Table I. Force Constants and Reference Values in the QMFF for Amides A. Bond length
E = 2K
b
(b - b0)2 + 3Kb
(b - b0)3 + 4Kb
(b - b0)4b0 2K
b
3Kb
4Kb
bond (Å) (kcal mol-1 Å-2) (kcal mol-1 Å-3) (kcal mol-1 Å-4) C'-O' 1.19201124.2 -2568.6
C'-N 1.3983 443.4
1415.3
C'-H 1.0934
-780.5 896.3
C-N 1.4416403.9-676.4 650.0
H*-N 1.0015
-1300.8 1908.3
C-C' 1.5103
-522.3 490.0
H-C 1.0845
-851.3 1040.0
C-C 1.5297
-586.1 758.0
B. Bond angle
E = 2K
q
(q - q0)2 + 3Kq
(q - q0)3 + 4Kq
(q - q0)4q0 2K
q
3Kq
4Kq
N-C'-O' 124.5 115.2 -41.0 -13.3
H-C'-O' 117.8 54.7 -26.8 0.0
H-C'-N 110.4 62.1 -22.0 0.0
C-N-C' 111.0 38.0 -8.0 -8.1
C'-N-H* 117.0 44.6 -9.7 0.0
C-N-H* 113.9 54.7 -23.9 0.0
C-C-N 114.3 50.8 -12.6 -11.1
H-C-N 108.9 68.3 3.5 0.0
C-C'-O 123.1 66.1 -20.5 0.2
C-C'-N 116.9 46.9 -13.1 -10.4
H*-N-H* 112.9 46.7 -11.3 0.0
C-C-C' 108.5 61.9 -11.3 -13.1
C'-C-H 107.7 48.3 -34.3 0.0
C-N-C 111.6 47.2 -10.2 -10.1
C'-C-N 100.6 62.0 -6.3 -12.7
N-C'-N 122.5 123.9 -43.7 -28.9
C'-N-C' 122.0 90.8 -31.3 -21.1
H-C-H 107.7 51.5 -8.9 -10.8
H-C-C 110.8 52.7 -10.9 -11.3
C. Torsion angle
E = 1K
f
(1 - cosf) + 2Kf
(1 - cos2f) + 3Kf
(1 - cos3f)1K
f
2Kf
3Kf
torsion (kcal mol-1) (kcal mol-1) (kcal mol-1)
O'-C'-N-C 0.830 3.723 -0.050
O'-C'-N-H* -1.882 3.043 -0.373
H-C'-N-C 0.334 2.584 -0.401
C-C-N-C' 0.020 -0.019 0.013
H-C-N-C' 0.031 -0.037 -0.058
C-C-N-H* -0.069 -0.011 -0.002
H-C-N-H* -0.021 -0.113 -0.175
C-C-C-N 0.108 0.080 -0.287
H-C-C-N -0.025 0.031 -0.207
C-C'-N-H* -0.915 2.385 -0.238
C-C-C'-O' 0.063 0.042 0.080
H-C-C'-O' -0.258 0.002 0.053
C-C-C'-N -0.053 0.056 -0.076
H-C-C'-N 0.241 -0.013 -0.098
C-C-C-C' 0.108 0.080 -0.287
C'-C-C-H -0.025 0.031 -0.207
C-C-N-C -0.002 -0.010 0.001
H-C-N-C 0.058 0.051 -0.236
C'-C-C-C' 0.108 0.080 -0.287
C'-C-C-N 0.108 0.080 -0.287
C'-C-N-C' -0.098 0.109 -0.088
C'-C-N-H* 0.078 0.108 0.104
N-C-C'-O' 0.128 0.174 0.129
N-C-C'-N -0.127 0.180 -0.126
C-C'-N-C -0.753 2.739 0.090
C'-C-N-C -0.005 0.007 0.006
N-C'-N-H* -0.818 0.516 -1.233
O'-C'-N-C' -0.452 1.390 -0.834
H-C'-N-C' 0.212 1.246 0.047
H-C-C-H -1.152 0.012 -0.179
2K
c
out-of-plane (kcal mol-1 rad-2)
H-C'-N-O' 42.126
C-N-C'-H* 1.250
C-C'-N-O' 44.242
C'-N-H*-H* 0.000
C-N-C-C' 4.247
N-C'-N-O' 49.203
C'-N-C'-H* 0.000
E. Van der Waals interaction
E = e [2(r*/r)9 - 3(r*/r)6] where r* = [(r
i
*6 + rj
*6) /2] 1/6and e = (e
i
ej) 1/2 2 (ri
*rj
*)3/(ri
*6 + rj
*6)r
i
* ei
atom (Å) (kcal mol-1)
H* 1.098 0.013
O' 3.535 0.267 C' 3.308 0.120
N 4.070 0.106
H 2.995 0.020
C 4.010 0.054
E = 332.0 q
i
qj
/ rij
; where qi
=d
ik
. Each dik
donates positive charge to the first atom listed and corresponding negative charge to the second atom.d
ik
bond (electrons)C'–O' 0.396
C–N 0.211
H*–N 0.440
C'–N 0.000
H–C' 0.046 C'–C 0.000
H–C 0.053
C–C 0.000
G. Bond/bond
E = K
bb'
(b - b0)(b' - b'0)K
bb'
bond/bond (kcal mol-1 Å-2)
O'-C'/C'-N 200.718
O'-C'/C'-H 61.122 N-C'/C'-H 4.058 C'-N/N-C 17.563 C'-N/N-H* -6.250 C-N/N-H* -5.030 N-C/C-C 5.137
C-C'/C'-O 66.766
C-C'/C'-N -9.389
N-H*/H*-N -0.820
C'-C/C-C 7.855
C'-C/C-H 1.031
N-C/C-N -2.171
N-C/C-C' -5.558
N-C'/C'-N 37.613
H-C/H-C 11.769
H-C/C-C 11.310
H. Bond/angle
E = K
bq
(b - b0)(q - q0)K
bq
bond/angle (kcal mol-1Å-1rad-1)
C'-O'/N-C'-O' 75.9
C'-N/N-C'-O' 90.9
C'-O'/H-C'-O' 71.1
C'-H/H-C'-O' 22.1
C'-N/H-C'-N 45.4
C'-H/H-C'-N 32.4
C'-N/C-N-C' 21.5
C-N/C-N-C' 5.5
C'-N/C'-N-H* 42.9
H*-N/C'-N-H* 15.7
C-N/C-N-H* 17.2
H*-N/C-N-H* 8.6
C-N/C-C-N 6.7
C-C/C-C-N -7.9
C-H/H-C-N 15.5
C'-O'/C-C'-O' 53.8
C-C'/C-C'-O' 50.7
C'-N/C-C'-N 8.8
C-C'/C-C'-N 36.8
H*-N/H*-N-H* 28.7
C-C'/C-C-C' 23.0
C-C/C-C-C' 26.3
C-C'/C'-C-H 18.1
