www.elsevier.com / locate / bres
Research report
The cardiac sodium channel mRNA is expressed in the developing
and adult rat and human brain
a ,
*
b b cLaurel M. Donahue
, Penelope W. Coates , Vaughan H. Lee , Denise C. Ippensen ,
c c
Steven E. Arze , Shirley E. Poduslo
a
Cascade Biologics, Inc., 4475 SW Scholls Ferry Road, Portland, OR 97225, USA
b
Department of Cell Biology and Biochemistry, Texas Tech University Health Sciences Center, Lubbock, TX 79430, USA
c
Division of Neurology, Texas Tech University Health Sciences Center, Lubbock, TX 79430, USA Accepted 26 September 2000
Abstract
Expression of the rat (RH-I / SkM2) and human (hH1 / SCN5A) tetrodotoxin-resistant (TTX-R), voltage-sensitive sodium channels is thought to be specific to cardiac tissue. We detected RH-I / SkM2 mRNA in newborn rat brain using both RNase protection assay analysis and in situ hybridization and in adult rat brain using RNase protection assay analysis. This expression was observed primarily in developing limbic structures of the cerebrum and diencephalon, and in the medulla of the brain stem. Using RT-PCR analysis, we detected hH1 / SCN5A mRNA in both fetal and adult human brain. Interestingly, mutations in the human cardiac sodium channel are known to lead to cardiac abnormalities, which result in arrhythmias and frequently in sudden cardiac death. If these mutant channels were also expressed in limbic regions of the brain, alterations in channel function could have drastic effects on the brain’s signaling ability, possibly promoting seizure activity. 2000 Elsevier Science B.V. All rights reserved.
Theme: Excitable membranes and synaptic transmission
Topic: Sodium channels
Keywords: Cardiac sodium channel; RH-1 / SkM2; hH1 / SCN5A; Limbic; mRNA expression; Long QT3 syndrome
1. Introduction
cardiac cells [2,3,6,10,11,20,22,23,34,42,46,48,49,54,60].
In addition, they are also expressed in glial cells
Voltage gated sodium channels (sodium channels) are
[3,6,10,24,48,51]
and
other
peripheral
tissues
essential for the electrical excitability of neurons and other
[3,8,20,23,24,26,28,35,36,42]. Although the sodium
chan-excitable cells. Sodium channels comprise a multigene
nel isoforms were originally defined based on the tissue or
family with two subfamilies determined by sequence
cell type from which they were isolated (rat brain I, II, and
homology [10]. Within each family, the channel subtypes
III, the cardiac channel, and the glial channel), they are
are distinguished by their single channel conductance and
actually expressed in a variety of tissues. In fact, many
sensitivity to pharmacological agents [22,34,60]. Sodium
tissues
express
multiple
channel
isoforms
[5,7–
channels are expressed in a number of different cell types,
10,17,18,21,22,24,29,39,47,53,56,57]. For example, the rat
including excitable cells, such as central nervous system
brain I and II sodium channels, as well as the glial sodium
(CNS) neurons, peripheral nervous system (PNS) sensory
channel, are expressed in rat spinal sensory neurons [9].
neurons, neuroendocrine cells, skeletal muscle cells, and
Sodium channels vary in their sensitivity to tetrodotoxin
(TTX), a guanidinium neurotoxin that interacts exclusively
with sodium channels [22,28,60]. Sodium channels are
broadly divided into three classes based on their sensitivity
to TTX [18,62]. Those demonstrating the highest TTX
*Corresponding author. Tel.: 11-503-292-9521; fax: 1
1-503-292-sensitivity are designated TTX-sensitive (TTX-S) channels
0566.
E-mail address: [email protected] (L.M. Donahue).
and are blocked with low concentrations of TTX (1–5
nM); those with intermediate sensitivity, TTX-resistant
2. Materials and methods
(TTX-R) channels, require higher concentrations of TTX
(0.2–1
m
M) for inactivation; and those with the lowest
2.1. RNA isolation and quantification
sensitivity, TTX-insensitive (TTX-I) channels, are blocked
only with very high concentrations of TTX (30–100
m
M).
Total RNA from adult rat heart, and adult and postnatal
To date, there are five cloned sodium channels that are not
day 0 (P0) Sprague–Dawley rat brains was isolated as
1
TTX-S: the rat RH-I / SkM2 and the human HH1 / SCN5A
described previously [19]. Poly (A)
RNA from HeLa
cardiac sodium channel genes, whose gene products are
cells (human cervical epithelial carcinoma cells; American
TTX-R [60], the rat and the human NaN / SNS2 sensory
Type Culture Collection, Rockville, MD) was also isolated
neuron sodium channel gene products which are TTX-I
as described previously [17]. Total RNA from adult human
[13], and the rat SNS / PN3 sensory neuron sodium channel
heart, and adult and fetal human brain was obtained from
gene product, which is TTX-I [2,46].
