22 Feuillet, C., Schachermayr, G. and Keller, B. (1997) Molecular cloning of a new receptor-like kinase gene encoded at the Lr10 disease resistance locus of wheat, Plant J. 11, 45–52
23 Zhou, J. et al. (1995) The tomato gene Pti1 encodes a serine/threonine kinase that is phosphorylated by Pto and is involved in the hypersensitive response,
Cell 83, 925–935
24 Zhou, J., Tang, X. and Martin, G.B. (1997) The Pto kinase conferring resistance to tomato bacterial speck disease interacts with proteins that bind a
cis-element of pathogenesis-related genes, EMBO J. 16, 3207–3218
25 Ligterink, W. et al. (1997) Receptor-mediated activation of a MAP kinase in pathogen defense of plants, Science 276, 2054–2057
26 Xing, T. Higgins, V.J. and Blumwald, E. (1996) Regulation of plant defense response to fungal pathogens: two types of protein kinases in the reversible phosphorylation of the host plasma membrane H+-ATPase, Plant Cell 8, 555–564
27 Roberts, D.M. and Harmon, A.C. (1992) Calcium-modulated proteins: targets of intracellular calcium signals in higher plants, Annu. Rev. Plant Physiol.
Plant Mol. Biol. 43, 375–414
28 Sheen, J. (1996) Ca2+-dependent protein kinase and stress signal transduction, Science 274, 1900–1902
29 Tavernier, E. et al. (1995) Involvement of free calcium in action of cryptogein, a proteinaceous elicitor of hypersensitive reaction in tobacco cells,
Plant Physiol. 109, 1025–1031
30 Xu, H. and Heath, M.C. (1998) Role of calcium in signal transduction during the hypersensitive response caused by basidiospore-derived infection of the cowpea rust fungus, Plant Cell 10, 585–597
31 Atkinson, M.M. and Baker, C.J. (1989) Role of the plasmalemma H+-ATPase
in Pseudomonas syringae-induced K+
/H+
exchange in suspension-cultured tobacco cells, Plant Physiol. 91, 298–303
32 De Wit, P.J.G.M. (1995) Fungal avirulence genes and plant resistance genes: unravelling the molecular basis of gene for gene interactions, Adv. Bot. Res. 21, 148–177
33 De Koninck, P. and Schulman, H. (1998) Sensitivity of CaM Kinase II to the frequency of Ca2+
oscillations, Science 279, 227–230
34 Allen, G.J. and Sanders, D. (1997) Vacuolar ion channels, Adv. Bot. Res. 25, 217–252
35 Mehdy, M.C. (1994) Active oxygen species in plant defense against pathogens, Plant Physiol. 105, 671–681
36 Segal, A.W. and Abo, A. (1993) The biochemical basis of the NADPH oxidase of phagocytes, Trends Biochem. Sci. 18, 43–47
37 Dwyer, S.C. et al. (1996) Plant and human neutrophil oxidative burst complexes contain immunologically related proteins, Biochim. Biophys. Acta 1289, 231–237
38 Groom, Q.J. et al. (1996) rbohA, a rice homologue of the mammalian gp91phox respiratory burst oxidase gene, Plant J. 10, 515–522
39 Keller, T. et al. (1998) A plant homolog of the neutrophil NADPH oxidase
gp91phoxsubunit gene encodes a plasma membrane protein with Ca2+binding
motifs, Plant Cell 10, 255–266
C
yanobacteria (blue-green algae) are a diverse group of pro-karyotes. A common feature is their oxygenic photosyn-thesis, which is similar to that in algae and higher plants and is the most important biological mechanism for capturing solar energy. As sunlight is their energy source and water the reductant, they generate oxygen in the light. Energy and reductant generated by photosynthesis are usually used for carbon dioxide reduction. Some strains are strict photoautotrophs, whereas others can use exogenous carbon sources such as fructose and glucose.Nitrogen fixation occurs only in prokaryotes and one line of evidence for the common origin of the nitrogen fixation mecha-nism is the similar physical, chemical and biological characteris-tics of the nitrogen-fixing enzyme system in otherwise dissimilar organisms. Many cyanobacteria are able to reduce atmospheric
dinitrogen to ammonia. In some filamentous cyanobacteria nitro-gen-fixing heterocysts are formed. Heterocysts are terminally dif-ferentiated cells whose interior becomes anaerobic, mainly as a consequence of respiration, allowing the oxygen-sensitive process of nitrogen fixation to continue. Heterocysts are spaced at semi-regular intervals along the filament with approximately 7% of the cells differentiating into heterocysts in free-living Anabaena/Nostoc species. The regulation of dinitrogen fixation has been extensively studied in the heterocyst system of diazotrophic cyanobacteria1
.
Nitrogen fixation and heterocyst formation
During differentiation of a vegetative cell into a heterocyst, major structural and biochemical changes occur that affect nitrogen fix-ation. Upon nitrogen deprivation phycobiliproteins are broken Eduardo Blumwald*, Gilad S. Aharon and Bernard C-H. Lam are at the Dept of Botany, University of Toronto, 25 Willcocks St, Toronto, Ontario, Canada M5S 3B2.