C-H/C'-C-H 13.3
C-N/C-N-C -3.0
C-N/C'-C-N 25.2
C-C'/C'-C-N -7.5
C'-N/N-C'-N 98.7
C'-N/C'-N-C' 29.1
H-C/H-C-H 22.7
H-C/H-C-C 13.9
C-C/H-C-C 34.5
I. Angle/angle
E = K
qq'
(q - q0)( q' - q'0)K
C-C-N/H-C-N C-N -1.00
C-C-H/C-C-N C-C -4.88
H-C-N/H-C-N C-N 3.71
C-C-H/H-C-N C-H -1.29
C-C-C'/C'-C-H C-C' 2.96
C-C-C'/C-C-H C-C -2.64
C'-C-H/C'-C-H C-C' -5.07
C-C-H/C'-C-H C-H 1.57
C'-C-H/H-C-H C-H -4.91
C-C-N/C-C-N C-N -0.72
C-C-C/C-C-N C-C 0.22
C'-C-N/H-C-N C-N 4.39
C'-C-H/C'-C-N C-C' 0.08
C'-C-H/H-C-N C-H -2.39
C-C-C'/C'-C-N C-C' 8.24
C-C-N/C'-C-N C-N 5.76
C-C-C'/C-C-N C-C -1.23
H-C-H/H-C-H H-C 0.93
H-C-H/H-C-C H-C 1.57
H-C-C/H-C-C H-C 3.12
H-C-C/H-C-C C-C -2.18
J. Angle/angle/torsion
E = K
fqq’
(q - q0)( q' - q'0) cosfK
fqq
' angle/angle/torsion (kcal mol-1rad-2) N-C'-O'/C-N-C'/O'-C'-N-C -22.5H-C-N/C-N-H*/H-C-N-H* -9.7
C-C-C/C-C-N/C-C-C-N -1.5
C-C-H/C-C-N/H-C-C-N -18.5
C-C'-N/C'-N-H*/C-C'-N-H* -1.9 C-C-C'/C-C'-O'/C-C-C'-O' -11.6 C'-C-H/C-C'-O'/H-C-C'-O' -22.2 C-C-C'/C-C'-N/C-C-C'-N -7.9 C'-C-H/C-C'-N/H-C-C'-N -17.7 C-C-C/C-C-C'/C-C-C-C' -0.6 C-C-C'/C-C-H/C'-C-C-H -7.8
C-C-N/C-N-C/C-C-N-C -2.5
H-C-N/C-N-C/H-C-N-C -17.7
C-C-C'/C-C-C'/C'-C-C-C' 0.2
C-C-C'/C-C-N/C'-C-C-N 0.5
C'-C-N/C-N-C'/C'-C-N-C' -13.4 C'-C-N/C-N-H*/C'-C-N-H* -1.4 C'-C-N/C-C'-O'/N-C-C'-O' -9.5 C'-C-N/C-C'-N/N-C-C'-N -2.6 C-C'-N/C-N-C'/C-C'-N-C -9.5
C'-C-N/C-N-C/C'-C-N-C 0.2
N-C'-N/C'-N-H*/N-C'-N-H* -2.2 N-C'-O'/C'-N-C'/O'-C'-N-C' -4.9 H-C'-N/C'-N-C'/H-C'-N-C' -1.1
H-C-C/H-C-C/H-C-C-H -14.8
K. Bond/torsion(Type 1)
E = (b - b0) [1K
fb
cosf + 2Kfb
cos2f + 3Kfb
cos3f]1K
fb
2Kfb
3Kfb
bond/torsion (kcal mol-1Å-1) (kcal mol-1Å-1) (kcal mol-1Å-1)
C'-N/O'-C'-N-C -12.797 20.736 -2.568
C'-N/H-C'-N-C -16.993 4.645 -4.365
C'-N/H-C'-N-H* -1.399 7.542 -3.346
C-N/C-C-N-C' -5.725 -0.580 -0.985
C-N/H-C-N-C' -0.100 -3.282 1.678
C-N/C-C-N-H* -5.131 -4.908 0.051
C-N/H-C-N-H* -1.703 4.066 1.171
C-C/C-C-C-N -6.134 -4.786 -1.919
C-C/H-C-C-N -5.946 -0.861 -0.068
C'-N/C-C'-N-H* -0.768 6.863 -1.542
C-C'/C-C-C'-O' 0.491 -0.159 0.177
C-C'/H-C-C'-O' 0.342 1.324 1.390
C-C'/C-C-C'-N -2.907 -2.236 2.767
C-C'/H-C-C'-N 0.333 -0.601 1.160
C-C/C-C-C-C' -2.311 0.328 -1.002
C-C/C'-C-C-H -5.078 1.806 -1.103
C-N/C-C-N-C -6.822 -1.530 -4.325
C-N/H-C-N-C -3.323 0.472 1.400
C-C/C'-C-C-C' 2.139 0.848 -0.270
C-C/C'-C-C-N -1.770 -5.899 -4.924
C-N/C'-C-N-C' -6.875 -7.398 -7.869
C-N/C'-C-N-H* -2.104 5.859 -0.765
C-C'/N-C-C'-O' -6.527 7.099 0.619
C-C'/N-C-C'-N -6.327 -3.236 -7.968
C'-N/C-C'-N-C -13.408 4.941 -4.111
C-N/C'-C-N-C 1.828 -5.164 -4.479
C'-N/N-C'-N-H* -1.803 -6.429 3.201
C'-N/O'-C'-N-C' -0.162 -1.738 0.983
C'-N/H-C'-N-C' -0.699 -0.856 -1.197
C-C/H-C-C-H -16.582 -0.784 -0.901
L. Bond/torsion(type 2)
1K
fb'
2Kfb'
3Kfb'
bond/torsion (kcal mol-1Å-1) (kcal mol-1Å-1) (kcal mol-1Å-1)
C'-O'/O'-C'-N-C 0.178 -3.091 0.809
C-N/O'-C'-N-C 0.232 1.051 -0.146
C'-O'/O'-C'-N-H* -1.102 -3.831 1.807
H*-N/O'-C'-N-H 0.176 0.281 0.118
C'-H/H-C'-N-C -0.859 -0.007 0.215
C-N/H-C'-N-C -0.773 0.830 -0.094
C'-H/H-C'-N-H* -0.372 0.693 0.517
H*-N/H-C'-N-H* 0.143 0.808 0.170
C'-N/C-C-N-C' -0.180 -1.358 1.128
C-C/C-C-N-C' -0.295 0.005 0.081
C'-N/H-C-N-C' 0.332 1.700 -0.084
C-H/H-C-N-C' -0.531 1.188 0.194
H*-N/C-C-N-H* -0.144 -0.105 -0.600
C-C/C-C-N-H* 0.191 0.002 0.192
H*-N/H-C-N-H* -0.709 0.238 0.450
C-H/H-C-N-H* -1.302 0.410 0.128
C-N/C-C-C-N -0.115 -0.059 0.037
C-C/C-C-C-N 0.108 0.015 0.075
C-N/H-C-C-N 0.438 0.364 0.673
C-H/H-C-C-N -0.087 -0.545 -0.272
C-C'/C-C'-N-H* -1.011 0.814 0.610
H*-N/C-C'-N-H* 0.929 0.243 0.209
C'-O'/C-C-C'-O' -0.407 0.118 -0.221
C-C/C-C-C'-O' 0.385 0.073 0.152
