SELECTION AND CHARACTERIZATION OF
SIDEROPHORE-PRODUCING RHIZOBACTERIA AND
POTENTIAL ANTAGONISTIC ACTIVITY
TOWARD
Ralstonia solanacearum
ABDJAD ASIH NAWANGSIH , IDA PARIDA , SURYO WIYONO and 1* 1 1 JUANG GEMA KARTIKA2
1Department of Plant Protection, Faculty of Agriculture, Institut Pertanian Bogor, Bogor 16680, Indonesia 2Department of Agronomy and Horticulture, Faculty of Agriculture, Institut Pertanian Bogor, Bogor 16680, Indonesia
Received 24 June 2014/Accepted 9 November 2016
ABSTRACT
Ralstonia solanacearum is an important disease of tomato. An alternative method to control the disease is the application of biocontrol agents. Plant Growth-Promoting Rhizobacteria (PGPR) could be used as potential biocontrol agents. PGPR with siderophores is among compounds having important role in disease suppression. This experiment was conducted to select and characterize the siderophore-producing rhizobacteria from tomato and to determine their potential as antagonistic agents for R. solanacearum. Candidates of the PGPR were isolated from tomato grown in West Java Province, Indonesia. The isolates were detected as siderophore-producing bacteria using CAS medium. Among 29 isolates producing siderophore and having negative result on hypersensitivity reaction, two isolates provided the widest diameter of inhibition zone toward R. solanacearum. Both isolates were CP1C and CP2D with diameter of inhibition zone up to 3.6 and 7.0 mm, respectively. Based on the sequence of 16S rDNA, isolate CP1C was identified as Brevundimonas sp., while isolate CP2D was identified as Enterobacter sp. Both bacteria did not cause negative effect on the increasing plant height and dry weight of the plants, compared with control.
Keywords: Bacterial wilt, biocontrol, Brevundimonas, Enterobacter, PGPR
* Corresponding author: [email protected]
INTRODUCTION
Bacterial wilt of tomato caused by Ralstonia solanacearum is one of important diseases of tomato in tropics and subtropics area (Jeung et al.
2007). The bacteria survive for a long time in the soil (Wang & Lin 2005). The bacteria also multiply in the xylem as well as attacking the xylem and affect the water and nutrient translocation, causing wilting and death of the plant (Agrios 2005). Among the control methods to control the disease, is the application of biological control (biocontrol) agents (Yuliar et al. 2015). According to Jenifer et al. (2013), one of the most important mechanisms responsible for suppressing
Pseudomonas sp. (a plant pathogen) is siderophore-mediated competitions for iron.
Siderophores (from the Greek “iron carriers”) are small ferric-ion-specific chelating agents produced by bacteria and fungi which grow in low iron condition causing them to scavenge iron from the environment and to make iron available to the microbial cell (Neilands 1995; Pal & Gokarn 2010). Siderophores are also known to bind molybdenum and lead (Pal & Gokarn 2010).
Sayyed et al. (2005) reported that siderophore-producing Pseudomonas sp. plays vital role in stimulating plant growth and in controlling several plant diseases. In Pseudomonas fluorescens
the pigment produced is a siderophore. Pigment production is increased in the presence of sodium, potassium, lead, molybdenum, cadmium and ammonium sulphate ((NH ) SO ) 4 2 4
Java Province, Indonesia. From each village, 1,500 g of rhizosphere soil samples (with tomato roots) were collected from five different plots (300 g per plot as subsample) and thoroughly mixed using trowel, until becoming a composite sample. From each composite sample, 10 g of rhizosphere soil was immersed in 90 mL 0.85% NaCl. After a serial dilution, 0.1 mL of each dilution was inoculated on Chrome Azurol Sulphate (CAS) medium (Gross 1990; Louden et al. 2011). Inoculation was repeated three times. Plates containing the inoculation were incubated at room temperature (±28 C). Numbers of o
colony of siderophore-producing rhizobacteria were calculated at 24 - 48 hours after incubation, based on the production of orange zone around colony of the bacteria. Successfully isolated bacteria were transferred to King's B agar medium and separated from each other to obtain pure culture. Each isolate was preserved in Nutrient Broth (NB) with 20% glycerol and kept in -20 C environment. For daily maintenance, o
isolates were preserved in NB and kept at room temperature (±28 C)o .