*Author for correspondence (tel +1 416 978 2378;
fax +1 416 978 5878; e-mail [email protected]).
Regulation of nitrogen fixation in
heterocyst-forming cyanobacteria
Herbert Böhme
down. At the same time, around the outer membrane of the hetero-cyst a double layered envelope is formed, which decreases the dif-fusion of oxygen. Connection to vegetative cells occurs through a pore, equipped with microplasmodesmata. Heterocysts import carbohydrates – these act as reductant and energy sources for nitrogen fixation – an in turn, export glutamine. Changes in the thylakoid structure of heterocysts are associated with a photosys-tem II that lacks oxygen evolving activity and Rubisco (the main enzyme complex responsible for CO2fixation). Therefore,
reduc-tant is almost exclusively channelled to the reduction of nitrogen to ammonia, which in turn reacts with glutamate derived from the imported carbohydrates. Both oxygen and nitrogen diffuse into the cells, but increased respiratory activity in membranes near to the polar ends of heterocysts depletes the oxygen concentration. Hydrogen produced by the nitrogenase reaction feeds into an up-take hydrogenase system, which is induced upon heterocyst for-mation. This reacts with oxygen to produce water, contributing to the ATP pool required for biosynthetic reactions such as nitrogen fixation1.
Requirements for dinitrogen fixation
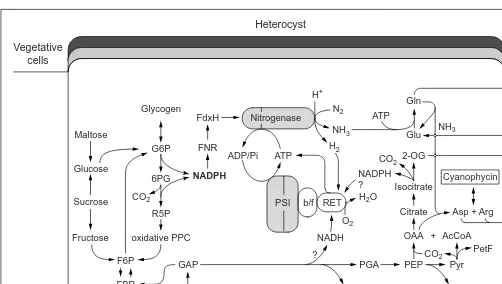
All diazotrophs, including nitrogen-fixing cyanobacteria, have the same general requirements for nitrogen fixation: a nitrogenase complex; ATP; a source of low-potential electrons and a partially anaerobic environment (Fig. 1). It has been shown that heterocysts contain the nitrogenase complex, which consists of an iron protein (dinitrogenase-reductase) and a iron-molybdenum protein (dini-trogenase, with the iron-molybdenum cofactor) the latter of which catalyzes nitrogen reduction1. Subsequent studies led to the cloning
and sequencing of nifH, nifD and nifK genes in Anabaena, which encode the structural genes of the nitrogenase complex. Sequence comparison revealed that the nitrogenase proteins of Anabaena were very similar to those of other diazotrophic bacteria2.
ATP formation in heterocysts is not completely understood. The rate of N2-fixation in the dark by heterocystous cyanobacteria is a
fraction of the rate in the light. ATP can be produced in the light by either cyclic photophosphorylation or oxidative phosphorylation, the latter process consumes oxygen and uses pyridine nucleotides or hydrogen as electron sources. Reduced pyridine nucleotides can
Fig. 1. Heterocyst metabolism and nitrogen fixation. The scheme of a heterocyst with adjacent vegetative cells is shown. The outer and inner layers of the heterocyst envelope consist of polysaccharides and glycolipids, respectively. In this scheme the pore region is not drawn to scale and shown enlarged to accommodate metabolite exchange between the cells. Cell wall and cell membranes are not drawn separately. Heterocysts import carbohydrates from vegetative cells, with glutamine moving in the opposite direction. In a cell-free system derived from heterocysts, the following substrates supported nitrogenase activity43
: glycogen, maltose, sucrose (less active), glucose and fructose; glucose 6-phosphate (G6P) and other intermediates of the oxidative pentose-phosphate cycle (PPC), including dihydroxyacetone phosphate (DAP), glyceraldehyde 3-phosphate (GAP) and fructose-1,6-bisphosphate (FBP), were particularly active. Glycolytic substrates, such as phospho-enolpyruvate (PEP) and pyruvate (Pyr) were inactive or inhibitory in acetylene reduction by the heterocyst extract. In the dark, reductant for nitrogen and oxygen is generated by the activity of the oxidative PPC and possibly by isocitrate dehydrogenase. NADPH thus formed donates electrons via ferredoxin:NADP reductase (FNR) to a heterocyst-specific ferredoxin (FdxH) and then to the two components of nitrogenase (Fe-protein and FeMo-protein) as indicated. NAD(P)H and hydrogen are also electron donors to the respiratory electron transport (RET) gen-erating the necessary ATP for the nitrogenase reaction. In the light, ATP is formed by cyclic photophosphorylation mediated by photosystem I (a PSI-dimer, as indicated). Ferredoxin could be also photoreduced by PSI at the expense of hydrogen and NAD(P)H as electron donors. Abbreviations: AcCoA, acetyl coenzyme A; Arg, arginine; Asp, aspartate; b/f, cytochrome b6f complex; F6P, fructose 6-phosphate; PetF,
vegetative cell type ferredoxin; Glu, glutamate; Gln, glutamine; OAA, oxaloacetate; 2-OG, 2-oxoglutarate; 6PG, 6-phosphogluconate; PGA, 3-phosphoglycerate; Pi, inorganic phosphate; R5P, ribose 5-phosphate.