C'-O'/H-C-C'-O' -0.333 0.051 0.558
C-H/H-C-C'-O' 1.760 0.410 0.568
C'-N/C-C-C'-N -0.399 -0.443 -0.256
C-C/C-C-C'-N -0.381 -0.011 -0.166
C-H/H-C-C'-N -0.039 1.136 0.005
C-C'/C-C-C-C' 0.008 0.009 -0.001
C-C/C-C-C-C' 0.009 0.000 0.005
C-C'/C'-C-C-H -0.030 0.526 -0.642
C-H/C'-C-C-H -0.014 -0.046 -0.109
C-N/C-C-N-C -0.119 -0.011 -0.135
C-C/C-C-N-C -0.181 -0.164 -0.112
C-N/H-C-N-C 0.181 0.470 -0.108
C-H/H-C-N-C -1.954 1.159 0.979
C-C'/C'-C-C-C' 0.008 -0.001 -0.006
C-N/C'-C-C-N -0.244 -0.214 -0.214
C-C'/C'-C-C-N -0.191 -0.161 -0.168
C'-N/C'-C-N-C' -0.645 0.617 -0.631
C-C'/C'-C-N-C' -0.259 0.311 -0.406
H*-N/C'-C-N-H* 1.305 0.430 0.085
C-C'/C'-C-N-H* 0.158 0.487 0.134
C-N/N-C-C'-O' -0.110 0.379 0.234
C'-O'/N-C-C'-O' 1.456 -0.479 0.945
C-N/N-C-C'-N -0.252 0.104 -0.865
C'-N/N-C-C'-N -0.018 0.028 -1.389
C-C'/C-C'-N-C 0.135 -0.671 0.418
C-N/C-C'-N-C 0.333 -0.165 -0.206
C-N/C'-C-N-C 0.026 -0.131 -0.117
C-C'/C'-C-N-C 0.097 -0.115 -0.134
H*-N/N-C'-N-H* -0.707 -1.033 0.191
C'-N/N-C'-N-H* 0.112 -0.767 -0.006
C'-O'/O'-C'-N-C' -1.017 1.204 -0.996
C'-N/O'-C'-N-C' 0.250 -0.699 0.386
C'-N/H-C'-N-C' -0.306 -0.609 -0.330
M. Angle/torsion
E = (q - q0) [1K
fq
cosf + 2Kfq
cos2f + 3Kfq
cos3f]1K
fq
2Kfq
3Kfq
angle/torsion (kcal mol-1rad-1) (kcal mol-1rad-1) (kcal mol-1rad-1)
N-C'-O'/O'-C'-N-C 6.444 5.843 2.482
C-N-C'/O'-C'-N-C 10.787 3.117 -0.320
N-C'-O'/O'-C'-N-H* -3.803 0.523 0.793
C'-N-H*/O'-C'-N-H* 6.115 1.803 0.359
H-C'-N/H-C'-N-C 0.197 4.539 0.310
C-N-C'/H-C'-N-C 8.960 -0.511 -0.311
H-C'-N/H-C'-N-H* -3.163 2.963 -0.211
C'-N-H*/H-C'-N-H* 5.694 1.463 -0.046
C-N-C'/C-C-N-C' -2.207 1.637 1.039
C-C-N/C-C-N-C' -1.095 0.082 1.765
C-N-C'/H-C-N-C' 0.054 -0.495 -0.112
H-C-N/H-C-N-C' -2.197 3.012 0.777
C-N-H*/C-C-N-H* -1.489 0.523 -0.355
C-C-N/C-C-N-H* -4.198 3.925 -0.054
C-N-H*/H-C-N-H* -0.992 0.523 0.114
H-C-N/H-C-N-H* -5.365 2.011 0.347
C-C-N/C-C-C-N 0.295 0.232 -1.152
C-C-C/C-C-C-N -0.797 -2.461 0.360
C-C-N/H-C-C-N -2.872 0.337 -0.569
C-C-H/H-C-C-N -1.807 2.454 -1.751
C-C'-N/C-C'-N-H* -3.208 1.871 1.410
C'-N-H*/C-C'-N-H* 4.950 0.540 0.143
C-C'-O'/C-C-C'-O' 0.978 0.864 0.975
C-C-C'/C-C-C'-O' 0.128 -1.986 -0.790
C-C'-O'/H-C-C'-O' -2.166 1.059 -0.302
C-C'-N/C-C-C'-N 3.076 0.729 -0.111
C-C-C'/C-C-C'-N 3.160 -0.049 -2.002
C-C'-N/H-C-C'-N 2.900 0.735 1.218
C'-C-H/H-C-C'-N 10.283 0.011 1.001
C-C-C'/C-C-C-C' 0.075 -0.098 -0.069
C-C-C/C-C-C-C' -0.378 0.464 -0.331
C-C-C'/C'-C-C-H -1.082 -1.369 -0.916
C-C-H/C'-C-C-H 0.023 2.060 -2.042
C-N-C/C-C-N-C 1.874 2.990 -0.223
C-C-N/C-C-N-C -0.417 2.571 0.397
C-N-C/H-C-N-C 0.513 -0.577 -0.283
H-C-N/H-C-N-C -2.541 1.884 0.733
C-C-C'/C'-C-C-C' 5.002 1.813 -0.065
C-C-N/C'-C-C-N -2.292 -1.202 -2.330
C-C-C'/C'-C-C-N -1.959 -3.449 -0.383
C-N-C'/C'-C-N-C' -3.491 1.270 -1.604
C'-C-N/C'-C-N-C' 6.929 2.668 1.883
C-N-H*/C'-C-N-H* 0.159 -0.812 -3.753
C'-C-N/C'-C-N-H* -5.324 2.945 0.548
C'-C-N/N-C-C'-O' -2.636 -4.079 3.410
C-C'-O'/N-C-C'-O' 1.425 5.477 7.985
C'-C-N/N-C-C'-N -1.188 -1.194 3.564
C-C'-N/N-C-C'-N -0.065 0.924 -9.223
C-C'-N/C-C'-N-C 6.106 4.247 4.769
C-N-C'/C-C'-N-C 8.574 2.588 0.587
C-N-C/C'-C-N-C -0.650 3.641 3.724
C'-C-N/C'-C-N-C 5.071 -5.186 -3.202
C'-N-H*/N-C'-N-H* 1.178 -1.183 0.899
N-C'-N/N-C'-N-H* -3.877 2.284 0.106
N-C'-O'/O'-C'-N-C' -2.282 3.478 -0.413
C'-N-C'/O'-C'-N-C' -0.440 -0.079 -0.462
C'-N-C'/H-C'-N-C' 2.356 -1.832 0.859
[image:15.612.69.542.192.721.2]H-C-C/H-C-C-H -1.622 0.739 -0.449
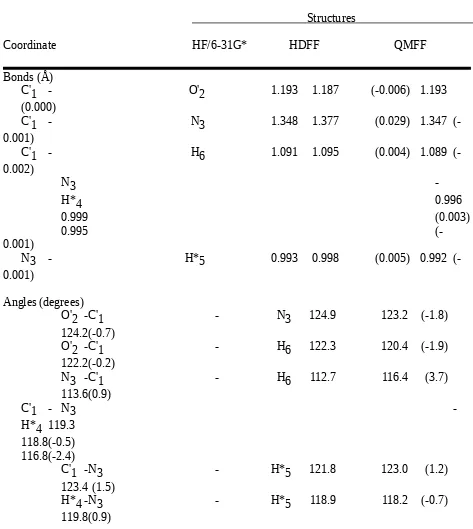
Table II.A. Comparison of Formamide Structures Optimized by Ab Initio Calculation (HF/6-31G*) and by the HDFF and QMFF Force Fields.