Hypersensitive Reaction Test
Hypersensitive Reaction (HR) test was conducted to select the nonpathogenic bacteria as candidates of biocontrol agents. The test was
8 9
conducted by inoculating 10 – 10 cfu/mL of rhizobacteria on tobacco leaves. One milliliter of suspension was injected into tobacco leaf using 5 mL sterile syringe. Successful injection was indicated by water soaked zone around the point of injection. Inoculation of each isolate was conducted in duplo. Inoculated leaves were
o
incubated at room temperature (±28 C) for 24 hours. Leaves showing necrotic symptoms before 24 hours were counted as having positive reaction to the hypersensitive reaction and thus, be eliminated.
Antagonistic Activities of Siderophore-producing Rhizobacteria against In vitro Ralstonia solanacearum
Bacteria isolates having negative reaction on the previous hypersensitive test were tested on their antagonistic activity toward R. solanacearum. Both bacteria, i.e. R. solanacearum and rhizobacteria, were grown on a King's B agar 2011). Sayyed et al. (2005) also reported that Mn,
Hg and Co showed inhibitory effect on siderophores growth and production. Presence of potassium, magnesium and calcium had little inhibitory effect on siderophores production compared to controls (Battacharya 2010).
Sayyed et al. (2005) reported that production of siderophores by bacteria was affected by growing media. In growing media such as Nutrient Broth (NB) and MacKonkey's Broth (MB), siderophores were not produced because both media are luxurious media having high content of Fe. Fe-free succinic acid medium (SM) was found to provide maximum (92.25%) siderophores production in comparison to 88.00% in Barbhayya Rao broth (BR), 79.00% in Cassamino Acid broth (CAA) and 48.00% in Enrichment Medium (EM) (Sayyed et al. 2010). A study on pH effect to the production of siderophores by
Rhizobium sp. showed that the growth and production of siderophores started at pH 4.5, reaching maximum at neutral pH; at pH 10.0, there was no siderophores production (Sridevi et al. 2008). Study on genetic diversity of tobacco rhizosphere which produces siderophore demonstrated that 85% of the total 354 isolates produced siderophores in iron limited liquid medium; some of them are Pseudomonas,
Enterobacter Serratia Pantoea Erwinia , , , and
Stenotrophomonas which belong to γ-Proteobacteria
(Tian et al. 2009).
This experiment was conducted to select and characterize the siderophore-producing rhizobacteria from tomato and to determine their potential as antagonistic agents for R. solanacearum.
MATERIALS AND METHODS Isolation and Quantification of Siderophore-producing Rhizobacteria
plate. Five hundred micro liter suspension of R. solanacearum (10 - 10 cfu/mL) was spread on the 8 9
surface of the King's B agar plate. After being air dried, three sterilized filter papers having 0.5 cm diameter were placed besides the agar plate with 2 cm distance from each other. Filter paper in the center was inoculated with 50 µL of sterilized distilled water and designated as control. The two other filter papers were each inoculated with 50 µL suspension of one isolate of rhizobacteria. Each treatment was repeated three times. Inoculated plates were incubated at room temperature (±28 C). The antagonistic activity o
was observed at 24 hours after incubation. Antagonistic activity was indicated by the production of inhibition zone around the filter paper inoculated with siderophore-producing rhizobacteria.
