R
EVIEWAutism spectrum disorder: An omics perspective
Alisa G. Woods
1,2∗, Kelly L. Wormwood
1, Armand G. Ngounou Wetie
1, Roshanak Aslebagh
1,
Bernard S. Crimmins
2, Thomas M. Holsen
3and Costel C. Darie
11
Biochemistry & Proteomics Group, Department of Chemistry & Biomolecular Science, Clarkson University,
Potsdam, NY, USA
2
SUNY Plattsburgh Neuropsychology Clinic and Psychoeducation Services, Plattsburgh, NY, USA
3Department of Civil & Environmental Engineering, Clarkson University, Potsdam, NY, USA
Received: August 22, 2014 Revised: September 11, 2014 Accepted: October 7, 2014
Current directions in autism spectrum disorder (ASD) research may require moving beyond
genetic analysis alone, based on the complexity of the disorder, heterogeneity and
conver-gence of genetic alterations at the cellular/functional level. Mass spectrometry (MS) has been
increasingly used to study CNS disorders, including ASDs. Proteomic research using MS is
directed at understanding endogenous protein changes that occur in ASD. This review focuses
on how MS has been used to study ASDs, with particular focus on proteomic analysis. Other
neurodevelopmental disorders have been investigated using MS, including fragile X syndrome
(FXS) and Smith-Lemli-Opitz Syndrome (SLOS), genetic syndromes highly associated with
ASD comorbidity.
Keywords:
Autism spectrum disorder / Neurodevlopmental disorders
1
Autism overview
Autism spectrum disorder (ASD) consists of social deficits,
communication problems, and repetitive behaviors [1]. ASDs
were previously separated in the DSM-IV-TR into Asperger’s
syndrome, autism, and pervasive developmental disorder not
otherwise specified [2, 3], but the DSM-5 collapsed these into
one term, ASD. Clinical aspects are now described as a dyad:
(i) persistent deficits in social communication and (ii)
re-stricted, repetitive behavior, interests, and activities [1, 2]. An
estimated one in every 68 US children has ASD [4],
com-pared to year 2002 estimates of one in 150 children [5] and
worldwide prevalence rates are similar [6]. The reasons for
this increase are not known. Early detection of ASD is highly
desirable to promote early behavioral interventions and more
functional outcomes [7].
Correspondence: Dr. Costel C. Darie, Biochemistry & Proteomics Group, Department of Chemistry & Biomolecular Science Clark-son University, 8 ClarkClark-son Avenue, Potsdam, NY, 13699-5810, USA
E-mail: [email protected] Fax:+1-315-268-6610
Abbreviations: ADHD, attention-deficit hyperactivity disorder; ASD, autism spectrum disorder;FMR1, fragile X mental retarda-tion;FXS, fragile X syndrome;PON1, paraoxanase/arylesterase 1;PPI, protein–protein interaction;SLOS, Smith-Lemli-Opitz syn-drome
ASD is likely a heterogeneous disorder with multiple
causes, involving genes and the environment [8–10].
Numer-ous genetic studies of ASD have been conducted [11],
support-ing not only a strong genetic link in many cases, but also a
high likely degree of heterogeneity of cause [12]. Many
impli-cated genes are involved in nervous system development and
neurotransmitter systems [13]. ASD is more than 20-times
increased in first-degree relatives [14], based on a twin study.
In identical twins, between 60 and 92% ASD concordance
occurs and from 0 to 10% in fraternal twins [14, 15]. Despite
probable genetic influences, numerous associated genes have
been suggested, without clear consistency in many instances.
[16]. Over 100 genes and 40 genomic loci appear to be
asso-ciated [17, 18]. A recent study described the first clear link in
ASD subtypes and the CHD8 gene mutation [19], indicating
that more progress may follow in the genetic understanding
of other subtypes of ASD. Genes involved in ASD have
com-mon molecular mechanisms, [20], suggesting convergence of
function at the protein level [21]. Currently genetic complexity
and genetics alone do not provide clear predictive markers,
limiting the current utility of genetic testing for ASD [22].
Clinical geneticists rule out known genetic syndromes when
evaluating ASD, and 80% of families are left without a
defini-tive diagnosis [23].
∗Additional corresponding author: Dr. Alisa G. Woods, E-mail:
Colour Online: See the article online to view Figs. 1–3 in colour.
In addition to genetic influences on ASD etiology,
envi-ronmental exposures likely play a major role. For example,
certain pesticides and industrial chemicals may influence
the development of ASD, particularly in susceptible
individ-uals [24]. Further studies investigating the timing, dosage or
mechanisms that induce ASD are needed [25]. Recent results
of the Childhood Autism Risks from Genetics and
Environ-ment (CHARGE) study reported that prenatal close
proxim-ity to organophosphates produced a 60% increased ASD risk.
The risk increased further for third trimester exposure [26].
Thousands of high production volume chemicals circulate
through commerce [27], and hundreds of these chemicals
have recently been detected in pregnant women’s tissues [28].
Understanding the link between environmental exposures of
these chemicals and the onset of ASD is a major knowledge
gap [24].
For example, a relatively new class of compounds that
have begun to receive a significant amount of attention is
organophosphorous flame retardants [29]. Currently slated as
a replacement for the pentabrominated diphenyl ether
mix-ture, this class of compounds is ubiquitous and exhibit
ele-vated concentrations relative to other neurotoxicants, such as,
brominated diphenyl ethers in indoor environments [30–32].
Similar to the OP pesticides, triaryl-, and trialkylphosphate
flame retardants hydrolyze in the blood resulting in
diaryl-and dialkylphosphates, respectively, [33,34] to conformations
conspicuously similar to OPs oxon metabolites, suggesting
organophosphorous flame retardants may have OP-like
neu-rotoxicities and contribute to the onset and severity of ASD
in vulnerable groups.