Vegetative cells
Vegetative cells Heterocyst
Glycogen
Cyanophycin
Carbohydrates
G6P FNR
ADP/Pi ATP
PSI b/f RET H2O O2
NADH
?
?
ATP
6PG NADPH
Maltose
Glucose
R5P
oxidative PPC Fructose
F6P GAP PGA PEP
OAA
Pyr
DAP FBP
CO2
Sucrose
FdxH
H+
N2
ATP
NH3 NH3
H2
ATP AcCoA
PetF
PetF
+
Citrate Asp + Arg Isocitrate
NADPH 2-OG
Glu Gln
Gln + 2-OG
Glu +
2-OG
? Glu
CO2
CO2
be generated from the metabolism of imported carbohydrates1
. The nature of these carbohydrates is unknown, but a disaccharide such as sucrose might be involved. The putative disaccharide would be phosphorylated, converted to glucose-6-phosphate and then further degraded via enzymes of the oxidative pentose-phosphate cycle, which are particularly active in heterocysts, to produce NADPH and CO2(Ref. 1). It has been suggested that NADPH provides
elec-trons for nitrogenase reductase via a heterocyst-specific ferredoxin
(FdxH) and ferredoxin:NADP+
oxidoreductase (FNR) through re-versed electron transport. For this to occur, the ratio of NADPH to
NADP+
must be high, consistent with the levels measured in iso-lated heterocysts under nitrogen-fixing conditions3. In heterocysts,
expression of FNR is at an enhanced rate, about tenfold higher than in vegetative cells4. By the oxidative pentose-phosphate cycle,
glucose-6-phosphate can be almost completely oxidized to CO2,
yielding mainly NADPH for nitrogen reduction.
Inactivation of the zwf and opcA genes, encoding glucose-6-phosphate dehydrogenase (and a nearby gene of unknown function) has been shown to inactive nitrogen fixation in a mutant of Nostoc sp. ATCC 29133. This underlines the importance of this enzyme in the oxidative pentose-phosphate cycle of heterocysts in relation to nitrogen fixation5.
Because of the incomplete tricarboxylic acid cycle in cyano-bacteria, isocitrate dehydrogenase is the terminal step in carbon flow. This enzyme generates 2-oxoglutarate in an NADP-dependent re-action. The icd gene, encoding isocitrate dehydrogenase, shows a fivefold increase in transcription and enzymic activity under nitro-gen-fixing conditions. Upstream of the translation start region is a DNA binding site for NtcA, a global nitrogen regulator, which is re-quired for the expression of genes involved in nitrogen assimilation6.
From 2-oxoglutarate and glutamine, two molecules of glutamate are produced by glutamate synthase (GltS) in a ferredoxin-dependent reaction. Glutamine synthetase (GlnA) converts glutamate to gluta-mine using ammonia, generated upon nitrogen reduction, and ATP. The main route of nitrogen assimilation in heterocysts occurs via glutamine which is exported to vegetative cells1. In an in vitro assay,
isolated glutamate synthase (GltS) with heterocyst ferredoxin (FdxH) as electron source had only 10% of the electron transfer ac-tivity of vegetative cell ferredoxin (PetF). Flavodoxin, thought partly to replace the functions of PetF under iron limiting-conditions, was inactive in this reaction7
. Western blots of glutamate synthase indicated that this enzyme is only present at low concentrations in heterocysts. Furthermore, the enzyme only has very low activity in heterocysts, suggesting that glutamate is synthesized in veg-etative cells and exported to heterocysts for glutamine biosynthe-sis (F.J. Florencio, unpublished).
Glycogen granules found in cyanobacteria and especially in het-erocysts are used as carbohydrate reserves that can be phosphoryl-ated and broken down to glucose-6-phosphate as a substrate for the oxidative pentose-phosphate cycle. Another polymer is cyanophy-cin, consisting of asparagine and arginine; similar to phycobilipro-teins, it is a nitrogen reserve polymer that can be mobilized quickly1.
Genes involved in nitrogen fixation
Most of the genes involved in nitrogen fixation (nif genes), orig-inally described in Klebsiella, have also been detected in other diazo-trophic organisms. The structural genes of the nitrogenase complex,
nifH, nifD and nifK, represent one of the most highly conserved gene
groups in bacteria. On the basis of sequence similarity to
Kleb-siella DNA probes, nifH,D,K and nifS were cloned and mapped in Anabaena sp. PCC 7120 (Anabaena 7120). Subsequent sequence
analysis of adjacent genes and comparison to other nif genes from
Klebsiella, Azotobacter and Rhizobium species led to the current
picture of nif gene organization in Anabaena (Fig. 2)8.