E f f e c t o f S i d e r o p h o r e - p r o d u c i n g Rhizobacteria on the Viability of Tomato Seeds
Tomato seeds (Arthaloka and Ratna varieties) were dipped in 10 - 10 cfu/mL suspension of 7 8
siderophore-producing rhizobacteria for 16 hours before being planted on sterilized mixture of soil and compost (1 : 1 ratio). The soil mixture was put in 30 - 50 cm polyethylene pot tray having 128 holes. The experiment was arranged as completely randomized factorial design with 7 isolates of biocontrol agents and one control (without bacteria) as the first factor and two tomato varieties (Arthaloka and Ratna) as the second factor. Thus, 16 treatments were applied in this experiment. Each treatment was replicated three times. There were 48 units in total; each unit
contained 15 seeds. One tomato seed was grown in one hole of polyethylene pot tray. The isolates of biocontrol agents used in this experiment were CP1C (code of one isolate of bacteria), CP2B, CP2D, CP3E, LB1A, LB1C and LB1L. Seeds dipped in sterilized distilled water were used as control. Layout of the treatments was shown in Table 1.
Total of the emerging seedlings were calculated every day. Seed viability (SV) was calculated using the following formula:
Total normal seedlings
SV (%) = x 100% Total seeds sown
E f f e c t o f S i d e r o p h o r e - p r o d u c i n g Rhizobacteria on the Height, Fresh and Dry Weight of Tomato Plants
An experiment to test the effect of siderophore-producing rhizobacteria toward the height, fresh and dry weight of tomato plants was conducted in a green house. Tomato seeds of Arthaloka and Ratna varieties were dipped in suspension of siderophore-producing rhizobacteria (10 - 10 cfu/mL) for 16 hours 7 8
before being planted in 10 x 15 cm polybag filled with 2.5 – 3 kg of sterilized mixture of soil and compost (1 : 1 ratio). The experimental design applied was completely randomized factorial design having similar layout with the one presented in Table 1. The only difference was that each unit contained 5 seeds. Plant height was measured every 5 days starting from the day when the first leaf was fully opened. Total area under height of plant growth curve (AUHPGC) was calculated using
Table 1 Layout of experiment to test the effect of siderophore-producing rhizobacteria toward the viability of tomato seeds
Tomato Varieties Isolate’s code
CP1C CP2B CP2D CP3E LB1A LB1C LB1L Control
Arthaloka Repl 1Repl 2 15 seeds15 seeds 15 seeds15 seeds 15 seeds15 seeds 15 seeds15 seeds 15 seeds15 seeds 15 seeds15 seeds 15 seeds15 seeds 15 seeds15 seeds
Repl 3 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds
Ratna
Repl 1 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds
Repl 2 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds 15 seeds
modification of formula reported by Van der Plank (1963):
where:
y = increasing of plant height at the next i+1
observation
y = increasing of plant height at the time of i
observation
t = the next observation (II, III, …, VI) i+1
t = time of observation (I, II, …, V)i
Two months after planting, all plants (five plants) from each replication of each treatment were rooted and weighted using digital balance. The average of the five plants represented data of fresh weight for each replication. Dry weight of plant was determined by drying the fresh plants in the oven at 100 C. The weight of the sample was o
checked periodically until the weight of the sample was constant.