Whether genetically or environmentally induced (or both
in collaboration) numerous investigations indicate
biolog-ical disturbances in ASDs [35–38], but a clear diagnostic
biomarker for ASD is not available. ASDs are diagnosed
based on behavior, and despite the current existence of
gold-standard behavioral tests [39, 40], biomarkers could further
improve screening and diagnosis. ASDs can be
unrecog-nized in children and even in adults, and current screening
for ASDs can produce false positive or false negative results
[41]. Moving beyond genetic analysis alone may provide new
inroads to diagnosis and understanding of ASD biomarkers.
MS-based proteomics could provide one route for exploration.
2
MS analysis of endogenous molecules
in ASDs
2.1 Differential proteomics: qualitative and
quantitative analysis of proteins in ASD
Current analyses using MS to study ASDs have focused on
endogenously produced biomarkers that could aid in
diagno-sis or understanding of ASDs. Despite a variety of different
proteins identified, some consistency has emerged in the
liter-ature, specifically complement proteins and apolipoproteins
(apos).
One study analyzed blood serum in individuals with
ASD and comorbid attention-deficit hyperactivity disorder
(ADHD) (n
=
9 ASD
+
ADHD, ASD alone
n
=
7) compared
to age-matched controls (n
=
12). Three peaks that were
dif-ferent in ASD versus controls were identified, but the amino
acid sequence information of the peptides that corresponded
to these peaks was not identified [42], however, the
investi-gators speculated that the peptides corresponding to these
peaks may be part of an apo protein [43]. Our group obtained
the sera from these investigators, and reanalyzed them in
our lab. We confirmed increased levels of apoA1 and apoA4,
in individuals with ASD [44]. We further found significant
elevations in the high-density lipoprotein associated enzyme
[45] serum paraoxanase/arylesterase 1 (PON1) [44], which is
also involved in toxin metabolism and detoxification (such as
due to organophosphate exposure), and could help prevent
oxidative stress [46]. It interacts with the cholesterol-carrying
proteins known as apos, which bind PON1 increasing the
stability and activity of PON1 [47]. Interestingly PON1 gene
mutations have been associated with ASD [48]. Notably, one
study of 50 children with ASD found that PON1 protein
ac-tivity (but not gene polymorphisms) was associated with ASD
[49], underscoring the need to analyze biomarkers at the
pro-tein level.
Further suggesting the possibility that apos (and
lipid-associated molecules) are indeed ASD biomarkers,
eleva-tions in apo B-100 and apo A-IV were observed in an earlier
proteomic study that compared high functioning ASD with
low-functioning ASD [50]. Significantly elevated complement
factor H-related protein, complement C1q and fibronectin 1
and apoB-100 was also measured in children with ASD
com-pared to typically developing controls in the same study [50]. A
proteomic study of individuals with Asperger’s syndrome has
also implicated apo dysregulation (apoE, apoC2, and apo A1),
although this change seems to be more specific to females.
Specific proteomic studies examining Asperger’s syndrome
compared to non-Asperger’s ASD could help shed light on
this discrepancy [51].
Consistent with the possible presence of dysregulated
complement proteins in ASD, a different MS analysis of blood
plasma used surface-enhanced laser desorption/ionization
TOF MS to examine peptides in plasma of children with ASD
compared to typically developing controls. Increases in three
peptides were measured, corresponding to C3 complement
protein fragments [52,53]. Additional proteomic studies need
to be conducted to confirm the existence of complement
pro-tein and apo disturbances in ASDs. These studies may help to
identify proteomic biomarker signatures, and possible ASD
subtypes associated with specific biomarkers.
of the few published proteomic studies of salivary biomarkers
in ASD [55].
Our group has recently used 2D differential in-gel
elec-trophoresis (2D-DIGE) to investigate the differences between
the salivary proteomes of children with ASD and matched
controls. DIGE is primarily used in protein expression
pro-filing experiments of at least two samples or conditions
allowing the determination of the relative abundance of
pro-teins [56]. In 2D-DIGE, propro-teins are labeled with
fluores-cent cyanine dyes (Cy2, 3, and 5) prior to 2D-PAGE [57],
run on 2D-PAGE and then scanned for quantitative
analy-sis of the two proteomes investigated [58, 59]. In our
2D-DIGE experiments, we observed that many proteins were
differentially expressed between the ASD and control
con-ditions mostly observed as either green or red colored gel
spots; the yellow colored gel spots contained proteins with
unchanged levels (Fig. 1). Nano-LC-MS/MS and
MALDI-MS/MS analysis of the proteins contained in the red or
green-colored gel spots identified many of the
differen-tially expressed proteins. One such protein upregulated in
ASD was S100-A9 also called migration inhibitory
factor-related protein 14 (MRP-14) or calgranulin-B (Fig. 1). This
protein has not been previously reported to be associated
with ASD.
[image:3.595.55.534.311.634.2]In considering proteomic biomarker analysis in ASDs,
specific attributes of the individuals studied, including age
and gender also need to be taken into consideration. Two
studies have emphasized the possibility of gender-specific
differences in ASD biomarkers, finding that males with
Asperger’s syndrome tend to have altered levels of cytokines
and other inflammatory molecules, whereas biomarkers in
fe-males with Asperger’s seem to be growth factors, hormones
and factors associated with lipid transport, and metabolism
[51,60]. Further investigation into gender specific biomarkers
in ASDs is warranted, particularly since there is a gender bias
in ASDs, favoring diagnosis in males. Indeed, neuroimaging
Figure 1. Difference gel electrophoresis (DIGE). (A)Analytical DIGE gel, where protein samples were labeled with fluorescent cyanine dyes (Cy2, 3, and 5) prior to 2D PAGE. The differentially expressed proteins in ASD (cy3 or green) and matched controls (cy5 or red) are either green or red; the yellow ones are unchanged between ASD and controls. (B)Preparative DIGE gel, from which the differentially expressed proteins were picked and digested by trypsin and analyzed by nano-LC-MS/MS or MALDI-MS. The spots that were specific to ASD (left) or control (right) samples are indicated. (C) MALDI-MS/MS spectrum, whose analysis led to the identification of a peptide with the sequence LGHPDTLNQGEFK which is part of protein S100-A9, also named migration inhibitory factor-related protein 14 (MRP-14) or calgranulin-B. This protein was found to be upregulated in ASD in the DIGE experiments The y- and b-ion series, the amino acid sequence of the peptide (top of the spectrum and the name of the protein (left) are indicated. Reprinted with permission from [56].
evidence has suggested that the neuroanatomy of ASD may
differ in males versus females [61].