Structures
Coordinate HF/6-31G* HDFF QMFF
Bonds (Å)
C'1 - O'2 1.193 1.187 (-0.006) 1.193
(0.000)
C'1 - N3 1.348 1.377 (0.029) 1.347
(-0.001)
C'1 - H6 1.091 1.095 (0.004) 1.089
(-0.002)
N3
-H*4 0.996
0.999 (0.003)
0.995
(-0.001)
N3 - H*5 0.993 0.998 (0.005) 0.992
(-0.001)
Angles (degrees)
O'2 -C'1 - N3 124.9 123.2 (-1.8)
124.2(-0.7)
O'2 -C'1 - H6 122.3 120.4 (-1.9)
122.2(-0.2)
N3 -C'1 - H6 112.7 116.4 (3.7)
113.6(0.9)
C'1 - N3
-H*4 119.3 118.8(-0.5) 116.8(-2.4)
C'1 -N3 - H*5 121.8 123.0 (1.2)
123.4 (1.5)
H*4 -N3 - H*5 118.9 118.2 (-0.7)
Torsions (degrees)
O'2 - C'1 - N3 - H*4 0.0 0.0 (0.0) 0.0
(0.0)
O'2 - C'1 - N3 - H*5 180.0 180.0 (0.0)
180.0 (0.0)
H6
-C'1
-N3
-H*4 180.0 180.0 (0.0) 180.0 (0.0)
H6 - C'1 - N3 - H*5 0.0 0.0 (0.0) 0.0
(0.0)
Out-of-Planes (degrees)
O'2 - C'1 - N3 - H6 0.0 0.0 (0.0) 0.0
(0.0)
C'1 - N3 - H*4 - H*5 0.0 0.0 (0.0) 0.0
(0.0)
Table II.B. Comparison of Planar Acetamide Structures Optimized by Ab Initio Calculation (HF/6-31G*) and by the HDFF and QMFF Force Fields.
Structures
Coordinate HF/6-31G* HDFF QMFF
Bonds (Å)
O'1 - C'2
1.198 1.187
(-0.011) 1.196
(-0.002)
C'2 - N3
1.356 1.369
[image:16.612.70.542.84.442.2](-0.004)
C'2 - C4
1.514 1.509
(-0.005) 1.503
(-0.011)
N3
-H*5 0.995
0.998 (0.003)
0.995 (0.000)
N3
-H*6 0.993
0.996 (0.003)
0.990
(-0.002)
C4 - H7
1.085 1.085
(0.000) 1.084
(-0.001)
C4 - H8
1.085 1.085
(0.000) 1.084
(-0.001)
C4 - H9
1.080 1.085
(0.005) 1.079
(-0.001) Angles (degrees)
O'1 - C'2
-N3 122.2 121.3 (-0.9) 121.4 (-0.8)
O'1 - C'2
-C4 122.9 123.4 (0.5) 123.0 (0.1)
N3 - C'2
-C4 114.9 115.3(0.4) 115.7(0.7)
C'2 - N3
118.8(0.3) 116.5(-2.1)
C'2 - N3
-H*6 122.7
122.6 (-0.1) 123.5 (0.8)
H*5 - N3
-H*6 118.7 118.6(-0.1) 120.1 (1.3)
C'2 - C4
-H7 110.6 109.4 (-1.2) 110.7(0.1)
C'2 - C4
-H8 110.6 109.4 (-1.2) 110.7(0.1)
C'2 - C4
-H9 108.9 111.2(2.3) 109.3 (0.4)
H7 - C4
-H8 108.0 106.9 (-1.1) 108.9 (0.9)
H7 - C4
-H9 109.4 109.9 (0.5) 108.6 (-0.8)
H8 - C4
-H9 109.4 109.9 (0.5) 108.6 (-0.8) Torsions (degrees)
O'1
-C'2
-N3
0.0
(0.0) 0.0
(0.0)
O'1
-C'2
-N3
-H*6 180.0 180.0 (0.0) 180.0 (0.0)
C4
-C'2
-N3
-H*5 180.0 180.0 (0.0) 180.0 (0.0)
C4
-C'2
-N3
-H*6 0.0
0.0
(0.0) 0.0
(0.0)
O'1
-C'2
-C4
-H7 120.2 121.6 (1.4)
119.6
(-0.6)
O'1
-C'2
-C4
238.4 (-1.4) 240.4
(0.6)
O'1
-C'2
-C4
-H9 0.0
0.0
(0.0) 0.0
(0.0)
N3
-C'2
-C4
-H7 300.2 301.6 (1.4)
299.6
(-0.6)
N3
-C'2
-C4
-H8 59.8
58.4
(-1.4) 60.4 (0.6)
N3
-C'2
-C4
-H9 180.0 180.0 (0.0) 180.0 (0.0) Out-of-Planes (degrees)
O'1
-C'2
-C4
0.0
(0.0) 0.0
(0.0)
C'2
-N3
-H*6
-H*5 0.0
0.0
(0.0) 0.0
(0.0)
Appendix II.C. Comparison of Planar N-Methylformamide Trans and Cis Structures Optimized by Ab Initio Calculation (HF/6-31G*) and by the HDFF and QMFF Force Fields.
trans cis Coordinate HF/6-31G* HDFF QMFF HF/6-31G* HDFF QMFF
Bonds (Å)
O'1 -C'2 1.196 1.190 (-0.006) 1.196 (0.001) 1.195 1.189 (-0.006) 1.194 (-0.001) C'2 -N3 1.345 1.401 (0.056) 1.348 (0.003) 1.347 1.389 (0.041) 1.349 (0.002) C'2 -H5 1.091 1.094 (0.003) 1.088 (-0.003) 1.091 1.095 (0.004) 1.092 (0.000) N3 -C4 1.448 1.452 (0.004) 1.445 (-0.003) 1.442 1.450 (0.007) 1.440 (-0.002) N3 -H*6 0.993 0.999 (0.006) 0.995 (0.002) 0.997 1.001 (0.004) 1.000 (0.003) C4 -H7 1.081 1.085 (0.004) 1.083 (0.002) 1.082 1.086 (0.003) 1.082 (0.000) C4 -H8 1.083 1.086 (0.003) 1.085 (0.002) 1.084 1.086 (0.001) 1.085 (0.001) C4 -H9 1.083 1.086 (0.003) 1.085 (0.002) 1.084 1.086 (0.001) 1.085 (0.001)
Angles (degrees)
O'1 -C'2 -N3 124.8 125.8 (1.0) 124.6 (-0.2) 124.9 123.4 (-1.5) 125.1 (0.2) O'1 -C'2 -H5 122.2 119.6 (-2.7) 121.8 (-0.4) 122.1 120.0 (-2.2) 121.9 (-0.2) N3 -C'2 -H5 113.0 114.6 (1.6) 113.6 (0.6) 112.9 116.6 (3.7) 113.0 (0.0) C'2 -N3 -C4 121.8 122.5 (0.7) 120.8 (-0.9) 125.4 124.5 (-0.9) 124.5 (-0.9) C'2 -N3 -H*6 118.7 120.0 (1.3) 120.7 (2.0) 115.5 117.1 (1.6) 115.4 (-0.1) C4 -N3 -H*6 119.6 117.5 (-2.0) 118.5 (-1.1) 119.1 118.4 (-0.7) 120.1 (1.0) N3 -C4 -H7 108.7 110.2 (1.5) 109.0 (0.3) 109.3 110.7 (1.5) 108.3 (-1.0) N3 -C4 -H8 110.9 110.7 (-0.2) 111.1 (0.2) 111.4 110.5 (-0.8) 111.4 (0.0) N3 -C4 -H9 110.9 110.7 (-0.2) 111.1 (0.2) 111.3 110.5 (-0.8) 111.4 (0.1) H7 -C4 -H8 109.0 107.9 (-1.1) 108.4 (-0.5) 108.1 107.9 (-0.2) 108.5 (0.4) H7 -C4 -H9 109.0 107.9 (-1.1) 108.4 (-0.5) 108.1 107.9 (-0.1) 108.5 (0.4) H8 -C4 -H9 108.4 109.3 (1.0) 108.7 (0.3) 108.7 109.1 (0.5) 108.7 (0.0)
Torsions (degrees)
H*6 -N3 -C4 -H7 0.0 0.0 (0.0) 0.0 (0.0) 179.9 180.0 (0.1) 180.0 (0.1) H*6 -N3 -C4 -H8 119.8119.3 (-0.5) 119.4 (-0.3) 299.2 299.5 (0.4) 299.2 (0.1) H*6 -N3 -C4 -H9 240.2240.7 (0.5) 240.6 (0.3) 60.6 60.5 (-0.2) 60.8 (0.1)
Out-of-Planes (degrees) O’ '1-C'2 -N3 -H5 0.0 0.0
0.0 (0.0)
0.0 (0.0 )0.0 (0.0) C'2 -N3 -H*6 -C4 0.0 0.0
0.0 (0.0)
0.0 (0.0 )0.0 (0.0)
Appendix II.D. Comparison of Planar N-Methylacetamide Trans and Cis Structures Optimized by Ab Initio Calculation (HF/6-31G*) and by the HDFF and QMFF Force Fields.