Characterization and Identification of the Siderophore-producing Rhizobacteria
Seven isolates of the siderophore-producing rhizobacteria were characterized based on microscopic and colony appearances, as well as on physiological and biochemical properties, following the methods of Klement et al. (1990) and Schaad et al. (2001). Two isolates of the siderophore-producing bacteria having potential as biocontrol agents were genetically identified by sequencing the 16S rDNA gene. DNA was extracted from log phase culture using phenolchloroform extraction procedure (Sambrook & Russel 2001). The 16S rDNA gene was amplified using universal primer for prokaryotes which were the forward primer 27F (5'-AGAGTTTGATCCTGGCTCAG-3') and reverse primer 1492R (5'GGTTACCTTACG-ACTT-3'). Total volume reaction for Polymerase Chain Reaction (PCR) was 25 µL consisted of 1 µL of DNA template; 12.5 µL of 1x Ready mix PCR, 1.5 mM MgCl , 0.2 mM dNTPs and Taq 2
polymerase 5 units/reaction; 9.5 µL of ddH O; 1 2
µL of 120 pmol Primer 27F; and 1 µL of 120 pmol Primer 1492R. PCR was performed under the following conditions: one cycle of
pre-o
denaturation at 95 C for 5 minutes, followed by 35
o
cycles of denaturation at 95 C for 1 minute,
o
annealing at 55 C for 1 minute and extension at
o
72 C for 2 minutes. The reaction was terminated
o
with a final extension at 72 C for 10 minutes. The PCR products were sent to the First Base Laboratory, Malaysia for sequencing. BLAST searches were performed for sequences obtained to find the similarity with sequence data in GeneBank.
Data Analysis
Data of the bacterial population was analyzed using t test of Minitab program version 13.3. Effects of the bacteria to the plant growth were statistically analyzed using ANOVA for completely randomized factorial design with isolates of biocontrol agents as the first factor and tomato varieties as the second factor. Treatment means were compared using the DMRT test at 5% level of significance. SAS Program version 9.1 was used for performing statistical analyses for completely randomized factorial design and for the DMRT test.
RESULTS AND DISCUSSION
Abundance and Antagonistic Activities of Siderophore-producing Bacteria
Based on the t test, the average of siderophore-producing bacteria isolated from Cipanas District
7
was 1.98 10 cfu/g, which was not significantly different with those from Lembang District
7
having average of 5.3 x 10 cfu/g. Colonies of the siderophore-producing bacteria on CAS medium were shown in Figure 1. Siderophore production was indicated by the orange color around the colony of the bacteria. Figure 1 shows that each colony produced different amount of siderophores, indicated by the diameter of orange area around each colony of bacteria. Among the 60 isolates of siderophore-producing rhizobacteria, 31 isolates positively showed hypersensitive reaction on tobacco, while 29 others showed negative reaction. Bacteria were also tested further for their antagonistic activities against Ralstonia solanacearum. Based on the antagonistic test, 16 isolates positively produced inhibition zone (Fig. 2) having diameter between 0.5 to 7.0 mm. Among those 16 isolates, 9 isolates were HR positive which had to be eliminated from being candidate of biocontrol agents. The
AUHPGC
n – 1
y y i i+ +1
2 t i+1 – ti
other 7 isolates were HR negative, having respective diameter of the inhibition zone of CP1C (3.6 mm), CP2B (2.3 mm), CP2D (7.0 mm), CP3E (5.0 mm), LB1A (1.6 mm), LB1C (1.6 mm) and LB1L (0.5 mm). The widest diameter of inhibition zone was produced by isolate CP2D which was isolated from Cipanas (Table 2).
Microorganisms growing under aerobic conditions need iron for a variety of functions, including reduction of oxygen for ATP synthesis, reduction of ribotide precursors of DNA, for formation of heme and for other essential purposes. A level of at least one micromolar iron is needed for optimum growth (Neilands 1995).
