva, R. Edwards, Y.J. Sun, G. Hrazdina, R.A. Dixon, Stress responses in alfalfa (Medicago sati6a L.)
11. Molecular cloning and expression of alfalfa
isoflavone reductase, a key enzyme of isoflavonoid phy-toalexin biosynthesis, Plant Mol. Biol. 17 (1991) 653 – 667.
[36] V.P. Wingate, M.A. Lawton, C.J. Lamb, Glutathione causes a massive and selective induction of plant defense genes, Plant Physiol. 87 (1988) 206 – 210.
[37] C.L. Steele, M. Gijzen, D. Qutob, R.A. Dixon, Molecu-lar characterization of the enzyme catalyzing the aryl migration reaction of isoflavonoid biosynthesis in soy-bean, Arch. Biochem. Biophys. 367 (1999) 146 – 150. [38] W. Jung, O. Yu, S.-M.C. Lau, D.P. O’Keefe, J. Odell,
G. Fader, B. McGonigle, Identification and expression of isoflavone synthase, the key enzyme for biosynthesis of isoflavones in legumes, Nat. Biotechnol. 18 (1999) 208 – 212.
[39] W. Jung, O. Yu, S.-M.C. Lau, D.P. O’Keefe, J. Odell, G. Fader, B. McGonigle, Supplement to: identification and expression of isoflavone synthase, the key enzyme for biosynthesis of isoflavones in legumes, Nat. Biotech-nol. Web Extras (http://biotech.nature.com/web –extras)
(1999).
[40] M. Hagmann, H. Grisebach, Enzymatic rearrangemant of flavanone to isoflavone, FEBS Lett. 175 (1984) 199 – 202.
[41] R. Edwards, R.A. Dixon, Isoflavone O
-methyltrans-ferase activities in elicitor-treated cell suspension cultures of Medicago sati6a, Phytochemistry 30 (1991) 2597 –
2606.
[42] X.Z. He, J.T. Reddy, R.A. Dixon, Stress responses in alfalfa (Medicago sati6aL.). XXII. cDNA cloning and characterization of an elicitor-inducible isoflavone 7-O-methyltransferase, Plant Mol. Biol. 36 (1998) 43 – 54. [43] T. Akashi, Y. Sawada, T. Aoki, S. Ayabe, New scheme
of biosynthesis of formononetin involving 2,7,4%
-trihy-droxyisoflavanone but not daidzein as the methyl accep-tor, Biosci. Biotechnol. Biochem., in press.
[44] S. Overkamp, W. Barz, Cloning of two Cicer arietinum L. cDNAs encoding cytochrome P450s highly ho-mologous to isoflavone 2%-hydroxylase from licorice (PGR99-102), Plant Physiol. 120 (1999) 935.
[45] W. Hinderer, U. Flentje, W. Barz, Microsomal
isoflavone 2%- and 3%-hydroxylases from chickpea (Cicer
arietinum L.) cell suspensions induced for pterocarpan phytoalexin formation, FEBS Lett. 214 (1987) 101 – 106.