trans cis Coordinate HF/6-31G* HDFF QMFF HF/6-31G* HDFF QMFF Bonds (Å)
C1 - C'2 1.514 1.514 (0.000) (-0.001) 1.514 1.515 (0.001) 1.512 (-0.001)
C1 - H7 1.083 1.085 (0.002) 1.084 (0.001) 1.085 1.084 0.001) 1.084 (-0.001)
C1 - H8 1.084 1.085 (0.001) (-0.002) 1.080 1.085 (0.005) 1.079 (-0.001)
C1 - H9 1.083 1.085 (0.002) (0.001) 1.085 1.084 (-0.001) 1.084 (-0.001)
C'2 -N3 1.350 1.398 (0.048) 1.351 (0.001) 1.357 1.389 (0.032) 1.358 (0.001) C'2 -O'5 1.201 1.190 (-0.011) 1.202 (0.001) 1.200 1.189(-0.011) 1.197 (-0.003) N3 - C4 1.447 1.452 (0.005) 1.449 (0.002) 1.445 1.450 (0.005) 1.444 (-0.001)
N3 - H*6 0.992 0.998 (0.006) (0.002) 0.995 1.000 (0.005) 0.999 (0.004)
C4 - H10 1.083 1.086 (0.003) (0.002) 1.085 1.086 (0.001) 1.085 (0.000)
C4 - H11 1.082 1.085 (0.003) 1.083 (0.001) 1.081 1.085 (0.004) 1.084 (0.003)
C4 - H12 1.083 1.086 (0.003) (0.002) 1.085 1.086 (0.001) 1.085 (0.000) Angles (degrees)
N3 - C4 - H12 111.0 110.7 (-0.3) 111.1 (0.1) 112.2 110.9 (-1.3) 111.5 (-0.7) H10 - C4 - H11 109.0 107.9 (-1.1) 108.4 (-0.6) 107.8 107.7 (-0.1) 108.2 (0.4)
H10 - C4 - H12 108.3 109.3 (1.0) 108.8 (0.5) 108.5 109.6 (1.1) 109.0 (0.5)
H11- C4 - H12 109.0 107.9 (-1.1) 108.4 (-0.6) 107.8 107.7 (-0.1) 108.2 (0.4) Torsions (degrees)
H7 - C1 - C'2 - N3 238.4 238.5 (0.1) 238.7 (0.3) 300.0 301.6 299.2 (-0.8)
H7 - C1 - C'2 - O'5 58.4 58.5 (0.1) 58.7 (0.3) 120.0 121.6 119.2 (-0.8)
H8 - C1 - C'2 - N3 0.0 0.0 (0.0) 0.0 (0.0) 180.0 180.0 180.0 (0.0)
H8 - C1 - C'2 - O'5 180.0 180.0 (0.0) 180.0 (0.0) 0.0 0.0(0.0) 0.0 (0.0)
H9 - C1 - C'2 - N3 121.6 121.5 (-0.1) 121.3 (-0.3) 60.0 58.4(-1.6)60.8 (0.8)
H9 - C1 - C'2 - O'5 301.6 301.5 (-0.1) 301.3 (-0.3) 240.0 238.4 240.8 (0.8)
C1 - C'2- N3 - C4 180.0 180.0 (0.0) 180.0 (0.0) 0.0 0.0 (0.0)0.0 (0.0) C1 - C' - N3 - H*6 0.0 0.0 (0.0) 0.0 (0.0) 180.0 180.0 (0.0)180.0(0.0) O'5- C'2 - N3 - C4 0.0 0.0 (0.0) 0.0 (0.0) 180.0 180.0 (0.0) 180.0 O'5- C'2 - N3 - H*6 180.0 180.0 (0.0) 180.0 (0.0) 0.0 0.0 (0.0) 0.0 (0.0) C'2 - N3 - C4 - H10 299.8 299.3 (-0.5) 299.4 (-0.4) 298.8 299.0 290.0 (0.2)
C'2 - N3 - C4 - H11 180.0 180.0 (0.0) 180.0 (0.0) 180.0 180.0 180.0 (0.0)
C'2 - N3 - C4 - H12 60.2 60.7 (0.5) 60.6 (0.4) 61.2 61.0(-0.2)61.0 (0.2)
H*6 - N3 - C4 - H11 0.0 0.0 (0.0) 0.0 (0.0) 0.0 0.0(0.0) 0.0 0.0)
H*6 - N3 - C4 - H12 240.2 240.7 (0.5) 240.6 (0.4) 241.2 241.0 241.0 (-0.2)
Out-of-Planes
[image:23.612.94.542.516.712.2]C1 - C'2-N3 - O'5 0.0 0.0 (0.0) 0.0 (0.0) 0.0 0.0 (0.0)0.0 (0.0) C'2 - N3 - H*6 -C4 0.0 0.0 (0.0) 0.0 (0.0) 0.0 0.0 (0.0)0.0 (0.0) ______________________________________________________________________________________________________________
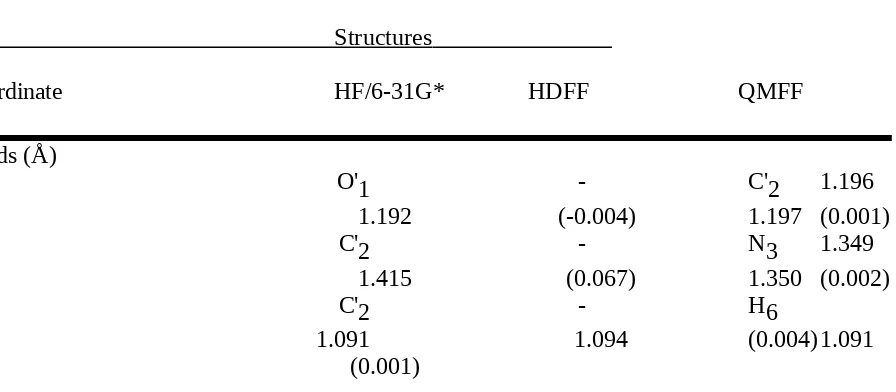
Table II.E. Comparison of N,N-Dimethylformamide Structures Optimized by Ab Initio Calculation (HF/6-31G*) and by the HDFF and QMFF Force Fields.