With regard to age differences, one study has reported that
12 proteins vary with age in individuals with ASD, including
those involved with inflammation, growth, and hormonal
sig-naling. Examples include higher levels of adiponectin in ASD
with increased age and a decrease with age in typically
de-veloping individuals. Other potential markers that increase
with age in ASD include C-reactive protein, haptoglobin,
TRAIL-R3, matrix metalloproteinase, thyroglobulin, and
can-cer antigen 19–9. Many of these molecules are suggestive of
an inflammatory condition. Age-related changes in
biomark-ers and the possible impact of these molecules on behavioral
phenotypes would be a valuable focus of future studies [62].
2.2 Differential proteomics: analysis of PTMs of
proteins in ASD
Modulation of the function of many proteins is achieved
mostly by transient or definitive PTMs of proteins. Transient
modifications include phosphorylation, acetylation,
myristoy-lation, etc., while stable modifications are presented as
disul-fide bonding formation, protein truncation, and/or
glycosy-lation [63–66]. Only one study investigated PTMs in ASD by
examining salivary peptides examined (n
=
27) compared
to age-matched controls using HPLC-ESI-Ion-Trap MS. In
this study, the investigators found hypophosphorylation of
statherin, histatin 1, and acidic proline-rich proteins in
sub-jects with ASDs compared with controls [55].
2.3 Differential proteomics: analysis of
protein-protein interactions (PPIs) in ASD
Most protein biomarkers or biomarker signatures for diseases
and/or disorders are identified using qualitative and
quanti-tative proteomics, reflecting proteins that are either over- or
under-expressed due to the diagnosed condition. However,
recently, additional protein modifications have been
recog-nized. Phosphorylation, acetylation or glycosylation are
sev-eral such examples. Therefore, both protein quantitation and
protein PTMs provide a more comprehensive picture of the
state of the disease/disorder. The same principle applies to
ASD. However, to our knowledge, no research group has
con-sidered, in addition to protein quantitation and protein PTMs,
stable or transient PPIs as a possible extra indicator of a
dis-ease or disorder. Therefore, we believe that both protein levels
(quantitation), protein PTMs, types and location of PTMs, as
well as stable or transient PPIs, with or without PTMs should
be measured as part of a comprehensive approach. In fact,
our lab recently reported that ApoA1, ApoA4, and PON1
pro-teins, in addition to being dysregulated, are also predicted to
interact with each other as part of HDLs (Figs. 2 and 3) [44].
Protein interactions and networks could provide an important
component of biomarker signature identification in ASD and
take into account a systems approach in understanding the
disorder, rather than focusing on a single molecule.
3
MS-based metabolomic and small
molecule analysis in ASD
LC-MS/MS was used to analyze the urine of young adults with
severe ASD and schizophrenia versus nondiagnosed controls
(n
=
15, 18, and 18, respectively). Butofenine, a molecule
re-lated to serotonin, was elevated versus controls in individuals
with ASD and schizophrenia and correlated with hyperactivity
[67]. This corresponds with serotonin abnormalities reported
in ASD in three MS studies [68–70].
Oxidative stress, the generation of ROS causing tissue
and cellular damage [71], has been consistently implicated in
ASD [72], supported by studies using MS. GC-MS identified
higher urinary lipid peroxidation markers and endothelium
activation in individuals with ASD (n
=
26) versus
non-ASD controls (n
=
12). Lipid peroxidation correlated with
endothelium and platelet activation [73]. MS analyses at the
metabolomic and small molecule level have supported that
oxidative stress and dysregulation of serotonin may be present
in ASD. Overall, proteomic evidence uncovered by MS
sup-ports a role for altered complement proteins and
apolipopro-teins in ASDs, and further supports the presence of oxidative
stress and alterations in serotonergic signaling.
4
MS Analysis in fragile X syndrome
[95]. MALDI-TOF MS was used to examine FMR1
methy-lation in blood of 62 premutation (between 55–200 CGG
repeats in FMR1 gene) or 18 full mutation (more than 200
CGG repeats) females compared to controls (n
=
74). FMR1
intron 1 hypermethylation was predictive of verbal cognitive
impairment [96]. In this study a MS-based biochemical
mea-surement was paired with a behavioral test, underscoring the
possibility of interdisciplinary research in
neurodevelopmen-tal disorders with MS. Such an interdisciplinary approach
may increase for neurodevelopmental disorder assessment
in the future, as MS-based proteomic use increases.
5
MS Analysis in Smith-Lemli-Opitz
syndrome
Smith-Lemli-Opitz syndrome (SLOS) is a genetically
in-herited deficit in cholesterol synthesis frequently
associ-ated with ASD, as well as intellectual/learning problems
and many physical problems [97–100]. Partial or total
defi-ciency of the
Dhcr7
gene causes SLOS [101]. Tissue
choles-terol and total scholes-terol levels are substantially depleted, and
7-dehydrocholesterol levels increase as a result of the gene
mutation. High 7DH levels inhibit
Hmgcr
further causing
cellular cholesterol deficits [102]. SLOS symptoms improve
marginally with cholesterol supplementation, and cholesterol
supplementation does not cure or even largely alleviate the
disorder. Notably, there are individuals with ASD without
SLOS with low total cholesterol [100], supporting the
pos-sible association of cholesterol disturbances and some ASD
subtypes.
[image:5.595.57.537.340.667.2]MS studies on SLOS and SLOS animal models have
fo-cused on sterols analysis, specifically 7-dehydrocholesterol
levels and cholesterol [103–107]. Recently the first MS
pro-teomic study in a SLOS model (rat retina) was published.