Clontech Laboratories, Inc. (Palo Alto, CA). The
con-One of the most extensively studied sodium channels is
centration of each RNA was estimated by optical density
the cardiac sodium channel. It has been cloned from rat
measurements and the relative amounts of RNAs used in
(RH-I / SkM2) and human (HH1 / SCN5A) cardiac cDNA
the RNase protection assays were normalized to
cyto-libraries [25,33,45,55]. Electrophysiological studies of rat
plasmic actin mRNA levels.
and human cardiac sodium channels expressed in Xenopus
oocytes demonstrated that RH-I / SkM2 and HH1 / SCN5A
are the functional TTX-R sodium channels expressed by
2.2. RNase protection assay
cardiac cells [22,32]. RH-I / SkM2 is expressed in neonatal
and adult rat heart: neonatal heart expresses solely RH-I /
RNase protection assays were performed as previously
SkM2, whereas adult heart expresses both RH-I / SkM2 and
described ([19]; RPA II kit, Ambion, Austin, TX). In
a TTX-S NaCh current [33,45,49]. RH-I / SkM2 expression
assays for RH-1 / SkM2 expression, 50
m
g of total RNA
is also present in neonatal skeletal muscle and in de-
from P0 and adult rat brain, 5
m
g of total RNA from adult
nervated adult muscle, but is absent in innervated adult
rat heart, and 5
m
g of total RNA from yeast were
4 32
muscle [33,45]. The expression of HH1 / SCN5A has only
hybridized to 5
3
10 cpm of the
P-labeled RH-1 / SkM2
been examined in adult tissues and while generally re-
antisense RNA probe. The 414 nucleotide (nt)
gene-spe-ported to be specific to cardiac tissue [25], a few reports
cific probe for RH-1 / SkM2 was generated by BamHI
demonstrate its localization in adult rat brain [18,30,59,61].
linearization of the plasmid SK- / 15-2-1 and transcription
To date, the sodium channel genes that have been cloned
with T7 polymerase. The Sk- / 15-2-1 plasmid ([33]; gift
from rat brain are TTX-S (I, II, III, and NaCh6
from Dr. Roland Kallen, University of Pennsylvania,
[4,16,27,38,50,53]). Although earlier studies failed to
Philadelphia, PA) contained nucleotides 6718–7076 from
distinguish between the TTX-R and TTX-I sodium channel
the 3
9
untranslated region of the mRNA. The expected size
currents, several studies suggest that both kinds of current
of the protected fragment was 358 nt. RPAs were also
exist in the CNS [14,31,43,52,58,60]. However, the genes
performed using a cytoplasmic actin antisense RNA probe
responsible for the TTX-R and the TTX-I currents in the
to control for RNA load. In this case, 5
m
g of each of the
4 32
CNS have not yet been identified. Because we demon-
RNAs were hybridized to 1
3
10 cpm of the
P-labeled
strated expression of the rat RH-I / SkM2 cardiac channel
actin antisense RNA probe. The template for cytoplasmic
mRNA in the neuronal cell lines, RT4-B8 and RT4-E5
actin (pTRI-
b
-Actin-125-Rat) was obtained from Ambion
[17] and others detected its expression in B104 neuro-
(Austin, TX); a 218 nt antisense RNA probe was
syn-blastoma cells [29], we hypothesized that expression of
thesized using SP6 polymerase. The predicted size of the
this gene might be responsible for the TTX-R current in
protected fragment was 126 nt. Both the RH-1 / SkM2 and
the nervous system. In fact, using RNase protection assays
actin antisense RNA probes were synthesized using the
of total brain RNA, we and others have observed RH-I /
Riboprobe System kit (Promega Corporation, Madison,
SkM2 mRNA expression in neonatal and adult rat brain
WI). RPAs were visualized both by autoradiography and
[18,59,61].
by using a Molecular Dynamics PhosphoImager 445SI
In this study, we demonstrate that RH-I / SkM2 and
(Sunnyvale, CA). Band intensities were quantified using
HH1 / SCN5A mRNAs were expressed in both developing
ImageQuaNT software (Molecular Dynamics).
and adult rat and human brain by RNase protection assay
and RT-PCR analyses. In situ hybridization of P0 rat brain
further demonstrated that RH-I / SkM2 mRNA expression
2.3. In situ hybridization
was localized to limbic and certain autonomic structures of
m
m serial coronal sections. Sections were de-paraffinized,
expression of the RH-I / SkM2 cardiac sodium channel
dehydrated through a series of ethanol washes, washed in
gene in P0 and adult rat brain (Fig. 1). When qualitatively
PBS, subjected to Proteinase K (20
m
g / ml) digestion, and
normalized to levels of cytoplasmic actin mRNA
expres-pre-hybridized for 2–4 h at room temperature [32,38].