Structures
Coordinate HF/6-31G* HDFF QMFF
Bonds (Å)
O'1 - C'2 1.196
1.192 (-0.004) 1.197 (0.001)
C'2 - N3 1.349
1.415 (0.067) 1.350 (0.002)
C'2 - H6
1.091 1.094 (0.004)1.091
N3 - C4 1.442
1.454 (0.012) 1.447 (0.005)
N3 - C5 1.446
1.457 (0.011) 1.448 (0.002)
C4 - H7 1.082
1.085 (0.003) 1.080
(-0.002)
C4 - H8 1.086
1.086 (0.000) 1.087 (0.001)
C4 - H9 1.086
1.086 (0.000) 1.087 (0.001)
C5 - H10 1.077
1.085 (0.008) 1.080 (0.003)
C5 - H11 1.086
1.085 (-0.001) 1.087 (0.001)
C5 - H12 1.086
1.085 (-0.001) 1.087 (0.001)
Angles (degrees)
O'1 - C'2 - N3
125.9 126.1 (0.2)
126.2 (0.3)
O'1 - C'2 - H6
121.5 118.8 (-2.7)
120.9 (-0.6)
N3 - C'2 - H6
112.6 115.0 (2.5)
112.9 (0.3)
C'2 - N3 - C4
122.0 121.3 (-0.7)
121.4 (-0.6)
C'2 - N3 - C5
120.6 122.9 (2.3)
122.1 (1.5)
C4 - N3 - C5
117.4 115.8 (-1.6)
116.6 (-0.9)
N3 - C4 - H7
110.2 111.4 (1.3)
109.3 (-0.9)
N3 - C4 - H8
110.9 110.4 (-0.5)
N3 - C4 - H9
110.9 110.4 (-0.5)
111.1 (0.2)
H7 - C4 - H8
108.3 108.4 (0.1)
108.3 (0.1)
H7 - C4 - H9
108.3 108.4 (0.1)
108.3 (0.1)
H8 - C4 - H9
108.2 107.6 (-0.6)
108.6 (0.3)
N3 - C5
-H10 109.0 111.7
(2.8) 109.9 (0.9)
N3 - C5
-H11 110.5 110.4
(-0.1) 111.0 (0.6)
N3 - C5
-H12 110.5 110.4
(-0.1) 111.0 (0.6)
H10 - C5
-H11 109.3 108.3
(-1.0) 108.1 (-1.1)
H10 - C5
-H12 109.3 108.3
(-1.0) 108.1 (-1.1)
H11 - C5
-H12 108.4 107.7
(-0.7) 108.5 (0.1)
Torsions (degrees)
O'1 - C'2 - N3
- C4 180.0
180.0 (0.0) 180.0
(0.0)
O'1 - C'2 - N3
- C5 0.0
0.0 (0.0) 0.0
(0.0)
- C4 0.0
0.0 (0.0) 0.0
(0.0)
H6 - C'2 - N3
- C5 180.0
180.0 (0.0) 180.0
(0.0)
C'2 - N3 - C4
- H7 0.0
0.0 (0.0) 0.0
(0.0)
C'2 - N3 - C4
- H8 240.1
239.4 (-0.7) 240.5
(0.4)
C'2 - N3 - C4
- H9 119.9
120.6 (0.7) 119.5
(-0.4)
C5 - N3 - C4
- H7 180.0
180.0 (0.0) 180.0
(0.0)
C5 - N3 - C4
- H8 60.1
59.4 (-0.7) 60.5
(0.4)
C5 - N3 - C4
- H9 299.9
300.6 (0.7) 299.5
(-0.4)
C'2 - N3 - C5
- H10 0.0
0.0 (0.0) 0.0
(0.0)
C'2 - N3 - C5
- H11 120.0
120.5 (0.5) 119.6
(-0.5)
C'2 - N3 - C5
- H12 240.0
(0.5)
C4 - N3 - C5
- H10 180.0
180.0 (0.0) 180.0
(0.0)
C4 - N3 - C5
- H11 300.0
300.5 (0.5) 299.6
(-0.5)
C4 - N3 - C5
- H12 60.0
59.5 (-0.5) 60.4
(0.5) Out-of-Planes (degrees)
O'1 - C'2 - N3
- H6 0.0
0.0 (0.0) 0.0
(0.0)
C'2 - N3 - C4
- C5 0.0
0.0 (0.0) 0.0
[image:27.612.72.540.69.436.2] [image:27.612.70.540.536.718.2](0.0)
Table II.F. Comparison of Planar N,N-Dimethylacetamide Structures Optimized by Ab Initio Calculation (HF/6-31G*) and by the HDFF and QMFF Force Fields.