Developmentally regulated genome rearrangements in heterocysts
Heterocyst differentiation in Anabaena 7120 is accompanied by de-velopmentally regulated genome rearrangements that affect fdxN,
nifD and hupL gene expression9
. These rearrangements of the veg-etative cell genome occur during late stages of differentiation at about the same time that the nitrogen fixation genes begin to be transcribed. All the DNA elements shown in Fig. 2a (fdxN is inter-rupted by a 55 kb DNA element, the nifD gene by an 11-kb el-ement and at an unknown distance to the nif-genes the hupL gene by 10.5 kb of chromosomal DNA) become excised upon hetero-cyst differentiation10. The corresponding genes encoding these site-specific recombinases are xisF, xisA and xisC, respectively, which are located within the excised DNA elements. Recently, it was demonstrated that xisF alone is not sufficient for the hetero-cyst specific excision of the fdxN element and xisH and xisI are also required11
. Anabaena 7120 mutants for xisA or xisF formed heterocysts but did not grow on nitrogen-free media12. The hupL-rearrangement in Anabaena 7120 was independently found by pulsed-field electrophoresis and by comparison of the restric-tion pattern of vegetative cell and heterocyst DNA (Ref. 13). The 10.5 kb hupL element is not present in Anabaena 29413 (T. Happe and H. Böhme, unpublished).
In contrast to Anabaena 7120, the closely related cyanobac-terium Anabaena 29413 does not contain the fdxN element in the
nif1 region; only the nifD element is present. In the nif2 region, the nifD element and the fdxN gene are absent (Fig. 2c). Both Pseudo-anabaena and Fischerella lack the 11-kb element1.
Functions of nif genes in Anabaena
The function of many nif genes in Anabaena has still not been determined, but a possible function can be inferred by analysing analogous genes described in other diazotrophic bacteria. The first operon on the left (Fig. 2a) includes the genes nifB, fdxN, nifS and
nifU, which are required for biosynthesis of the iron-molybdenum
(FeMo)- or the iron-vanadium (FeV) cofactor, but fdxN, nifS and
nifU of Anabaena 29413 were not essential for nitrogen fixation
to take place14,15. The glbN gene of Nostoc commune was discov-ered between nifS and nifU. It encodes cyanoglobin, the only known prokaryotic myoglobin that might scavenge for oxygen or act as a component of the membrane-associated, microaerobically induced terminal oxidase. Cyanoglobin was only detected in Nostoc, when, in addition to microaerobiosis, the cells were starved of nitrogen16
. The next operon to nifB,S,U consists of nifH, nifD and nifK (Fig. 2a). The nifH gene encodes the dinitrogenase reductase, a homodimer (2 330 kDa) with one [4Fe-4S]-cluster at the interface; nifD and
nifK encode the a- and b-subunits of dinitrogenase, respectively, an a2b2tetramer of 240 kDa associated with two FeMo-cofactors and two P-clusters. Because NifE and NifN show significant struc-tural similarity to NifD and NifK, respectively, it has been sug-gested that NifE and NifN generate the scaffold on which the FeMo-cofactor is assembled. NifE and NifN also form an a2b2 tetramer that binds the NifB cofactor, a small iron-sulfur-cluster protein and a precursor of the FeMo-cofactor. The precise func-tion of the nifX gene in cyanobacterial nitrogen fixafunc-tion remains to be determined.
The nifW gene is necessary for full stability or processing of the FeMo-protein. The functions of hesA and hesB are not known, although insertional inactivation of hesA impairs nitrogen fixation by approximately 55% (Ref. 17). The fdxH gene, which is tran-scribed late during heterocyst development together with the nitrogenase genes, encodes a unique [2Fe-2S]-ferredoxin, which is a specific electron donor for nitrogenase in vitro18
necessary for the magnitude of maximum nitrogenase activity and optimal growth under nitrogen-fixing conditions, but that fdxH is not essential for diazotrophic growth19.
The nifV, nifZ and nifT genes are separated from the main nif gene region in Anabaena 7120. The nifV gene encodes homo-citrate synthase and homohomo-citrate is an integral component of the FeMo- cofactor. The functions of nifZ and nifT are not clear; inac-tivation of nifV in Anabaena 7120 led to mutant strains that were still capable of diazotrophic growth (nitrogenase activity reduced by about 30–40%)20
.
The nifJ gene encoding a pyruvate:flavodoxin oxidoreductase, is not closely linked to other nif genes of Anabaena 7120. In
Kleb-siella, NifJ functions to degrade pyruvate and generate reduced
flavodoxin (NifF) as a specific electron donor to nitrogenase. An
Anabaena 7120 a nifJ mutant was unable to grow on medium
de-pleted of both iron and combined nitrogen. However, this strain was capable of diazotrophic growth when iron was present21. No
equivalent of the nifF gene has been found in Anabaena19
. Induction of the nitrogenase complex is accompanied by the induction of the hydrogen uptake system. The hupL gene encodes the large subunit of a membrane-bound [NiFe]-uptake hydrogen-ase and uses molecular hydrogen, a byproduct of nitrogenhydrogen-ase activity10. To improve the efficiency of nitrogen fixation,
hydro-gen becomes oxidized in a respiratory, ATP-forming reaction (Fig. 1).