sion (Fig. 1B), RH-I / SkM2 mRNA was detected at
Sections were hybridized at 50
8
C for 16–20 h with 2 ng of
roughly equal levels in both P0 and adult rat brain in three
35
either the
S-labeled RH-1 / SkM2 sense or antisense RNA
separate experiments (representative example shown in
probe from the plasmid SK- / 15-2-1 ([33]; specific activity
Fig. 1A). As expected, RH-I / SkM2 mRNA was detected
of the probes ranged from 12 to 23
m
Ci /
m
g). The sense
in the rat heart RNA positive control, but not in the
probe was generated by linearizing the SK- / 15-2-1 plas-
negative control, yeast RNA.
mid with SalI followed by transcription with T3
poly-merase. The antisense probe was generated by linearizing
the plasmid with SacI followed by transcription with T7
polymerase. After hybridization, the sections were washed
for 30 min in 50% formamide, 1
3
SSC (0.15 M NaCl /
0.015 M Na citrate), 10 mM DTT at 50
8
C, followed by a
30-min wash in 0.5
3
SSC at room temperature. The
sections were then treated for 30 min with 20
m
g / ml
RNase A at room temperature, washed for 2 h in 0.1
3
SSC at 75
8
C, and dehydrated through a series of ethanol
washes. Sections were dipped in NTB2 emulsion (Eastman
Kodak Company, Rochester, NY), exposed for 3 weeks at
4
8
C, developed, and counterstained with Harris modified
hematoxylin [37,44]. The sections were examined under
both brightfield and darkfield optics on an Olympus BX-60
microscope. Identification of rat P0 brain regions was
based on Paxinos et al. [41].
2.4. RT-PCR amplification
The GeneAmp RNA PCR Core Kit (Perkin Elmer
Cetus, Norwalk, CT) was used to reverse transcribe and
amplify 500 ng of DNase-treated human heart and brain
total RNAs, as well as 200–500 ng of DNase-treated HeLa
1
poly (A)
RNA. Here, 100 pmol of each primer were used
in 50
m
l reactions. The primers used were from the domain
I-II linker region of hH1 / SCN5A [55] and were used to
amplify a 410 base pair (bp) fragment. The forward primer
was 5
9
-(CAGGACTTCTATGAAGCCACG)-3
9
, which
ex-tends from nucleotides 1698 to 1718, and the reverse
primer
was
5
9
-(AAGCCATCTACACACGGAGC)-3
9
,
which extends from nucleotides 2089 to 2108 [55].
Ampli-fication conditions were 95
8
C for 5 min, 30 cycles of 94
8
C
for 1 min, 62
8
C for 1.5 min, and 72
8
C for 1 min, followed
by an incubation at 72
8
C for 5 min. The PCR products
were separated by electrophoresis on 2% agarose gels.
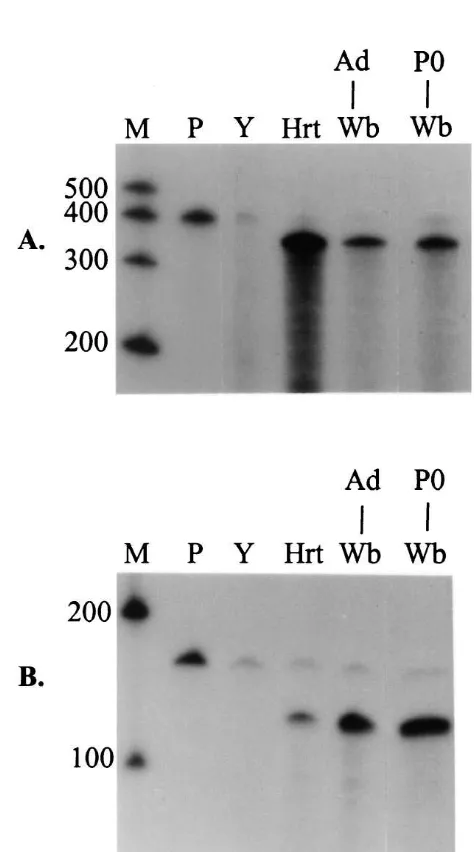
Fig. 1. RNase protection assays of RH-I / SkM2 and cytoplasmic actin. Lane designations: M, markers; P, probe; Y, yeast; Hrt, adult rat heart; Ad, adult; P0, postnatal day 0; Wb, whole brain. (A) RH-I / SkM2 RPA. Fifty
3. Results
mg of total RNA from P0 and adult rat brain were compared. Fivemg of total RNA from adult rat heart was used as a positive control. Fivemg of total RNA from yeast was used as a negative control. The predicted size3.1. The RH-I /SkM2 rat cardiac sodium channel mRNA
of the protected fragment was 358 nt. (B) Cytoplasmic actin RPA. Five
is expressed in P
0 and adult rat brain
mg of each of the RNAs were compared. The predicted size of theprotected fragment was 126 nt. In both (A) and (B), the asterisk (*) marks
3.2. Expression of the RH-I /SkM2 rat cardiac sodium
tions with the sense probe did not reveal any detectable
channel mRNA is localized to limbic regions of the P
0
expression in brain, as expected (Figs. 2C, 2F, 3C, 4D).