[108]. The model demonstrates some SLOS characteristics
of retinal degeneration and visual impairment generated by
treatment with AY9944. This drug (AY9944) inhibits DHCR7
(3 beta-hydroxysterol-Delta 7-reductase), the enzyme found
to be defective in SLOS [109]. Rat retinas from the SLOS
rat model (n
=
5) were compared to retinas from age- and
sex-matched controls (n
=
5), via nano-RPLC. Statistically
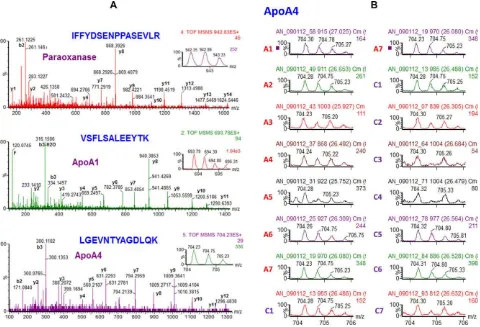
Figure 2. (A) Nano-LC-MS/MS analysis of the sera from children with ASD and matched controls. The MS/MS spectra whose interpretation led to the identification of PON1, ApoA1, and ApoA4. Them/zof the precursor ion, ions, the b and y product ions, the amino acid sequence of the identified peptides and the proteins that the peptides belong to are shown. (B)Relative quantitation of ApoA4 using the precursor ion intensity. The precursor ion investigated hadm/zof 704.23 (2+) and corresponded to the peptide with the amino acid sequence LGEVNTYAGDLQK. A1-A7 are ASD and C1-C7 are controls. Reprinted with permission from [44].
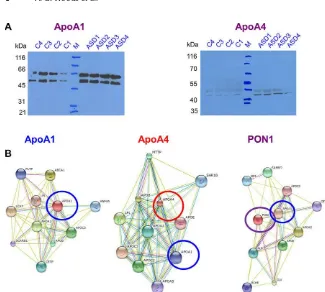
Figure 3. (A) Validation of the LCMS/MS data (shown in Fig. 2) by WB using anti-ApoA1 and anti-ApoA4 Ab. (B) String anal-ysis for ApoA1, ApoA4, and PON1 proteins for identification of the potential interac-tion partners for these proteins. Reprinted with permission from [44].
significant differences in 101 proteins were reported,
in-cluding those regulating lipid metabolism, oxidative stress,
vision, proteolysis, cell death, and vesicular/membrane
trans-port. Western blot and immunohistochemistry validated
spe-cific protein targets. Cathepsin D, glial fibrillary acidic
pro-tein, Stat3, and
-crystallin were elevated in the SLOS model
retina, and apoE was decreased. Further proteomic studies
of human biomaterials taken from individuals with SLOS
could help elucidate proteomic changes that occur and that
contribute to the disorder, in addition to deficits in sterols.
6
Conclusions
MS-based proteomics can be used for the analysis of
human biomaterials to further understand
neurodevelop-mental disorders. For ASD, MS analysis directed at proteomic
biomarker discovery may provide tools to understand ASD
etiology and potentially aid in ASD identification. As such,
proteomic biomarkers will likely be part of a comprehensive
biomarker signature, rather than individual identifiers.
Ulti-mately, they may provide the potential biological diagnostic
(or predictor) for ASD that does not currently exist, likely
complemented by additional measurements, such as
behav-ioral assessments [110]. Proteomic biomarkers may also be
used for treatment monitoring, and can help further
eluci-date the biological consequences of this disorder. In contrast,
FXS and SLOS, are diagnosed via genotyping. Using
MS-based proteomics methods could supplement the diagnosis
of these syndromes or provide further information, such as
indicators of symptom severity. MS-based analyses could be
employed clinically in FXS and SLOS for biomarker
identifi-cation, directed toward protein target discovery,
understand-ing the consequences of these disorders and for monitorunderstand-ing
medication effects. Although in early stages of
implementa-tion, the use of MS holds great potential in furthering the
general understanding of neurodevelopmental disorders.
The authors would like to thank Dr. Witold Winnik
(Envi-ronment Protection Agency, Research Triangle Park, Durham,
NC) for the MALDI MS analysis of the DIGE spots. This work
was supported in part by the Redcay Foundation (SUNY
Platts-burgh), the Alexander von Humboldt Foundation, SciFund
Chal-lenge, private donations (Ms. Mary Stewart Joyce & Mr. Kenneth
Sandler), and by the U.S. Army research office (DURIP grant
#W911NF-11-1-0304).
The authors have declared no conflict of interest.
7
References
[1] APA,Diagnostic and Statistical Manual of Mental Disor-ders. 5th ed. American Psychiatric Association, Arlington, VA 2013.
[2] Woods, A. G., Mahdavi, E., Ryan, J. P., Treating clients with Asperger’s syndrome and autism.Child Adolesc. Psychiatry Ment. Health2013,7, 32.
[4] Autism and Developmental Disabilities Monitoring Net-work Surveillance Year 2010 Principal Investigators, and Centers for Disease Control and Prevention, Prevalence of autism spectrum disorder among children aged 8 years— autism and developmental disabilities monitoring network, 11 sites, United States, 2010.MMWR Surveill. Summ.2014, 63, 1–21.
[5] Autism and Developmental Disabilities Monitoring Net-work Surveillance Year 2002 Principal Investigators, and Centers for Disease Control and Prevention, Prevalence of autism spectrum disorders–autism and developmental dis-abilities monitoring network, 14 sites, United States, 2002. MMWR Surveill. Summ. 2007,56, 12–28.
[6] Quaak, I., Brouns, M. R., Van de Bor, M., The dynamics of autism spectrum disorders: how neurotoxic compounds and neurotransmitters interact.Int. J. Environ. Res. Public Health, 2013,10, 3384–3408.