Wolk and co-workers used transposon mutagenesis, based on a Tn5-derivative bearing luxA,B (encoding luciferase) of Vibrio
fis-cheri as a transcriptional reporter, to identify mutants that exhibit
enhanced luciferase activity after removal of ammonia from the medium. Visualization of gene activation in single cells was made possible using constructs in which the promoter region of PnifHDK and PrbcLSwas fused to luxA,B (Ref. 22). Among the first genes to be activated by nitrogen deprivation (within 0.5 h) were the
nirA-nrtA,B,C,D-narB genes of the nir operon, encoding the structural
genes for nitrite reductase (nirA), nitrate permease (nrtA,B,C,D) and nitrate reductase (narB). Anabaena strains carrying a mutation in
nirA, nrtC or nrtD remained competent to make heterocysts and
fix nitrogen23.
Alternative nitrogenase systems
Recently it was shown with Anabaena 29413 that in anaerobic conditions a second, Mo-dependent nitrogenase system (nif2) is expressed in all vegetative cells some hours after induction and long before heterocysts begin to develop. In contrast to the nif1 system of heterocysts, which functions under both anaerobic or external aerobic conditions and is developmentally regulated, the
nif2 system is expressed in all cells only under anaerobic
con-ditions and is regulated by environmental factors24,25. Anabaena
29413 has a very similar nif1 and nif2 gene arrangement (Fig. 2b,c)26. The environmentally regulated nif2 system lacks fdxN, but
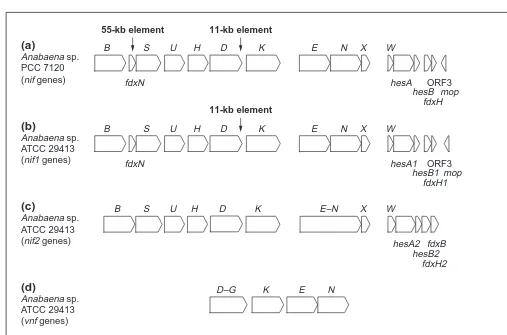
Fig. 2. Arrangement of nitrogen fixation genes from Anabaena ssp. and Klebsiella pneumoniae. The nif1 and nif2 systems of (a) Anabaena 7120 (nif genes), (b) Anabaena sp. ATCC 29413 (nif1 genes), (c) Anabaena 29413 (nif2 genes), and (d) the vnf system, encoding the alternative vanadium (V)-dependent nitrogenase. (e) The arrangement of genes involved in nitrogen fixation in Klebsiella pneumoniae. The major nif gene cluster of Anabaena 7120 heterocysts encompassing genes from nifB (left) to the mop gene (right) is separated from the nifVZT gene region. Vertical arrows indicate the positions of the 55-kb and the 11-kb DNA-elements of the vegetative cell genome of Anabaena 7120, which be-come excised during heterocyst differentiation. The nif2E-N and vnfD-G genes of Anabaena 29413 are fused into a single open reading frame27
.
(a)
Anabaena sp. PCC 7120 (nif genes)
55-kb element
V Z T W
N E K
D H U S
fdxN hesA ORF3
hesB mop fdxH
B X
11-kb element
(b)
Anabaena sp. ATCC 29413 (nif1 genes)
W N
E K
D H U S
fdxN hesA1 ORF3
hesB1 mop fdxH1
B X
11-kb element
(c)
Anabaena sp. ATCC 29413 (nif2 genes)
W E–N
K D H U S
hesA2 fdxB hesB2
fdxH2
B X
(d)
Anabaena sp. ATCC 29413 (vnf genes)
N E K D–G
(e)
Klebsiella pneumoniae (nif genes)
S V W Z M F L A B Q
N E K T Y D
H
contains the fdxB gene downstream of fdxH2. The fdxB gene en-codes a 2[4Fe-4S]-ferredoxin of unknown function24, and is
simi-lar to the corresponding gene from Rhodobacter.
In addition to the nif1 and nif2 genes, which encode nitrogenase-1 and -2, respectively, and which require the same FeMo-cofactor,
Anabaena 29413 also contains vnf genes encoding a V-dependent
nitrogenase27
, as found in some other diazotrophic organisms. The alternative, V-nitrogenase-encoding vnfD,G,K genes of Anabaena 29413 are organized much like those of Azotobacter spp. However, the gene for the dsubunit of the V-nitrogenase, vnfG, is fused to the
vnfD gene in Anabaena 29413. Two genes, vnfE and vnfN, which are
similar to vnfE,N genes of Azotobacter vinelandii were found down-stream from vnfD,G,K in Anabaena 29413 (Ref. 27). Insertional inactivation of the vnfN gene produced a mutant that grew poorly on a medium where vanadium replaced molybdenum (Ref. 28).
Genes involved in the regulation of nitrogen fixation Sigma factors
Upon deprivation of combined nitrogen, photosynthesizing veg-etative cells differentiate to form N2-fixing heterocysts. This requires
the coordinated regulation of many genes. These changes in gene expression involve modification of the transcription apparatus, although the nature of that modification remains unknown. In gen-eral, sigma factors play a major role in the progression of differ-entiation in prokaryotes. Sigma factors are modular components and can modify the major RNA polymerase to respond to nitrogen deficiency. In a search for similar factors in Anabaena, sigA, en-coding the major sigma factor in vegetative cells, and sigB and sigC, two nitrogen-regulated sigma factors, were isolated. However, in-activation of either sigB or sigC genes, which are expressed under nitrogen deficiency, still led to mutant strains capable of heterocyst differentiation and nitrogen fixation29. A new group 2 sigma-factor
gene, sigD, has been cloned recently. A sigD-minus mutant strain showed impaired diazotrophic growth and the appearance of hetero-cysts was delayed (I. Khudyakov and J.W. Golden, unpublished).