Structures
Coordinate HF/6-31G* HDFF QMFF
Bonds (Å)
C1 - C'2 1.518
1.520 (0.002) 1.521
(0.004)
C1 - H7 1.079
1.085 (0.006) 1.080
(0.001)
1.084 (0.000) 1.084 (-0.001)
C1 - H9 1.085
1.084 (0.000) 1.084
(-0.001)
C'2 - O'3 1.203
1.192 (-0.010) 1.201
(-0.001)
C'2 - N4
1.361 1.430 (0.069)
1.366 (0.006)
N4 - C5
1.449 1.461 (0.012)
1.455 (0.006)
N4 - C6 1.445
1.459 (0.014) 1.450
(0.005)
C5 - H10
1.076 1.085 (0.009)
1.079 (0.004)
C5 - H11 1.087
1.085 (-0.001) 1.087
(0.001)
C5 - H12 1.087
1.085 (-0.001) 1.087
(0.001)
C6 - H13 1.077
1.083 (0.006) 1.078
(0.001)
C6 - H14 1.087
1.086 (-0.001) 1.088
(0.001)
C6 - H15 1.087
1.086 (-0.001) 1.088
(0.001) Angles (degrees)
C'2 - C1 - H7
107.2 111.4 (4.2)
108.1 (0.9)
C'2 - C1 - H8
111.6 109.5 (-2.1)
111.4 (-0.3)
111.6 109.5 (-2.1)
111.4 (-0.3)
H7 - C1 - H8
109.1 109.5 (0.5)
108.3 (-0.8)
H7 - C1 - H9
109.1 109.5 (0.5)
108.3 (-0.8)
H8 - C1 - H9
108.1 107.3 (-0.9)
109.3 (1.1)
C1 - C'2 - O'3
119.9 119.0 (-0.9)
119.0 (-0.8)
C1 - C'2 - N4
118.0 118.3 (0.3)
118.8 (0.8)
O'3 - C'2 - N4
122.1 122.1 (0.6)
122.2 (0.1)
C'2 - N4 - C5
119.5 121.4 (1.8)
120.9 (1.4)
C'2 - N4 - C6
125.6 124.8 (-0.8)
124.9 (-0.7)
C5 - N4 - C6
114.9 113.9 (-1.0)
114.2 (-0.6)
N4 - C5
-H10 109.5 112.4
(3.0) 110.8 (1.4)
N4 - C5
-H11 110.3 110.3
(0.0) 110.8 (0.4)
N4 - C5
-H12 110.3 110.3
(0.0) 110.8 (0.4)
H10 - C5
-H11 109.3 108.0
(-1.2) 107.9 (-1.3)
-H12 109.3 108.0
(-1.2) 107.9 (-1.3)
H11 - C5
-H12 108.2 107.5
(-0.6) 108.5 (0.3)
N4 - C6
-H13 111.8 113.0
(1.1) 111.1 (-0.7)
N4 - C6
-H14 110.4 110.2
(-0.2) 110.6 (0.3)
N4 - C6
-H15 110.4 110.2
(-0.2) 110.6 (0.3)
H13 - C6
-H14 108.1 108.0
(-0.1) 108.0 (-0.1)
H13 - C6
-H15 108.1 108.0
(-0.1) 108.0 (-0.1)
H14 - C6
-H15 107.9 107.4
(-0.6) 108.4 (0.5)
Torsions (degrees)
H7 - C1 - C'2
- O'3 0.0
0.0 (0.0) 0.0
(0.0)
H7 - C1 - C'2
- N4 180.0
180.0 (0.0) 180.0
(0.0)
H8 - C1 - C'2
- O'3 119.4
121.3 (1.9) 118.9
(-0.6)
H8 - C1 - C'2
- N4 299.4
301.3 (1.9) 298.9
H9 - C1 - C'2
- O'3 240.6
238.7 (-1.9) 241.1
(0.6)
H9 - C1 - C'2
- N4 60.6
58.7 (-1.9) 61.1
(0.6)
C1 - C'2 - N4
- C5 180.0
180.0 (0.0) 180.0
(0.0)
C1 - C'2 - N4
- C6 0.0
0.0 (0.0) 0.0
(0.0)
O'3 - C'2 - N4
- C5 0.0
0.0 (0.0) 0.0
(0.0)
O'3 - C'2 - N4
- C6 180.0
180.0 (0.0) 180.0
(0.0)
C'2 - N4 - C5
- H10 0.0
0.0 (0.0) 0.0
(0.0)
C'2 - N4 - C5
- H11 120.3
120.7 (0.4) 119.8
(-0.5)
C'2 - N4 - C5
- H12 239.7
239.3 (-0.4) 240.2
(0.5)
C6 - N4 - C5
- H10 180.0
180.0 (0.0) 180.0
(0.0)
- H11 300.3
300.7 (0.4) 299.8
(-0.5)
C6 - N4 - C5
- H12 59.7
59.3 (-0.4) 60.2
(0.5)
C'2 - N4 - C6
- H13 0.0
0.0 (0.0) 0.0
(0.0)
C'2 - N4 - C6
- H14 120.4
120.9 (0.5) 119.9
(-0.5)
C'2 - N4 - C6
- H15 239.6
239.1 (-0.5) 240.1
(0.5)
C5 - N4 - C6
- H13 180.0
180.0 (0.0) 180.0
(0.0)
C5 - N4 - C6
- H14 300.4
300.9 (0.5) 299.9
(-0.5)
C5 - N4 - C6
- H15 59.6
59.1 (-0.5) 60.1
(0.5) Out-of-Planes (degrees)
C1 - C'2 - O'3
- N4 0.0 0.0
(0.0) 0.0 (0.0)
C'2 - N4 - C5
- C6 0.0 0.0
Table II.G. Comparison of Urea Structures Optimized by Ab Initio Calculation (HF/6-31G*) and by the HDFF and QMFF Force Fields.
Structures
Coordinate HF/6-31G* HDFF QMFF
Bonds (Å)
N1 - C'2
1.373 1.308
(-0.065) 1.372
(-0.001)
N1 - H*5
0.996 0.996
(0.000) 0.996
(0.000)
N1 - H*6
0.996 0.994
(-0.003) 0.993
(-0.003)
C'2 - O'3
1.197 1.180
(-0.017) 1.192
(-0.005)
C'2 - N4
1.373 1.308
(-0.065) 1.372
(-0.001)
N4 - H*7
0.996 0.996
(0.000) 0.996
(0.000)
N4 - H*8
0.996 0.994
(-0.003) 0.993
(-0.003) Angles (degrees)
C'2 - N1
- H*5
113.4 112.3
C'2 - N1
- H*6
117.7 114.5
(-3.3) 117.3 (-0.4)
H*5 - N1
- H*6
114.4 116.2
(1.7) 116.2
(1.8)
N1 - C'2
- O'3
123.0 122.5
(-0.5) 121.9 (-1.1)
N1 - C'2
- N4
114.0 115.0
(1.1) 116.2
(2.2)
O'3 - C'2
- N4
123.0 122.5
(-0.5) 121.9 (-1.1)
C'2 - N4
- H*7
113.4 112.3
(-1.0) 109.3 (-4.1)
C'2 - N4
- H*8
117.7 114.5
(-3.3) 117.3 (-0.4)
H*7 - N4
- H*8
114.4 116.2
(1.7) 116.2
(1.8) Torsions (degrees)
H*5 - N1
- C'2
-O'3 12.6
13.9 (1.3) 9.2
H*5 - N1
- C'2
-N4 192.6
193.9 (1.3)
189.2 (-3.4)
H*6 - N1
- C'2
-O'3 150.0
149.2 (-0.8)
144.2 (-5.8)
H*6 - N1
- C'2
-N4 330.0
329.2 (-0.8)
324.2 (-5.8)
N1 - C'2
- N4
-H*7 192.6
193.9 (1.3)
189.2 (-3.4)
N1 - C'2
- N4
-H*8 330.0
329.2 (-0.8)
324.2 (-5.8)
O'3 - C'2
- N4
-H*7 12.6
13.9 (1.3) 9.2
(-3.4)
O'3 - C'2
- N4
-H*8 150.0
149.2 (-0.8)
144.2 (-5.8)
Out-of Planes (degrees)
C'2 - N1
- H*6
-H*5 37.4
(-3.3)
N1 - C'2
- O'3
-N4 0.0 0.0
(0.0) 0.0
(0.0)
C'2 - N4
- H*7
-H*8 37.4
40.6 (3.2)
[image:36.612.66.541.359.716.2]40.8 (3.3)
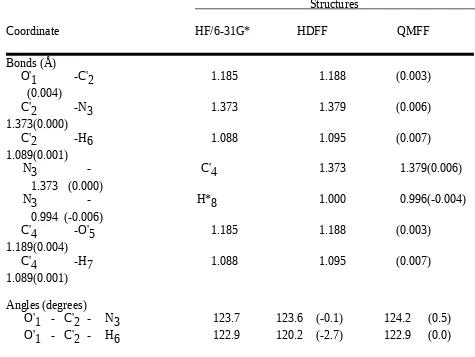
Table II.H. Comparison of N-Formylformamide Structures Optimized by Ab Initio Calculation (HF/6-31G*) and by the HDFF and QMFF Force Fields.