[7] Lai, M. C., Lombardo, M. V., Baron-Cohen, S., Autism. Lancet2014,383, 896–910.
[8] Herbert, M. R., Contributions of the environment and en-vironmentally vulnerable physiology to autism spectrum disorders.Curr. Opin. Neurol. 2010,23, 103–110.
[9] Ousley, O., Cermak, T., Autism Spectrum Disorder: defining Dimensions and Subgroups.Curr. Dev. Disord. Rep.2014, 1, 20–28.
[10] Xu, L. M., Li, J. R., Huang, Y., Zhao, M. et al., AutismKB: an evidence-based knowledgebase of autism genetics. Nu-cleic Acids Res.. 2012,40(Database issue), D1016–D1022. [11] State, M. W., Levitt, P., The conundrums of understanding
genetic risks for autism spectrum disorders.Nat. Neurosci. 2011,14, 1499–1506.
[12] Leblond, C. S., Heinrich, J., Delorme, R., Proepper, C. et al., Genetic and functional analyses of SHANK2 mutations sug-gest a multiple hit model of autism spectrum disorders. PLoS Genet. 2012,8, e1002521.
[13] Banerjee, S., Riordan, M., Bhat, M. A., Genetic aspects of autism spectrum disorders: insights from animal models. Front. Cell. Neurosci. 2014,8, 58.
[14] Bailey, A., Le Couteur, A., Gottesman, I., Bolton, P. et al., Autism as a strongly genetic disorder: evidence from a British twin study.Psychol. Med. 1995,25, 63–77. [15] Le Couteur, A., Bailey, A., Goode, S., Pickles, A. et al.,
A broader phenotype of autism: the clinical spectrum in twins.J. Child Psychol. Psychiatry1996,37, 785–801. [16] Veenstra-VanderWeele, J., Cook, E. H. Jr., Molecular
genet-ics of autism spectrum disorder.Mol. Psychiatry2004,9, 819–832.
[17] Betancur, C., Etiological heterogeneity in autism spectrum disorders: more than 100 genetic and genomic disorders and still counting.Brain Res. 2011,1380, 42–77.
[18] Nickl-Jockschat, T. and Michel, T. M., [Genetic and brain structure anomalies in autism spectrum disorders. Towards an understanding of the aetiopathogenesis?].Nervenarzt 2011,82, 618–627.
[19] Bernier, R., Golzio, C., Xiong, B., Stessman, H. A. et al., Disruptive CHD8 mutations define a subtype of autism early in development.Cell2014,158, 263–276.
[20] Lanz, T. A., Guilmette, E., Gosink, M. M., Fischer, J. E. et al., Transcriptomic analysis of genetically defined autism can-didate genes reveals common mechanisms of action.Mol. Autism2013,4, 45.
[21] Pinto, D., Delaby, E., Merico, D., Barbosa, M. et al., Con-vergence of genes and cellular pathways dysregulated in autism spectrum disorders.Am. J. Hum. Genet. 2014,94, 677–694.
[22] Gurrieri, F., Working up autism: the practical role of medical genetics.Am. J. Med. Genet. C Semin. Med. Genet. 2012, 160C, 104–110.
[23] Carter, M. T., Scherer, S. W., Autism spectrum disorder in the genetics clinic: a review.Clin. Genet. 2013,83, 399–407. [24] Landrigan, P., Lambertini, L., Birnbaum, L., A research strat-egy to discover the environmental causes of autism and neurodevelopmental disabilities.Environ. Health Perspect. 2012,120, a258–a260.
[25] Shelton, J. F., Hertz-Picciotto, I., Pessah, I. N., Tipping the balance of autism risk: potential mechanisms linking pesti-cides and autism.Environ. Health Perspect. 2012,120, 944– 951.
[26] Shelton, J. F., Geraghty, E. M., Tancredi, D. J., Delwiche, L. D. et al., Neurodevelopmental Disorders and Prenatal Res-idential Proximity to Agricultural Pesticides: the CHARGE Study.Environ. Health Perspect. 2014,122, 1103–1109. [27] Howard, P. H. and Muir, D. C., Identifying new persistent and
bioaccumulative organics among chemicals in commerce. Environ. Sci. Technol. 2010,44, 2277–2285.
[28] Woodruff, T. J., Zota, A. R., Schwartz, J. M., Environmen-tal chemicals in pregnant women in the United States: NHANES 2003–2004.Environ. Health Perspect. 2011,119, 878–885.
[29] van der Veen, I., de Boer, J., Phosphorus flame retardants: properties, production, environmental occurrence, toxicity, and analysis.Chemosphere2012,88, 1119–1153.
[30] Reemtsma, T., Quintana, J. B., Rodil, R., Garcia-Lopez, M., Rodriguez, I., Organophosphorus flame retardants and plasticizers in water and air I. Occurrence and fate.Trends Anal. Chem. 2008.27, 727–737.
[31] Stapleton, H. M., Klosterhaus, S., Eagle, S., Fuh, J. et al., Detection of organophosphate flame retardants in furniture foam and U.S. house dust.Environ. Sci. Technol. 2009,43, 7490–7495.
[32] Van den Eede, N., Dirtu, A. C., Neels, H., Covaci, A., An-alytical developments and preliminary assessment of hu-man exposure to organophosphate flame retardants from indoor dust.Environ. Int. 2011,37, 454–461.
[33] Kurebayashi, H., Tanaka, A., Yamaha, T., Metabolism and disposition of the flame retardant plasticizer, tri-p-cresyl phosphate, in the rat.Toxicol. Appl. Pharmacol. 1985,77, 395–404.
[34] Sasaki, K., Suzuki, T., Takeda, M., Uchiyama, M., Metabolism of phosphoric acid triesters by rat liver ho-mogenate.Bull. Environ. Contam. Toxicol. 1984,33, 281– 288.
[35] Nickl-Jockschat, T., Michel, T. M., The role of neurotrophic factors in autism.Mol. Psychiatry2011,16, 478–490.