Figure 1 Production of siderophores by tomato rhizobacteria on CAS agar was indicated by yellow-orange color around colony of bacteria
Figure 2 Production of inhibition zone (arrow sign) by the isolate of siderophore-producing rhizobacteria (Note: The inset shows magnification of the inhibition zone)
Table 2 Isolates of siderophore-producing rhizobacteria which produced inhibition zone against R. solanacearum on Kings’s B agar
Isolate code1) inhibition zone Diameter of
(mm)
Isolate
code inhibition zone (mm)Diameterof Isolatecode
Diameter of inhibition zone
(mm)
CP1B2)
CP1C CP2B
CP2C
CP2D
CP2H
6.8 3.6 2.3 0.6 7.0 4.5
CP2L CP2S CP3E
CP3M CP3T LB1A
1.6 2.6 5.0 2.2 4.0 1.6
LB1C
LB1D
LB1E
LB1L
1.6 0.5 0.5 0.5
Note: 1) CP = isolates from Cipanas; LB = isolates from Lembang
Table 3 Effect of siderophore-producing rhizobacteria on tomato seed viability of Arthaloka and Ratna varieties
Treatment Seed viability (%)*)
Var. Arthaloka
Note: *) Means in the same column followed by the same letter are not significantly different according to Duncan Multiple Range Test (p < 0.05)
Table 4 Effect of siderophore-producing rhizobacteria on the increase of tomato plant height of Arthaloka and Ratna varieties
Treatment Plant height increase (cm)
1)
Note: 1) Means in the same column followed by the same letter are not significantly different according to Duncan Multiple Range Test (p < 0.05)
2) DAP = Days After Planting
3) AUHPGC = Area Under Height of Plant Growth Curve
Seven isolates of the bacteria were tested for their effects on seed viability, plant height and the fresh and dry weight of two varieties of tomato plants, i.e. Arthaloka and Ratna, as presented in Table 3, 4, and 5, respectively. Data in Table 3, 4, and 5 show not only that the isolates of bacteria did not significantly increase
Table 6 Morphological characteristic of colony of the seven isolates of siderophore-producing rhizobacteria on King's B Agar
Isolate code1) Colony characteristic
Diameter Color Elevation Edge Form
CP1C
Note: 1) CP = isolates from Cipanas; LB = isolates from Lembang
Table 7 Physiological and biochemistry characteristics of siderophore-producing rhizobacteria having potential as antagonist for R. solanacearum
Isolate code1) Fluorescence Gram reaction Phosphate
solubilization Resistance to 80 oC
CP1C - - + +
Note: 1) CP = isolates from Cipanas; LB = isolates from Lembang
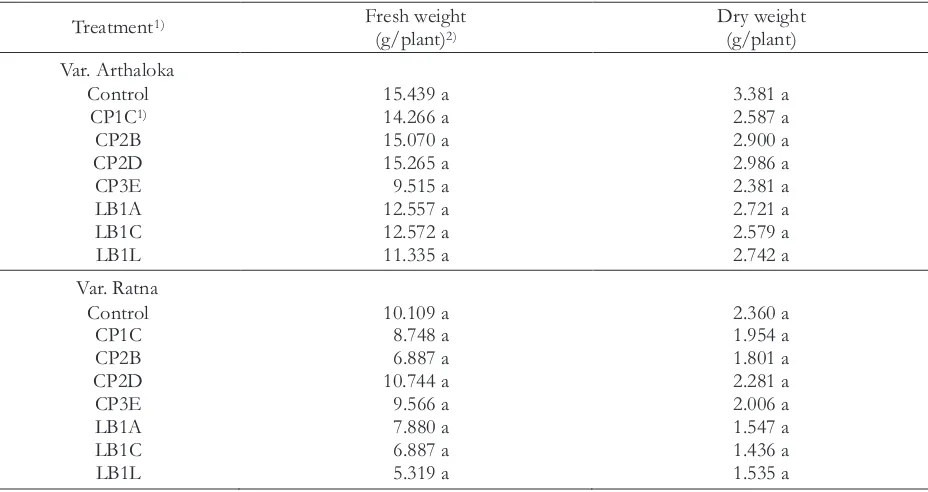
Table 5 Effect of siderophore-producing rhizobacteria on fresh and dry weight of tomato plants
Treatment1) Fresh weight
(g/plant)2) Dry weight (g/plant) Note: 1) CP = isolates from Cipanas; LB = isolates from Lembang
2) Means in the same column followed by the same letter are not significantly different according to Duncan Multiple Range Test (p < 0.05)
Based on the inhibition zone production, effect on seed viability, effect on plant height increase, and effect on fresh and dry weight of tomato product, two isolates of siderophore-producing bacteria having the best effects were