NtcA as a global nitrogen regulator
In cyanobacteria, ammonium exerts a negative control on proteins involved in the programme of assimilation of nitrogen from sources other than ammonium, such as nitrate and dinitrogen. NtcA is a glo-bal nitrogen regulator required for the activation of gene expression in response to removal of ammonia in diverse cyanobacteria. NtcA belongs to the family of bacterial transcriptional regulators, of which the cAMP receptor protein (Crp) of response regulators are the prototype. The amino acid sequence near the C-terminus of NtcA predicts a helix-turn-helix motif, characteristic of the formation of DNA–protein interactions30,31
. The bifA gene, discovered independ-ently as a DNA trans-acting factor from Anabaena 7120 on sites upstream of xisA, is identical to the ntcA gene32
. An ntcA mutant of
Anabaena 7120 failed to induce the nir operon and to express the
major glnA transcript induced under conditions of nitrogen deple-tion. In addition, the mutant did not develop heterocysts and was un-able to express nifH,D,K in response to nitrogen deprivation34–36
. In unicellular and filamentous strains, NtcA binds to target sequences
of glnA, nirA and ntcA, which have a palindromic GTA(N8)TAC
motif (where N is any nucleotide) upstream of the transcription start site. This sequence replaces the -35 promoter site of E. coli (Ref. 33). Using similar experiments, an alternative but overlapping BifA (= NtcA) binding site has been proposed: TGT(N9–10)ACA (Ref. 31).
In the unicellular diazotrophic cyanobacterium, Cyanothece sp.,
ntcA transcripts were weakly expressed during N2-fixation, but
expression increased in nitrate-grown and especially ammonium-grown cells. According to these data NtcA seems to be more im-portant for nitrogen assimilation than nitrogen fixation35.
GlnB (PII-protein) as sensor kinase of NtcA?
In enterobacteria, regulation of nitrogen metabolism is mediated by a two-component ntr-system (nitrogen regulation) in which the GlnB protein has the role of transmitting the nitrogen-status of the cell to Ntr-proteins. The GlnB protein controls both the activity and synthesis of glutamine synthetase (GlnA), a key enzyme in bacterial nitrogen assimilation. A cyanobacterial glnB gene has been isolated that is very similar to its bacterial counterpart36. In
Synechococcus sp. PCC 7942, a unicellular, non-nitrogen-fixing
cyanobacterium, the homotrimeric PII-protein is modified by
serine-phosphorylation. In the presence of ammonium, PIIis found in its
dephosphorylated state. A kinase- or phosphatase-activity can be separated by biochemical methods. The kinase activity depends on ATP as a phosphoryl donor and the presence of 2-oxoglutarate, as carbon skeleton required for nitrogen assimilation, to sense the nitrogen status of the cell37,38. ATP and 2-oxoglutarate were bound
by the PII-protein in a mutually dependent manner. Glutamine had
no effect on kinase or phosphatase activities. By studying inser-tional mutants of Synechococcus 7942 lacking the PII-protein and
mutants, where at the phosphorylation site serine was exchanged for alanine showed that in the presence of a dephosphorylated form of PIInitrate and nitrite transport was inhibited. However, a
pleio-tropic PII-deficient mutant suggested that PIIis not essential for
acti-vation of NtcA-dependent transcription39. In Synechocystis 6803,
a unicellular, non-nitrogen-fixing cyanobacterium, glnB expres-sion was specifically activated (tenfold) under nitrogen depri-vation. Induction of expression of the glnB gene might be under the control of NtcA. Constitutive levels of GlnB were detected from a
s70
-dependent E. coli-like promoter. This would ensure basal levels of the PII-protein were available to sense changes in
environmen-tal conditions at any time. Preliminary results indicate a correlation between PIIstate and GlnA activity
40. In the filamentous,
nitrogen-fixing cyanobacterium Nostoc 29133, glnB could not be in-sertionally inactivated41. In the nitrogen-fixing Calothrix 7504,
heterocyst differentiation correlated with the modified form of PII
(Ref. 42). More experiments concerning nitrogen fixation in cyanobacteria are necessary to clarify the relation between NtcA and PII.
Conclusions and future prospects
Although we have a fairly sophisticated level of understanding of nitrogen fixation in cyanobacteria there are still a number of uncertain aspects. For example, because photosynthetic oxygen evolution is absent from heterocysts, reductant must be provided by adjoining vegetative cells. However, the molecules that are transported into heterocysts to provide reductant and carbon skel-etons for fixed nitrogen are not known with any certainty. Another important question is the identity of the permeases that are present between heterocysts and vegetative cells and that participate in this process.
An oxygen sensor responsible for the regulated expression nif-genes has also yet to be described in cyanobacteria. The different pathways of electron-donation to nitrogenase are also not clearly known – an fdxH-minus strain was impaired in nitrogen fixation, but not completely inhibited. Finally, the nif-genes of Anabaena 7120 are interrupted and removed from the chromosome during heterocyst differentiation, but this process is not ubiquitous in cyanobacteria, and there is as yet no explanation for it. Clearly, there is more to be done to unravel the mysteries of nitrogen fixation in cyanobacteria.