rat brain
In the forebrain of the P0 rat brain, intense RH-I / SkM2
gene expression was localized to the developing septal
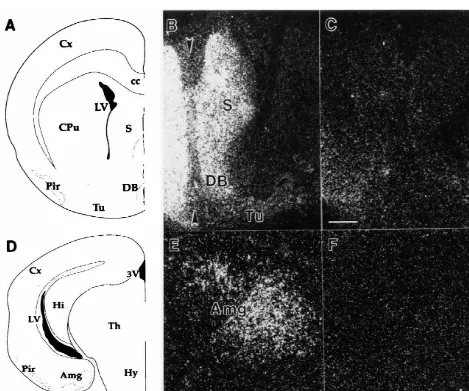
In situ hybridization was performed to examine the
region and the vertical and horizontal limbs of the diagonal
anatomic localization of RH-I / SkM2 cardiac sodium chan-
band of Broca (Fig. 2B). Within this section, signal
nel mRNA in the P0 rat brain (Figs. 2–4). Both sense and
continued ventrally in a diffuse, intense pattern into the
antisense RNA probes were used. In situ hybridizations
developing olfactory tubercule and the piriform cortex
with the antisense probe revealed that the expression of
(Fig. 2B). Strong, localized gene expression of RH-I /
mRNA for the RH-I / SkM2 cardiac sodium channel gene
SkM2 was observed in the region of the developing medial
was localized to three major regions in the P0 rat brain, all
basal amygdala (Fig. 2E). Furthermore, expression of
RH-of which are part RH-of the limbic system. In situ hybridiza-
I / SkM2 was also observed in the developing neocortex at
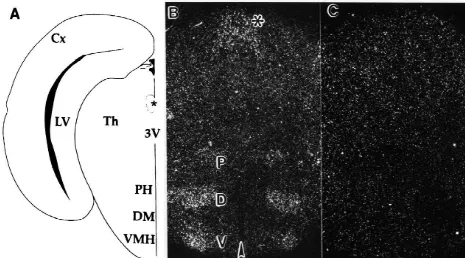
Fig. 3. In situ hybridization of the RH-I / SkM2 rat cardiac sodium channel in the diencephalon of P0 rats. (A–C) In situ hybridization of RH-I / SkM2 in developing hypothalamic and thalamic nuclei. (A) Drawing of the regions shown in (B) and (C). (B) Darkfield photomicrograph of a coronal section demonstrating RH-I / SkM2 expression in the VMH, DM, PH regions and the medial habenular-pretectal complex. Bar shown is 745mm. Arrowhead indicates the 3rd ventricle of the brain. The drawing (A) depicts an anterodorsal to ventrocaudal view, while the sections in the darkfield photomicrographs (B, C) are oriented ventrocaudal to posterodorsal. (C) Darkfield photomicrograph of the adjacent serial section to that seen in (B) hybridized with the sense RH-I / SkM2 probe. Abbreviations: Cx, cortex; Th, thalamus; VMH and V, ventral medial hypothalamic area; DM and D, dorsal medial hypothalamic area; PH and P, posterior hypothalamic area; asterisk (*), medial habenular–pretectal complex; 3V, 3rd ventricle; LV, lateral ventricle.
low, but detectable levels (data not shown). In this case,
performed with and without reverse transcriptase to ensure
the signal appeared to be localized over clusters of
that the 410 bp hH1 / SCN5A product did not arise from
immature neurons.
residual genomic DNA contamination of the RNA
sam-In the diencephalon of the P0 rat brain, relatively
ples. In reactions performed with the enzyme, expression
moderate levels of RH-I / SkM2 expression were localized
of the hH1 / SCN5A human cardiac sodium channel mRNA
to developing hypothalamic and thalamic nuclei (Fig. 3).
was detected in both human fetal and adult brain RNA, as
These included three hypothalamic groups corresponding
well as in the positive control sample of adult human heart
most closely to the ventral medial, dorsal medial and
RNA. Cloning and sequencing of the PCR products
posterior hypothalamic nuclei, as well as the medial
confirmed that they were hH1 / SCN5A (data not shown).
habenular–pretectal complex (Fig. 3B). Signal was located
No expression of hH1 / SCN5A mRNA was detected in
mostly over neurons (data not shown).
samples lacking reverse transcriptase or in the negative
1
In the brainstem, intense RH-I / SkM2 signal was local-
control sample of HeLa poly (A)
RNA.
ized to the ventrolateral medulla in a well-circumscribed
area that most closely approximated the lateral
paragigan-tocellular nuclei (Fig. 4B). Expression was observed in
cells with morphology typical of small, immature neurons
4. Discussion
(Fig. 4C) as observed in counterstained sections (data not
shown).