selected. The two isolates were CP1C and CP2D. The sequence of 16S rDNA of those isolates were referred to the gene bank. Using the BLAST program the isolate of CP1C was identified as
·
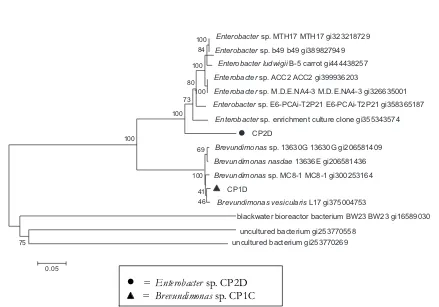
= Enterobacter sp. CP2D= Brevundimonas sp. CP1C
Figure 3 Phylogenetic tree of isolates Enterobacter sp. CP2D and Brevundimonas sp. CP1C
Enterobacter sp. MTH17 MTH17 gi323218729
Enterobacter sp. b49 b49 gi389827949
Enterobacter ludwigii B-5 carrot gi444438257
Enterobacter sp. ACC2 ACC2 gi399936203
Enterobacter sp. M.D.E.NA4-3 M.D.E.NA4-3 gi326635001
Enterobacter sp. E6-PCAi-T2P21 E6-PCAi-T2P21 gi358365187
Enterobacter sp. enrichment culture clone gi355343574 CP2D
Brevundimonas sp. 13630G 13630G gi206581409
Brevundimonas nasdae 13636E gi206581436
Brevundimonas sp. MC8-1 MC8-1 gi300253164 CP1D
Brevundimonas vesicularis L17 gi375004753
blackwater bioreactor bacterium BW23 BW23 gi16589030 uncultured bacterium gi253770558
uncultured bacterium gi253770269
100 84 100
100 80
100 73
100
69
100 41 46
75
0.05
Table 8 Maximum score, E value and percentage of similarities of siderophore-producing bacteria
Isolate Species homolog Identity Max score Query cover E value Accession number
CP1C Brevundimonas sp. 13630 G 16S ribosomal RNA gene, partial sequence
95% 2021 92% 0.0 EU741063.1
CP2D Enterobacter sp. enrichment culture clone DWSR 106 16S ribosomal RNA gene, partial sequence
88% 998 69% 0.0 JN944751.1
identified as Enterobacter sp. with percentage of similarity of 92% and 93%, respectively. Maximum score, E value and the percentage of similarities of siderophore-producing bacteria were presented in Table 8. Phylogenetic tree of the related bacteria is presented in Figure 3. Tian
et al. (2009) reported that Enterobacter and
Pseudomonas were dominant in the rhizosphere of tobacco, with 44.5% and 24.7% total frequency, respectively.
Siderophores are produced by various bacteria and fungi, usually classified by the ligands used to chelate the ferric iron. The major groups of siderophores include the catecholates
CONCLUSIONS
The average of siderophore-producing bacteria isolated from Cipanas District (1.98 x 10 7
cfu/g) was not significantly different from those isolated from Lembang District (5.3 x 10 cfu/g). 7
The highest diameter of inhibition zone to
Ralstonia solanacearum was 7.0 mm, produced by isolate CP2D. The selected bacteria producing the inhibition zone did not significantly affect seed viability, plant growth, fresh weight and dry weight of tomato compared to control. Based on the characteristics of colony morphology, physiology, biochemistry and partial sequence of 16S rDNA, two selected isolates, i.e. CP1C and CP2D were identified as Brevundimonas sp. and Enterobacter sp., respectively.
ACKNOWLEDGEMENTS
This research was funded by DIPA IPB with scheme of Decentralized Research Program (Program Penelitian Desentralisasi: Hibah Bersaing), Contract No. 27/I3.24.4/SPK/ PD/2010, on 5 March 2010. The authors also thank Dr Kikin Hamzah Mutaqin for the guidance in several molecular tests activities.