[36] Theoharides, T. C., Kempuraj, D., Redwood, L., Autism: an emerging ’neuroimmune disorder’ in search of therapy. Ex-pert Opin. Pharmacother. 2009,10, 2127–2143.
[37] Veenstra-VanderWeele, J., Blakely, R. D., Networking in autism: leveraging genetic, biomarker and model sys-tem findings in the search for new treatments. Neuropsy-chopharmacology2012,37, 196–212.
[38] Woods, A. G., Sokolowska, I., Taurines, R., Gerlach, M. et al., Potential biomarkers in psychiatry: focus on the cholesterol system.J. Cell Mol. Med. 2012,16, 1184–1195.
[39] Lord, C., Risi, S., Lambrecht, L., Cook, E. H. et al., The autism diagnostic observation schedule–generic: a standard mea-sure of social and communication deficits associated with the spectrum of autism.J. Autism Dev. Disord. 2000,30, 205–223.
[40] Lord, C., Risi, S., DiLavore, P. S., Shulman, C. et al., Autism from 2 to 9 years of age.Arch. Gen. Psychiatry2006,63, 694–701.
[41] Dereu, M., Roeyers, H., Raymaekers, R., Meirsschaut, M., Warreyn, P., How useful are screening instruments for tod-dlers to predict outcome at age 4? General development, language skills, and symptom severity in children with a false positive screen for autism spectrum disorder.Eur. Child Adolesc. Psychiatry2012,21, 541–551.
[42] Taurines, R., Dudley, E., Conner, A. C., Grassl, J. et al., Serum protein profiling and proteomics in autistic trum disorder using magnetic bead-assisted mass spec-trometry.Eur. Arch. Psychiatry Clin. Neurosci. 2010,260, 249–255.
[43] Taurines, R., Dudley, E., Grassl, J., Warnke, A. et al., Pro-teomic research in psychiatry.J. Psychopharmacol.2011, 25, 151–196.
[44] Ngounou Wetie, A. G., Wormwood, K., Thome, J., Dudley, E. et al., A pilot proteomic study of protein markers in autism spectrum disorder.Electrophoresis2014,35, 2046–2054. [45] Kotani, K., Yamada, T., Gugliucci, A., Paired measurements
of paraoxonase 1 and serum amyloid A as useful disease markers.Biomed. Res. Int. 2013,2013, 481–437.
[46] Furlong, C. E., Suzuki, S. M., Stevens, R. C., Marsillach, J. et al., Human PON1, a biomarker of risk of disease and exposure.Chem. Biol. Interact. 2010,187, 355–361. [47] Gaidukov, L., Viji, R. I., Yacobson, S., Rosenblat, M. et al.,
ApoE induces serum paraoxonase PON1 activity and sta-bility similar to ApoA-I.Biochemistry2010,49, 532–538. [48] Eskenazi, B., Huen, K., Marks, A., Harley, K. G. et al., PON1
and neurodevelopment in children from the CHAMACOS study exposed to organophosphate pesticides in utero. En-viron. Health Perspect. 2010,118, 1775–1781.
[49] Pasca, S. P., Dronca, E., Nemes, B., Kaucsar, T. et al., Paraox-onase 1 activities and polymorphisms in autism spectrum disorders.J. Cell Mol. Med. 2010,14, 600–607.
[50] Corbett, B. A., Kantor, A. B., Schulman, H., Walker, W. L. et al., A proteomic study of serum from children with autism showing differential expression of apolipoproteins and complement proteins.Mol. Psychiatry2007,12, 292– 306.
[51] Steeb, H., Ramsey, J. M., Guest, P. C., Stocki, P. et al., Serum proteomic analysis identifies sex-specific differences in lipid metabolism and inflammation profiles in adults di-agnosed with Asperger syndrome.Mol. Autism2014,5, 4. [52] Momeni, N., Brudin, L., Behnia, F., Nordstrom, B. et al., High complement factor I activity in the plasma of children with autism spectrum disorders.Autism. Res. Treat.2012.2012, 868576.
[53] Momeni, N., Bergquist, J., Brudin, L., Behnia, F. et al., A novel blood-based biomarker for detection of autism spec-trum disorders.Transl. Psychiatry2012,2, e91.
[54] Loo, J. A., Yan, W., Ramachandran, P., Wong, D. T., Compar-ative human salivary and plasma proteomes.J. Dent. Res. 2010,89, 1016–1023.
[55] Castagnola, M., Messana, I., Inzitari, R., Fanali, C. et al., Hypo-phosphorylation of salivary peptidome as a clue to the molecular pathogenesis of autism spectrum disorders. J. Proteome Res. 2008,7, 5327–5332.
[56] Wetie, A. G., Dekroon, R. M., Mocanu, M., Ryan, J. P. et al., Mass spectrometry for the study of autism and neurodevel-opmental disorders.Adv. Exp. Med. Biol. 2014.806, 525– 544.
[57] Unlu, M., Morgan, M. E., Minden, J. S., Difference gel electrophoresis: a single gel method for detecting changes in protein extracts. Electrophoresis 1997, 18, 2071–2077.
[58] Lilley, K. S., Razzaq, A., Dupree, P., Two-dimensional gel electrophoresis: recent advances in sample preparation, detection and quantitation.Curr. Opin. Chem. Biol. 2002, 6, 46–50.
[59] Tonge, R., Shaw, J., Middleton, B., Rowlinson, R. et al., Val-idation and development of fluorescence two-dimensional differential gel electrophoresis proteomics technology. Pro-teomics2001,1, 377–396.
[60] Schwarz, E., Guest, P. C., Rahmoune, H., Wang, L. et al., Sex-specific serum biomarker patterns in adults with Asperger’s syndrome.Mol. Psychiatry2011,16, 1213–1220.
[61] Lai, M. C., Lombardo, M. V., Suckling, J., Ruigrok, A. N. et al., Biological sex affects the neurobiology of autism. Brain2013,136, 2799–2815.