Expression of the rat TTX-R sodium channel gene
RH-I / SkM2 and its human ortholog, hH1 / SCN5A, is
3.3. The hH1 /SCN5A human cardiac sodium channel
generally considered to be specific for cardiac and skeletal
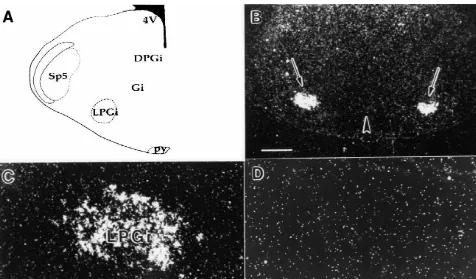
Fig. 4. In situ hybridization of the RH-I / SkM2 rat cardiac sodium channel in the medulla of P0 rats. (A–D) In situ hybridization of RH-I / SkM2 in the developing lateral paragigantocellular nucleus. (A) Drawing of the region in the medulla shown in panels (B) to (D). (B) Dark-field photomicrograph of a coronal section demonstrating RH-I / SkM2 expression in both LPGi nuclei (long arrows). Bar shown is 370mm. The arrowhead in (B) marks the brain midline. (C) Darkfield photomicrograph of a coronal section demonstrating RH-I / SkM2 expression in a single LPGi nucleus. Bar shown is 75mm. (D) Darkfield photomicrograph of the adjacent serial section to that seen in (C) hybridized with the sense RH-I / SkM2 probe. Abbreviations: LPGi, lateral paragigantocellular nucleus; Gi, gigantocellular nucleus; DPGi, dorsal paragigantocellular nucleus; Sp5, spinal trigeminal nucleus; 4V, 4th ventricle.
addition, we observed that hH1 / SCN5A mRNA was
the areas of RH-I / SkM2 expression, which were primarily
expressed in human fetal and adult brain.
in limbic structures. RH-I / SkM2 hybridization was
ob-RNase protection assays using an RH-I / SkM2-specific
served in the forebrain, the diencephalon, and the medulla.
probe demonstrated that the rat TTX-R cardiac sodium
In the forebrain, a high level of expression was detected in
channel is expressed in newborn and adult rat brain tissue.
developing limbic system regions, including the septal
In situ hybridization studies using P0 rat brain delineated
region, the diagonal band of Broca, and the medial basal
amygdala. Lower levels of expression were observed in the
developing olfactory tubercule, the piriform cortex, and the
neocortical region. Similar patterns have recently been
reported in adult rat forebrain regions [30]. The expression
we observed in the piriform cortex corresponds with the
1
heart-like Na
current recorded from acutely dissociated
neurons of the superficial adult rat medial entorhinal cortex
[58] and the neocortical expression we observed
corre-sponds to previous reports in the literature of both
neocortical expression of RH-I / SkM2 mRNA and of a
TTX-R current recorded from neocortical neurons [14].
RH-I / SkM2 mRNA expression was also observed by in
situ hybridization in diencephalic regions. As is typical,
boundaries in the diencephalon of P0 animals were
dif-Fig. 5. RT-PCR of hH1 / SCN5A. Lane designations: M, markers; Hrt,
ficult to establish, but three reasonably well circumscribed
adult human heart; HeLa, human cervical epithelial carcinoma cell line;hypothalamic areas and one (epi)thalamic area were
pres-Fb, fetal human brain; Ab, adult human brain. (1) and (2) indicate theent. The hypothalamic expression we observed correlates
presence or absence, respectively, of reverse transcriptase in the reaction
with a previous electrophysiological study wherein a
TTX-mix. The predicted size of the hH1 / SCN5A amplified fragment was 410
neuro-voltage-gated sodium channel expressed by sensory neurons, Nature
epithelial cell line derived from the mouse hypothalamus
379 (1996) 257–262.
[62]. Interestingly, intense RH-I / SkM2 signal was also
[3] A.N. Akopian, V. Souslova, L. Sivilotti, J.N. Wood, Structure and
observed in a well circumscribed area in the ventrolateral
distribution of a broadly expressed atypical sodium channel, FEBSmedulla. Due to the immaturity of the animals, boundaries
Lett. 400 (1997) 183–187.[4] V.G. Auld, A.L. Goldin, D.S. Krafte, J. Marshall, J.M. Dunn, W.A.
of nuclei in the brainstem of P0 animals were also difficult
1 Catterall, H.A. Lester, N. Davidson, R.J. Dunn, A rat brain Na
to establish; thus it is possible that the area also includes
channel a subunit with novel gating properties, Neuron 1 (1988)
the rostroventrolateral reticular nucleus that projects direct-
449–461.ly to the intermediolateral cell column in the spinal cord
[5] S. Beckh, Differential expression of sodium channel mRNAs in rat peripheral nervous system and innervated tissues, FEBS Lett. 262[40]. The ventrolateral medulla is part of the reticular
(1990) 317–322.
formation that regulates autonomic nervous system
func-[6] S.M. Belcher, C.A. Zerillo, R. Levenson, J.M. Ritchie, J.R. Howe,
tion, and it is known that certain hypothalamic and
Cloning of a sodium channel alpha subunit from rabbit Schwannautonomic centers in the brainstem are involved with
cells, Proc. Natl. Acad. Sci. USA 92 (1995) 11034–11038.cardiovascular regulation [40].