Acknowledgements
References
1 Wolk, C.P., Ernst, A. and Elhai, J. (1994) Heterocyst metabolism and
development, in The Molecular Biology of Cyanobacteria (Bryant, D.A., ed.), pp. 769–823, Kluwer
2 Haselkorn, R. and Buikema, W.J. (1992) Nitrogen fixation in cyanobacteria, in
Biological Nitrogen Fixation (Stacey, G., Burris, R.H. and Evans, H.J., eds),
pp. 166–190, Chapman & Hall
3 Böhme, H. (1987) Regulation of electron flow to nitrogenase in a cell-free system
from heterocysts of Anabaena variabilis, Biochim. Biophys. Acta 891, 121–128
4 Razquin, P. et al. (1996) Expression of ferredoxin-NADP+reductase in
heterocysts from Anabaena sp., Biochem. J. 316, 157–160
5 Summers, M.L. et al. (1995) Genetic evidence of a major role for
glucose-6-phosphate dehydrogenase in nitrogen fixation and dark growth of the cyanobacterium Nostoc sp. strain ATCC 29133, J. Bacteriol. 177, 6184–6194
6 Muro-Pastor, M.S. and Florencio, F.J. (1994) NADP+
-isocitrate dehydrogenase from the cyanobacterium Anabaena sp. strain PCC 7120: purification and characterization of the enzyme and cloning, sequencing, and disruption of the icd gene, J. Bacteriol. 176, 2718–2726
7 Schmitz, S. et al. (1996) Glutamate 94 of [2Fe-2S]-ferredoxin is important for
efficient electron transfer in the 1:1 complex formed with ferredoxin-glutamate synthase (GltS) from Synechocystis sp. PCC 6803, Biochim.
Biophys. Acta 1277, 135–140
8 Haselkorn, R. et al. (1991) Nitrogen fixation in filamentous cyanobacteria, in
Nitrogen Fixation (Polsinelli, M., Materassi, R. and Vincenzini, M., eds),
pp. 359–365, Kluwer
9 Buikema, W.J. and Haselkorn, R. (1993) Molecular genetics of cyanobacterial
development, Annu. Rev. Plant Physiol. Plant Mol. Biol. 44, 33–52
10 Carrasco, C.D., Buettner, J.A. and Golden, J.W. (1995) Programmed DNA
rearrangement of a cyanobacterial hupL gene in heterocysts, Proc. Natl. Acad.
Sci. U. S. A. 92, 791–795
11 Ramaswamy, K.S. et al. (1997) Cell-type specificity of the Anabaena
fdxN-element rearrangement requires xisH and xisI, Mol. Microbiol. 23, 1241–1249
12 Carrasco, C.D. et al. (1994) Anabaena xisF gene encodes a developmentally
regulated site-specific recombinase, Genes Dev. 8, 74–83
13 Matveyev, A.V. et al. (1994) A novel genome rearrangement involved in
heterocyst differentiation of the cyanobacterium Anabaena sp. PCC 7120,
FEMS Microbiol. Lett. 116, 201–208
14 Lyons, E.M. and Thiel, T. (1995) Characterization of nifB, nifS, and nifU
genes in the cyanobacterium Anabaena variabilis: NifB is required for the vanadium-dependent nitrogenase, J. Bacteriol. 177, 1570–1575
15 Masepohl, B. et al. (1997) The ferredoxin-encoding fdxN gene of the
filamentous cyanobacterium Anabaena variabilis ATCC 29413 is not essential for nitrogen fixation, New Phytol. 136, 419–423
16 Potts, M. et al. (1992) Myoglobin in a cyanobacterium, Science 256, 1690–1692 17 Borthakur, D. et al. (1990) Expression, nucleotide sequence and mutational
analysis of two open reading frames in the nif gene region of Anabaena sp. strain PCC 7120, Mol. Gen. Genet. 221, 227–234
18 Böhme, H. and Haselkorn, R. (1988) Molecular cloning and nucleotide
sequence analysis of a gene coding for heterocyst ferredoxin from the cyano-bacterium Anabaena sp. strain PCC 7120, Mol. Gen. Genet. 214, 278–285
19 Masepohl, B. et al. (1997) The heterocyst-specific fdxH gene product of the
cyanobacterium Anabaena sp. PCC 7120 is important but not essential for nitrogen fixation, Mol. Gen. Genet. 253, 770–776
20 Stricker, O. et al. (1997) Identification and characterization of the
nifV-nifZ-nifT-gene region from the filamentous cyanobacterium Anabaena sp.