REFERENCES
Agrios GN. 2005. Plant pathology. Fifth Edition. New York (US): Academic Press. 992 p.
Bhattacharya A. 2010. Siderophore mediated metal uptake by Pseudomonas fluorescens and its comparison to iron (III) chelation. Cey J Sci (Bio. Sci.)39(2):147-55. Gross M. 1990. Siderophores and fluorescent pigments.
In: Klement Z, Rudolph K, Sands DC, editors. Methods in phytobacteriology. Budapest (HU): Akadémiai Kiadó. 568 p.
Hu QP, Xu JG. 2011. A simple double-layered chrome azurol S agar (SD-CASA) plate assay to optimize the production of siderophores by a potential biocontrol agent Bacillus. Afr J Microbiol Res 5(25):4321-7.
Jenifer MRA, Reena A, Aysha OS, Valli S, Nirmala P, Vinothkumar P. 2013. Isolation of siderophore producing bacteria from rhizosphere soil and their antagonistic activity against selected fungal plant pathogens. Int J Curr Microbiol App Sci 2(1):59-65.
Jeung Y, Kim J, Kang Y. 2007. Genetic diversity and distribution of Korean isolates of Ralstonia solanacearum. Plant Dis 91(10):1277-87.
Klement Z, Rudolph K, Sands DC. 1990. Methods in phytobacteriology. Budapest (HU): Akadémiai Kiadó. 568 p.
Louden BC, Haarmann D, Lynne AM. 2011. Use of blue agar CAS assay for siderophore detection. J Microbiol Biol Educ 12(1):51-3.
Neilands JB. 1995. Siderophore: Structure and function of microbial iron transport compounds. J Biol Chem 270(45):26723-6.
Pal RP, Gokarn K. 2010. Siderophores and pathogenicity of microorganisms. J Biosci Tech 1:127-34.
Rachid D, Ahmed B. 2005. Effect of iron and growth inhibitors on siderophores production by
Pseudomonas fluorescens. Afr J Biotechnol 4(7):697-702. Available online at http://www. academicjournals. org/AJB.
Saharan BS, Nehra V. 2011. Plant growth-promoting Rhizobacteria: A critical review. Life Sciences and Medicine Research, Volume 2011:LSMR-21. http://astonjournals. com/lsmr [Retrieved on 24 October 2011].
Sambrook J, Russell DW. 2001. Molecular cloning. A laboratory manual. Third edition. New York (US): Cold Spring Harbor Lab Pr. p.6-62.
Sayyed RZ, Badgujar MD, Sonawane HM, Mhaske MM, Chincholkar SB. 2005. Production of microbial iron chelators (siderophores) by fluorescent Pseudomonads. Indian J Biotechnol 4:484-90.
Schaad NW, Jones JB, Chun W. 2001. Laboratory guide for identification of plant pathogenic bacteria. Third edition. St. Paul (US): The American Phytopathological Society. 373 p.
Sridevi M, Kumar KG, Mallaiah KV. 2008. Production of catechol-type of siderophores by Rhizobium sp. isolated from stem nodules of Sesbania procumbens
(Roxb.) W and A. Res J Microbiol 3(4):282-7. Tian F, Ding Y, Zhu H, Yao L, Du B. 2009. Genetic
diversity of siderophore-producing bacteria of tobacco rhizosphere. Braz J Microbiol 40:276-84. Wang JF, Lin CH. 2005. Integrated management of
tomato bacterial wilt. The World Vegetable Center. h t t p : / / w w w. a v r d c . o r g / p d f / P R O D 5 -management_bacterial_wilt.pdf. [Retrieved on 26 September 2011].
Yuliar, Nion YA, Toyota K. 2015. Recent trends in control methods for bacterial wilt diseases caused by