[62] Ramsey, J. M., Guest, P. C., Broek, J. A., Glennon, J. C. et al., Identification of an age-dependent biomarker signature in children and adolescents with autism spectrum disorders. Mol. Autism2013,4, 27.
[63] Wetie, A. G., Woods, A. G., Darie, C. C., Mass spectro-metric analysis of post-translational modifications (PTMs) and protein-protein interactions (PPIs).Adv Exp. Med. Biol. 2014,806, 205–235.
[64] Ngounou Wetie, A. G., Sokolowska, I., Woods, A. G., Roy, U. et al., Protein-protein interactions: switch from classi-cal methods to proteomics and bioinformatics-based ap-proaches.Cell Mol. Life Sci. 2014,71, 205–228.
[66] Ngounou Wetie, A. G., Sokolowska, I., Woods, A. G., Roy, U. et al., Investigation of stable and transient protein-protein interactions: past, present, and future.Proteomics2013,13, 538–557.
[67] Emanuele, E., Colombo, R., Martinelli, V., Brondino, N. et al., Elevated urine levels of bufotenine in patients with autistic spectrum disorders and schizophrenia.Neuro. Endocrinol. Lett. 2010,31, 117–121.
[68] Kałuzna-Czaplinska, J., Michalska, M., Rynkowski, J., De-termination of tryptophan in urine of autistic and healthy children by gas chromatography/mass spectrometry.Med. Sci. Monit.2010,16, CR488–CR492.
[69] Lam, K. S., Aman, M. G., Arnold, L. E., Neurochemical cor-relates of autistic disorder: a review of the literature.Res. Dev. Disabil. 2006,27, 254–289.
[70] Pedersen, O. S., Liu, Y., Reichelt, K. L., Serotonin uptake stimulating peptide found in plasma of normal individuals and in some autistic urines.J. Pept. Res.53, 641–646. [71] Sokolowska, I., Woods, A. G., Wagner, J., Dorler, J. et al.,
Mass spectrometry for proteomics-based investigation of oxidative stress and heat shock proteins, in: Andreescu, S., Hepel, M. (Eds.),Oxidative Stress: Diagnostics, Preven-tion, and Therapy. American Chemical Society, Washing-ton, D.C. 2011.
[72] Ghanizadeh, A., Akhondzadeh, S., Hormozi, M., Makarem, A. et al., Glutathione-related factors and oxidative stress in autism, a review.Curr. Med. Chem. 2012,19, 4000–4005. [73] Yao, Y., Walsh, W. J., McGinnis, W. R., Pratico, D., Altered
vascular phenotype in autism: correlation with oxidative stress.Arch. Neurol. 2006,63, 1161–1164.
[74] McCary, L. M., Roberts, J. E., Early identification of autism in fragile X syndrome: a review.J. Intellect. Disabil. Res. 2012,57, 803–814.
[75] Boyle, L., Kaufmann, W. E., The behavioral phenotype of FMR1 mutations.Am. J. Med. Genet. C Semin. Med. Genet. 2010,154C, 469–476.
[76] Hagerman, R., Lauterborn, J., Au, J., Berry-Kravis, E., Frag-ile X syndrome and targeted treatment trials.Results Probl. Cell Differ. 2012,54, 297–335.
[77] McCary, L. M. and Roberts, J. E., Early identification of autism in fragile X syndrome: a review.J. Intellect. Disabil. Res. 2013,57, 803–814.
[78] Smith, L. E., Barker, E. T., Seltzer, M. M., Abbeduto, L., Greenberg, J. S., Behavioral phenotype of fragile X syn-drome in adolescence and adulthood.Am. J. Intellect. Dev. Disabil. 2012,117, 1–17.
[79] Brown, W., The molecular biology of fragile X mutation, in: Hagerman, R., Hagerman, P. J., (Eds.),Fragile X Syndrome: Diagnosis, Treatment, and Research, Johns Hopkins Uni-versity Press: Baltimore, MD 2002, pp. 110–135.
[80] De Rubeis, S., Fernandez, E., Buzzi, A., Di Marino, D., Bagni, C., Molecular and cellular aspects of mental retardation in the Fragile X syndrome: from gene mutation/s to spine dys-morphogenesis.Adv. Exp. Med. Biol. 2012,970, 517–551. [81] Hagerman, R. J., Berry-Kravis, E., Kaufmann, W. E., Ono, M.
Y. et al., Advances in the treatment of fragile X syndrome. Pediatrics2009,123, 378–390.
[82] Berry-Kravis, E. M., Hessl, D., Rathmell, B., Zarevics, P. et al., Effects of STX209 (arbaclofen) on neurobehavioral function in children and adults with fragile X syndrome: a random-ized, controlled, phase 2 trial.Sci. Transl. Med. 2012,4, 152ra127.
[83] Dziembowska, M., Pretto, D. I., Janusz, A., Kaczmarek, L. et al., High MMP-9 activity levels in fragile X syndrome are lowered by minocycline.Am. J. Med. Genet. A2013,161A, 1897–1903.
[84] Leigh, M. J., Nguyen, D. V., Mu, Y., Winarni, T. I. et al., A ran-domized double-blind, placebo-controlled trial of minocy-cline in children and adolescents with fragile x syndrome. J. Dev. Behav. Pediatr. 2013,34, 147–155.
[85] Jacquemont, S., Berry-Kravis, E., Hagerman, R., von Rai-son, F. et al., The challenges of clinical trials in frag-ile X syndrome. Psychopharmacology (Berl) 2013, 231, 1237–1250.
[86] Berry-Kravis, E., Hessl, D., Abbeduto, L., Reiss, A. L. et al., Outcome measures for clinical trials in fragile X syndrome. J. Dev. Behav. Pediatr.2013,34, 508–522.