PCC 7120, J. Bacteriol. 179, 2930–2937
21 Bauer, C., Scappino, L. and Haselkorn, R. (1993) Growth of the
cyanobacterium Anabaena on molecular nitrogen: NifJ is required when iron is limited, Proc. Natl. Acad. Sci. U. S. A. 90, 8812–8816
22 Elhai, J. and Wolk, C.P. (1990) Developmental regulation and spatial pattern
of expression of structural genes for nitrogenase in the cyanobacterium
Anabaena, EMBO J. 9, 3379–3388
23 Cai, J. and Wolk, C.P. (1997) Nitrogen deprivation of Anabaena sp. strain
PCC 7120 elicits rapid activation of a gene cluster that is essential for uptake
and utilization of nitrate, J. Bacteriol. 179, 258–266
24 Schrautemeier, B., Neveling, U. and Schmitz, S. (1995) Distinct and
differently regulated Mo-dependent nitrogen-fixing systems evolved for heterocysts and vegetative cells of Anabaena variabilis ATCC 29413: characterization of the fdxH1/2 gene regions as part of the nif1/2 gene clusters,
Mol. Microbiol. 18, 357–369
25 Thiel, T. et al. (1995) A second nitrogenase in vegetative cells of a
heterocyst-forming cyanobacterium, Proc. Natl. Acad. Sci. U. S. A. 92, 9358–9362
26 Thiel, T., Lyons, E.M. and Erker, J.C. (1997) Characterization of genes for a
second Mo-dependent nitrogenase in the cyanobacterium Anabaena variabilis,
J. Bacteriol. 179, 5222–5225
27 Thiel, T. (1993) Characterization of genes for an alternative nitrogenase in the
cyanobacterium Anabaena variabilis, J. Bacteriol. 175, 6276–6286
28 Thiel, T. (1996) Isolation and characterization of the nifEN genes of the
cyanobacterium Anabaena variabilis, J. Bacteriol. 178, 4493–4499
29 Brahamsha, B. and Haselkorn, R. (1992) Identification of multiple RNA
polymerase sigma factor homologs in the cyanobacterium Anabaena sp. strain PCC 7120: cloning, expression, and inactivation of sigB and sigC genes,
J. Bacteriol. 174, 7273–7282
30 Luque, I., Flores, E. and Herrero, A. (1994) Molecular mechanism for the
operation of nitrogen control in cyanobacteria, EMBO J. 13, 2862–2869
31 Ramasubramanian, T.S., Wei, T.F. and Golden, J.W. (1994) Two Anabaena
sp. strain PCC 7120 DNA-binding factors interact with vegetative cell- and heterocyst-specific genes, J. Bacteriol. 176, 1214–1223
32 Wei, T.F., Ramasubramanian, T.S. and Golden, J.W. (1994) Anabaena sp.
strain PCC 7120 ntcA gene required for growth on nitrate and heterocyst development, J. Bacteriol. 176, 4473–4482
33 Frías, J.E., Flores, E. and Herrero, A. (1994) Requirement of the regulatory
protein NtcA for the expression of nitrogen assimilation and heterocyst development genes in the cyanobacterium Anabaena sp. PCC 7120, Mol.
Microbiol. 14, 823–832
34 Frías, J.E., Flores, E. and Herrero, A. (1997) Nitrate assimilation gene cluster
from the heterocyst-forming cyanobacterium Anabanena sp. strain PC 7120,
J. Bacteriol. 179, 477–486
35 Bradley, R.L. and Reddy, K.J. (1997) Cloning, sequencing, and regulation of
the global nitrogen regulator gene ntcA in the unicellular diazotrophic cyanobacterium Cyanothece sp. strain BH68K, J. Bacteriol. 179, 4407–4410
36 Tsinoremas, N.F. et al. (1991) Photosynthetic electron transport controls nitrogen
assimilation in cyanobacteria by means of posttranslational modification of the
glnB gene product, Proc. Natl. Acad. Sci. U. S. A. 88, 4565–4569
37 Forchhammer, K. and Tandeau de Marsac, N. (1994) The PIIprotein in the
cyanobacterium Synechococcus sp. strain PCC 7942 is modified by serine phosphorylation and signals the cellular N-status, J. Bacteriol. 176, 84–91
38 Forchhammer, K. and Hedler, A. (1997) Phosphoprotein PIIfrom
cyanobacteria. Analysis of functional conservation with the PII
signal-transduction protein from Escherichia coli, Eur. J. Biochem. 244, 869–875
39 Lee, H.M. et al. (1998) A role for the signal transduction protein PIIin the
control of nitrate/nitrite uptake in a cyanobacterium, FEBS Lett. 427, 291–295
40 García-Domínguez, M. and Florencio, F.J. (1997) Nitrogen availability and
electron transport control the expression of glnB gene (encoding PIIprotein) in
the cyanobacterium Synechocystis sp. PCC 6803, Plant Mol. Biol. 35, 723–734
41 Hanson, T.E. et al. (1998) Characterization of the glnB gene product of Nostoc
punctiforme strain ATCC 29133: glnB or the PIIprotein may be essential,
Microbiology 144, 1537–1547
42 Liotenberg, S. et al. (1996) Modification of the PIIprotein in response to
carbon and nitrogen availability in filamentous heterocystous cyanobacteria,
FEMS Microbiol. Lett. 144, 185–190
43 Böhme, H. and Schrautemeier, B. (1987) Electron donation to nitrogenase in a
cell-free system from heterocysts of Anabaena variabilis, Biochim. Biophys.
Acta 891, 115–120