[7] J.A. Black, S. Yokoyama, S.G. Waxman, Y. Oh, K.B. Zur, H.Sontheimer, H. Higashida, B.R. Ransom, Sodium channel mRNAs
While we have demonstrated that mRNA for the TTX-R
in cultured spinal cord astrocytes: in situ hybridization in identified
cardiac sodium channel is expressed in rat and human
cell types, Mol. Brain Res. 23 (1994) 235–245.
brain, whether this mRNA is translated is unclear. For
[8] J.A. Black, R.E. Westenbroek, W.A. Catterall, S.G. Waxman, Type IIexample, it has been reported that the GABA A receptor
brain sodium channel expression in non-neuronal cells: embryonicrat osteoblasts, Mol. Brain Res. 34 (1995) 89–98.
mRNA and the NMDA R1 mRNA are present in some
[9] J.A. Black, S. Dib-Hajj, K. McNabola, S. Jeste, M.A. Rizzo, J.D.
neurons, but are not translated [9]. However, it is likely
Kocsis, S.G. Waxman, Spinal sensory neurons express multiple
that at least in the piriform cortex the RH-I / SkM2 mRNA
sodium channel alpha-subunit mRNAs, Mol. Brain Res. 43 (1996)we detected is translated, since a heart-like sodium current
117–131.[10] J.A. Black, S.G. Waxman, Sodium channel expression: a dynamic
has been previously recorded in medial entorhinal cortex
process in neurons and non-neuronal cells, Dev. Neurosci. 18 (1996)
neurons from adult rats [58].
139–152.
Mutations in the human cardiac sodium channel gene
[11] W.A. Catterall, Cellular and molecular biology of voltage-gatedare known to lead to two cardiac abnormalities: congenital
sodium channels, Physiol. Rev. 72 (1992) S15–48.Long QT3 (LQT3) syndrome and idiopathic ventricular
[12] Q. Chen, G.E. Kirsch, D. Zhang, R. Brugada, J. Brugada, P. Brugada, D. Potenza, A. Moya, M. Borggrefe, G. Breithardt, R.fibrillation with right bundle branch block and ST segment
Ortiz-Lopez, Z. Wang, C. Antzelevitch, R.E. O’Brien, E.
Schulze-elevation [1,12]. Electrophysiological analysis shows that
Bahr, M.T. Keating, J.A. Towbin, Q. Wang, Genetic basis and
these mutations affect inactivation of the channels and lead
molecular mechanism for idiopathic ventricular fibrillation, Nature 1to small, but significant increases in Na
influx. These
392 (1998) 293–296.conditions can lead to arrhythmias and sudden cardiac
[13] T.R. Cummins, S.D. Dib-Hajj, J.A. Black, A.N. Akopian, J.N. Wood, S.G. Waxman, A novel persistent tetrodotoxin-resistantdeath. Sudden unexpected death also occurs in epilepsy.
sodium current in SNS-null and wild-type small primary sensory
The causes are unknown, but may include cardiac
arrhyth-neurons, J. Neurosci. 19 (24) (1999) RC43.
mias generated by epileptic seizures arising from stimula-
[14] R.A. Deisz, A tetrodotoxin-insensitive sodium current initiates bursttion of the autonomic nervous system [15]. While it
firing of neocortical neurons, Neuroscience 70 (1996) 341–351.remains to be investigated, it is possible that the expression
[15] O. Devinsky, B.H. Price, S.I. Cohen, Cardiac manifestations of complex partial seizures, Am. J. Med. 80 (1986) 195–202.of a mutant cardiac sodium channel gene in the limbic
[16] P.S. Dietrich, J.G. McGivern, S.G. Delgado, B.D. Koch, R.M. Eglen,
system has some impact on cardiac instability and / or
J.C. Hunter, L. Sangameswaran, Functional analysis of a
voltage-seizure activity.
gated sodium channel and its splice variant from rat dorsal rootganglia, J. Neurochem. 70 (6) (1998) 2262–2272.
[17] L.M. Donahue, K. Schaller, N. Sueoka, Segregation of Na(1 )-channel gene expression during neuronal-glial branching of a rat
Acknowledgements
PNS-derived stem cell line, RT4-AC, Dev. Biol. 147 (1991) 415– 424.
This study was supported by grants from the Muscular
[18] L.M. Donahue, The tetrodotoxin-insensitive sodium current in ratDystrophy Association and the National Institutes of
dorsal root ganglia is unlikely to involve the expression of the tetrodotoxin-resistant sodium channel, SkM2, Neurochem. Res. 20Health (HD29400) to L.M.D. We wish to thank Drs. Kurt
(1995) 713–717.
Droms and John Orem for critical reading of the
manu-[19] L.M. Donahue, P.W. Coates, A.J. Reinhart, Characterization of
script and valuable discussions and Ms. Anne Carpenter
developmental stage and neuronal potential of the rat PNS-derivedfor technical assistance.
stem cell line, RT4-AC, Dev. Brain Res. 94 (1996) 67–80.[20] A. Felipe, T.J. Knittle, K.L. Doyle, M.M. Tamkun, Primary structure and differential expression during development and pregnancy of a novel voltage-gated sodium channel in the mouse, J. Biol. Chem.