[87] Wormwood, K., Sokolowska, I., Ryan, J. P., Russell, S. et al., The potential for proteomics in understanding neurodevel-opmental disorders.J. Proteomics Bioinform. 2013,S5. [88] Darie, C. C., Biniossek, M. L., Gawinowicz, M. A., Milgrom,
Y. et al., Mass spectrometric evidence that proteolytic pro-cessing of rainbow trout egg vitelline envelope proteins takes place on the egg.J. Biol. Chem. 2005,280, 37585– 37598.
[89] Klemmer, P., Meredith, R. M., Holmgren, C. D., Klychnikov, O. I. et al., Proteomics, ultrastructure, and physiology of hip-pocampal synapses in a fragile X syndrome mouse model reveal presynaptic phenotype.J. Biol. Chem. 2011,286, 25495–25504.
[90] Liao, L., Park, S. K., Xu, T., Vanderklish, P., Yates, J. R., 3rd, Quantitative proteomic analysis of primary neurons reveals diverse changes in synaptic protein content in fmr1 knockout mice.Proc. Natl. Acad. Sci. USA2008,105, 15281– 15286.
[91] Monzo, K., Dowd, S. R., Minden, J. S., Sisson, J. C., Pro-teomic analysis reveals CCT is a target of Fragile X men-tal retardation protein regulation in Drosophila.Dev. Biol. 2010,340, 408–418.
[92] Zhang, Y. Q., Friedman, D. B., Wang, Z., Woodruff, E., 3rd et al., Protein expression profiling of the drosophila fragile X mutant brain reveals up-regulation of monoamine syn-thesis.Mol. Cell. Proteomics2005,4, 278–290.
[93] Zhang, Y. Q., Matthies, H. J., Mancuso, J., Andrews, H. K. et al., The Drosophila fragile X-related gene regulates ax-oneme differentiation during spermatogenesis.Dev. Biol. 2004,270, 290–307.
[94] Kaufmann, W. E., Cohen, S., Sun, H. T., Ho, G., Molecular phenotype of Fragile X syndrome: FMRP, FXRPs, and pro-tein targets.Microsc. Res. Tech.2002,57, 135–144. [95] Godler, D. E., Slater, H. R., Bui, Q. M., Ono, M. et al., FMR1
intron 1 methylation predicts FMRP expression in blood of female carriers of expanded FMR1 alleles.J. Mol. Diagn. 2011,13, 528–536.
[96] Godler, D. E., Slater, H. R., Bui, Q. M., Storey, E. et al., Fragile X mental retardation 1 (FMR1) intron 1 methylation in blood predicts verbal cognitive impairment in female carriers of expanded FMR1 alleles: evidence from a pilot study.Clin. Chem. 2012,58, 590–598.
[97] Aneja, A., Tierney, E., Autism: the role of cholesterol in treatment.Int. Rev. Psychiatry2008,20, 165–170.
[98] Bukelis, I., Porter, F. D., Zimmerman, A. W., Tierney, E., Smith-Lemli-Opitz syndrome and autism spectrum disor-der.Am. J. Psychiatry2007,164, 1655–1661.
[99] Diaz-Stransky, A., Tierney, E., Cognitive and behavioral as-pects of Smith-Lemli-Opitz syndrome.Am. J. Med. Genet. C Semin. Med. Genet. 2012,160C, 295–300.
[100] Tierney, E., Bukelis, I., Thompson, R. E., Ahmed, K. et al., Abnormalities of cholesterol metabolism in autism spec-trum disorders. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2006,141B, 666–668.
[101] DeBarber, A. E., Eroglu, Y., Merkens, L. S., Pappu, A. S., Steiner, R. D., Smith-Lemli-Opitz syndrome. Expert Rev. Mol. Med. 2011,13, e24.
[102] Fitzky, B. U., Moebius, F. F., Asaoka, H., Waage-Baudet, H. et al., 7-Dehydrocholesterol-dependent proteolysis of HMG-CoA reductase suppresses sterol biosynthesis in a mouse model of Smith-Lemli-Opitz/RSH syndrome.J. Clin. Invest. 2001,108, 905–915.
[103] Corso, G., Gelzo, M., Barone, R., Clericuzio, S. et al., Sterol profiles in plasma and erythrocyte membranes in patients with Smith-Lemli-Opitz syndrome: a six-year experience. Clin. Chem. Lab. Med. 2011,49, 2039–2046.
[104] Griffiths, W. J., Wang, Y., Karu, K., Samuel, E. et al., Po-tential of sterol analysis by liquid chromatography-tandem mass spectrometry for the prenatal diagnosis of Smith-Lemli-Opitz syndrome.Clin. Chem. 2008,54, 1317–1324. [105] Meljon, A., Watson, G. L., Wang, Y., Shackleton, C. H.,
Griffiths, W. J., Analysis by liquid chromatography-mass spectrometry of sterols and oxysterols in brain of the newborn Dhcr7(Delta3–5/T93M) mouse: a model of Smith-Lemli-Opitz syndrome. Biochem. Pharmacol. 2013, 86, 43–55.
[106] Paglia, G., D’Apolito, O., Gelzo, M., Dello Russo, A., Corso, G., Direct analysis of sterols from dried plasma/blood spots by an atmospheric pressure thermal desorption chemical ionization mass spectrometry (APTDCI-MS) method for a rapid screening of Smith-Lemli-Opitz syndrome.Analyst 2010,135, 789–796.
[107] Patti, G. J., Shriver, L. P., Wassif, C. A., Woo, H. K. et al., Nanostructure-initiator mass spectrometry (NIMS) imag-ing of brain cholesterol metabolites in Smith-Lemli-Opitz syndrome.Neuroscience2010,170, 858–864.
[108] Tu, C., Li, J., Jiang, X., Sheflin, L. G. et al., Ion current based proteomic profiling of the retina in a rat model of Smith-Lemli-Opitz syndrome.Mol. Cell Proteomics2013,12, 3583– 3598.
[109] Fliesler, S. J., Retinal degeneration in a rat model of Smith-Lemli-Opitz Syndrome: thinking beyond cholesterol defi-ciency.Adv. Exp. Med. Biol. 2010,664, 481–489.