References
269 (1994) 30125–30131.1
[22] H.A. Fozzard, D.A. Hanck, Structure and function of voltage- voltage-sensitive Na channel mRNAs in astrocytes, Mol. Brain dependent sodium channels: comparison of brain II and cardiac Res. 23 (1994) 57–65.
isoforms, Physiol. Rev. 76 (1996) 887–926. [40] G. Paxinos (Ed.), The Rat Nervous System, Forebrain and Midbrain; [23] J. Garcia-Anoveros, B. Derfler, J. Neville-Golden, B.T. Hyman, D.P. Hindbrain and Spinal Cord, Vols. I and II, Academic Press, San
Corey, BnaC1 and BmaC2 constitute a new family of human Diego, 1985.
neuronal sodium channels related to degenerins and epithelial [41] G. Paxinos, I. Tork, L.H. Tecott, K.L. Valentino (Eds.), Atlas of the sodium channels, Proc. Natl. Acad. Sci. USA 94 (1997) 1459–1464. Developing Rat Brain, Academic Press, San Diego, 1991. [24] S. Gautron, G. Dos Santos, D. Pinto-Henrique, A. Koulakoff, F. [42] N.W. Plummer, M.W. McBurney, M.H. Meisler, Alternative splicing
Gros, Y. Berwald-Netter, The glial voltage-gated sodium channel: of the sodium channel SCN8A predicts a truncated two-domain cell- and tissue-specific mRNA expression, Proc. Natl. Acad. Sci. protein in fetal brain and non-neuronal cells, J. Biol. Chem. 272
USA 89 (1992) 7272–7276. (1997) 24008–24015.
[25] M.E. Gellens, A.L. George Jr., L.Q. Chen, M. Chahine, R. Horn, [43] M. Raggenbass, M. Goumaz, E. Sermasi, E. Tribollet, J.J. Dreifuss, R.L. Barchi, R.G. Kallen, Primary structure and functional expres- Vasopressin generates a persistent voltage-dependent sodium current sion of the human cardiac tetrodotoxin-insensitive voltage-depen- in a mammalian motorneuron, J. Neurosci. 11 (1991) 1609–1616. dent sodium channel, Proc. Natl. Acad. Sci. USA 89 (1992) 554– [44] S.E. Ravnik, D.J. Wolgemuth, The developmentally restricted pattern
558. of expression in the male germ line of a murine Cyclin A, Cyclin
[26] A.L. George Jr., T.J. Knittle, M.M. Tamkun, Molecular cloning of A2, suggests roles in both mitotic and meiotic cell cycles, Dev. Biol. an atypical voltage-gated sodium channel expressed in human heart 173 (1996) 69–78.
and uterus: evidence for a distinct gene family, Proc. Natl. Acad.
[45] R.B. Rogart, L.L. Cribbs, L.K. Muglia, D.D. Kephart, M.W. Kaiser,
Sci. USA 89 (1992) 4893–4897. 1
Molecular cloning of a putative tetrodotoxin-resistant rat heart Na [27] A. Goldin, T. Snutch, H. Lubbert, A. Dowsett, A. Marshall, V. Auld,
channel isoform, Proc. Natl. Acad. Sci. USA 86 (1989) 8170–8174. W. Downey, L. Fritz, H. Lester, R. Dunn, W. Catterall, N. Davidson,
[46] L. Sangameswaran, S.G. Delgado, L.M. Fish, B.D. Koch, L.B. Messenger RNA coding for only theasubunit of the rat brain Na
Jakeman, G.R. Stewart, P. Sze, J.C. Hunter, R.M. Eglen, R.C. channel is sufficient for expression of functional channels in
Herman, Structure and function of a novel voltage-gated, tetrodotox-Xenopus oocytes, Proc. Natl. Acad. Sci. USA 83 (1986) 7503–
in-resistant sodium channel specific to sensory neurons, J. Biol. 7507.
Chem. 271 (1996) 5953–5956. [28] D.V. Gordienko, H. Tsukahara, Tetrodotoxin-blockable
de-1 [47] K.L. Schaller, D.M. Krzemien, N.M. McKenna, J.H. Caldwell,
polarization-activated Na currents in a cultured endothelial cell line
Alternatively spliced sodium channel transcripts in brain and derived from rat interlobar artery and human umbilical vein, Pflug.
muscle, J. Neurosci. 12 (1992) 1370–1381. Arch. Eur. J. Physiol. 428 (1994) 91–93.
[48] K.L. Schaller, D.M. Krzemien, P.J. Yarowsky, B.K. Krueger, J.H. [29] X.Q. Gu, S. Dib-Hajj, M.A. Rizzo, S.G. Waxman, TTX-sensitive
1 Caldwell, A novel, abundant sodium channel expressed in neurons
and -resistant Na currents, and mRNA for the TTX-resistant rH1
and glia, J. Neurosci. 15 (1995) 3231–3242. channel, are expressed in B104 neuroblastoma cells, J.
Neuro-[49] M.N. Sills, Y.C. Xu, E. Baracchini, R.H. Goodman, S.S. Cooperman, physiol. 77 (1997) 236–246.
1
G. Mandel, K.R. Chien, Expression of diverse Na channel mes-[30] H.A. Hartmann, L.V. Colom, M.L. Sutherland, J.F. Noebels,
Selec-senger RNAs in rat myocardium. Evidence for a cardiac-specific tive localization of cardiac SCN5A sodium channels in limbic
1
Na channel, J. Clin. Invest. 84 (1989) 331–336. regions of rat brain, Nature Neurosci. 2 (1999) 593–595.
[50] M.R. Smith, R.D. Smith, N.W. Plummer, M.H. Meisler, A.L. Goldin, [31] M. Hay, K.A. Lindsley, Membrane properties of area postrema
Functional analysis of the mouse Scn8a sodium channel, J. Neuro-neurons, Brain Res. 705 (1995) 199–208.
sci. 18 (16) (1998) 6093–6102. [32] S. Ji, W. Sun, A.L. George Jr., R. Horn, R.L. Barchi,
Voltage-1 [51] H. Sontheimer, J.A. Black, S.G. Waxman, Voltage-gated Na dependent regulation of modal gating in the rat SkM1 sodium
channel expressed in Xenopus oocytes, J. Gen. Physiol. 104 (1994) channels in glia: properties and possible functions, Trends Neurosci.
625–643. 19 (1996) 325–331.
[33] R.G. Kallen, Z.H. Sheng, J. Yang, L.Q. Chen, R.B. Rogart, R.L. [52] C.E. Stafstrom, P.C. Schwindt, W.E. Crill, Negative slope conduct-Barchi, Primary structure and expression of a sodium channel ance due to a persistent subthreshold sodium current in cat neocorti-characteristic of denervated and immature rat skeletal muscle, cal neurons in vitro, Brain Res. 236 (1982) 221–226.
Neuron 4 (1990) 233–242. [53] H. Suzuki, S. Beckh, H. Kubo, N. Yahagi, H. Ishida, T. Kayano, M. [34] R.G. Kallen, S.A. Cohen, R.L. Barchi, Structure, function and Noda, S. Numa, Functional expression of cloned cDNA encoding
expression of voltage-dependent sodium channels, Mol. Neurobiol. sodium channel III, FEBS Lett. 228 (1988) 195–200.
7 (1993) 383–428. [54] J.J. Toledo-Aral, B.L. Moss, Z.J. He, A.G. Koszowski, T.
[35] N. Kizer, X.L. Guo, K. Hruska, Reconstitution of stretch-activated Whisenand, S.R. Levinson, J.J. Wolf, I. Silos-Santiago, S. Halegoua, cation channels by expression of the alpha-subunit of the epithelial G. Mandel, Identification of PN1, a predominant voltage-dependent sodium channel cloned from osteoblasts, Proc. Natl. Acad. Sci. USA sodium channel expressed principally in peripheral neurons, Proc.
94 (1997) 1013–1018. Natl. Acad. Sci. USA 94 (1997) 1527–1532.
[36] N. Klugbauer, L. Lacinova, V. Flockerzi, F. Hofmann, Structure and [55] Q. Wang, Z. Li, J. Shen, M.T. Keating, Genomic organization of the functional expression of a new member of the tetrodotoxin-sensitive human SCN5A gene encoding the cardiac sodium channel, Gen-voltage-activated sodium channel family from human neuroendoc- omics 34 (1996) 9–16.
rine cells, EMBO J. 14 (1995) 1084–1090. [56] S.G. Waxman, J.A. Black, Expression of mRNA for a sodium [37] V.H. Lee, A.B. Lee, E.B. Phillips, J.K. Roberts, H.M. Weitlauf, channel in subfamily 2 in spinal sensory neurons, Neurochem. Res.
Spatio-temporal pattern of expression of Galectin-3 in the murine 21 (1996) 395–401.
utero-placental complex: evidence for differential regulation, Biol. [57] R.E. Weiss, N. Sidell, Sodium currents during differentiation in a Reprod. 58 (1998) 1277–1282. human neuroblastoma cell line, J. Gen. Physiol. 97 (1991) 521–539.
1
[38] M. Noda, T. Ikeda, H. Suzuki, H. Takeshima, T. Takahashi, M. [58] J.A. White, A. Alonso, A.R. Kay, A heart-like Na current in the Kuno, S. Numa, Expression of functional sodium channels from medial entorhinal cortex, Neuron 11 (1993) 1037–1047.
rat cerebral cortex, Proc. Natl. Acad. Sci. USA 88 (1991) 9453– sodium channels expressed in a peripheral neurotumor-derived cell
9457. line, RT4-B8, Am. J. Physiol. 270 (1996) C1522–1531.