Structures
Coordinate HF/6-31G* HDFF QMFF
Bonds (Å)
O'1 -C'2 1.185 1.188 (0.003)
(0.004)
C'2 -N3 1.373 1.379 (0.006)
1.373(0.000)
C'2 -H6 1.088 1.095 (0.007)
1.089(0.001)
N3 - C'4 1.373 1.379(0.006)
1.373 (0.000)
N3 - H*8 1.000 0.996(-0.004)
0.994 (-0.006)
C'4 -O'5 1.185 1.188 (0.003)
1.189(0.004)
C'4 -H7 1.088 1.095 (0.007)
1.089(0.001) Angles (degrees)
N3 -C'2 - H6 113.3 116.2 (2.9) 112.9 (-0.4)
C'2 - N3 - C'4 125.0 125.7 (0.7) 125.4
(0.4)
C'2 - N3 - H*8 117.5 117.2 (-0.3) 117.3
(-0.2)
C'4 - N3 - H*8 117.5 117.2 (-0.3) 117.3
(-0.2)
N3 -C'4 - O'5 123.7 123.6 (-0.1) 124.2 (0.5) N3 -C'4 - H7 113.3 116.2 (2.9) 112.9 (-0.4) O'5 - C'4 - H7 122.9 120.2 (-2.7) 122.9 (0.0) Torsions (degrees)
O'1 - C'2 - N3 - C'4 180.0 180.0 (0.0) 180.0 (0.0) O'1 - C'2 - N3 - H*8 0.0 0.0 (0.0) 0.0 (0.0) H6 -C'2 - N3 - C'4 0.0 0.0 (0.0) 0.0 (0.0) H6 -C'2 - N3 - H*8 180.0 180.0 (0.0) 180.0 (0.0)
C'2 - N3 - C'4 - O'5 180.0 180.0 (0.0)
180.0 (0.0)
C'2 - N3 - C'4 - H7 0.0 0.0 (0.0)
0.0 (0.0)
H*8 - N3 - C'4 - O'5 0.0 0.0 (0.0)
0.0 (0.0)
H*8 - N3 - C'4 - H7 180.0 180.0 (0.0)
180.0 (0.0) Out-of-Planes (degrees)
O'1 - C'2 - N3 - H6 0.0 0.0 (0.0) 0.0 (0.0)
C'2 - N3 - H*8 - C'4 0.0 0.0 (0.0)
0.0 (0.0)
[image:37.612.72.535.65.557.2]O'5 - C'4 - N3 - H7 0.0 0.0 (0.0) 0.0 (0.0)
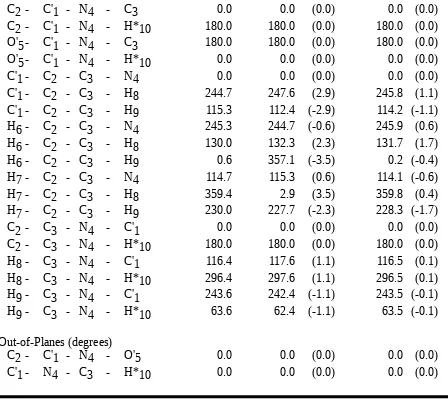
Table II.I. Comparison of Butyrolactam Structures Optimized by Ab Initio Calculation (HF/6-31G*) and by the HDFF and QMFF Force Fields.
Structures
Bonds (Å)
C'1 - C2 1.533 1.527 (-0.006) 1.527 C'1 - N4 1.358 1.384 (0.027) 1.354 C'1 - O'5 1.186 1.189 (0.002) 1.186 C2 -C3 1.549 1.533 (-0.016) 1.542 C2 -H6 1.082 1.085 (0.003) 1.080 C2 -H7 1.082 1.085 (0.003) 1.080 C3 -N4 1.455 1.429 (-0.026) 1.450 C3 -H8 1.083 1.086 (0.003) 1.082 C3 -H9 1.083 1.086 (0.003) 1.082 N4 - H*10 0.996 0.999 (0.003) 0.994 Angles (degrees)
C2 - C'1 - N4 91.2 93.9 (2.8) 91.2 (0.0) C2 - C'1 - O'5 135.9 134.3 (-1.6) 136.2 (0.3) N4 - C'1 - O'5 133.0 131.8 (-1.2) 132.7 (-0.3) C'1 - C2 - C3 85.7 82.5 (-3.1) 85.7 (0.0) C'1 - C2 - H6 114.1 116.0 (1.9) 113.5 (-0.6) C'1 - C2 - H7 114.1 116.0 (1.9) 113.5 (-0.6) C3 - C2 - H6 115.7 115.0 (-0.7) 116.2 (0.4) C3 - C2 - H7 115.7 115.0 (-0.7) 116.2 (0.4) H6 - C2 - H7 109.9 110.1 (0.2) 110.1 (0.2) C2 - C3 - N4 86.9 91.9 (4.9) 87.1 (0.1) C2 -C3 - H8 115.3 115.2 (-0.1) 115.5 (0.2) C2 - C3 - H9 115.3 115.2 (-0.1) 115.5 (0.2) N4 - C3 - H8 114.1 109.3 (-4.8) 113.1 (-1.0) N4 - C3 - H9 114.1 109.3 (-4.8) 113.1 (-1.0) H8 - C3 - H9 109.7 113.6 (3.9) 110.8 (1.1) C'1 - N4 - C3 96.2 91.7 (-4.6) 96.1 (-0.1) C'1 - N4 - H*10 131.0 137.5 (6.5) 130.5 (-0.4) C3 - N4 - H*10 132.8 130.8 (-2.0) 133.4 (0.6) Torsions (degrees)
C2 - C'1 - N4 - C3 0.0 0.0 (0.0) 0.0 (0.0) C2 - C'1 - N4 - H*10 180.0 180.0 (0.0) 180.0 (0.0) O'5- C'1 - N4 - C3 180.0 180.0 (0.0) 180.0 (0.0) O'5- C'1 - N4 - H*10 0.0 0.0 (0.0) 0.0 (0.0) C'1 - C2 - C3 - N4 0.0 0.0 (0.0) 0.0 (0.0) C'1 - C2 - C3 - H8 244.7 247.6 (2.9) 245.8 (1.1) C'1 - C2 - C3 - H9 115.3 112.4 (-2.9) 114.2 (-1.1) H6 - C2 - C3 - N4 245.3 244.7 (-0.6) 245.9 (0.6) H6 - C2 - C3 - H8 130.0 132.3 (2.3) 131.7 (1.7) H6 - C2 - C3 - H9 0.6 357.1 (-3.5) 0.2 (-0.4) H7 - C2 - C3 - N4 114.7 115.3 (0.6) 114.1 (-0.6) H7 - C2 - C3 - H8 359.4 2.9 (3.5) 359.8 (0.4) H7 - C2 - C3 - H9 230.0 227.7 (-2.3) 228.3 (-1.7) C2 - C3 - N4 - C'1 0.0 0.0 (0.0) 0.0 (0.0) C2 - C3 - N4 - H*10 180.0 180.0 (0.0) 180.0 (0.0) H8 - C3 - N4 - C'1 116.4 117.6 (1.1) 116.5 (0.1) H8 - C3 - N4 - H*10 296.4 297.6 (1.1) 296.5 (0.1) H9 - C3 - N4 - C'1 243.6 242.4 (-1.1) 243.5 (-0.1) H9 - C3 - N4 - H*10 63.6 62.4 (-1.1) 63.5 (-0.1) Out-of-Planes (degrees)
[image:39.612.74.522.70.472.2]C2 - C'1 - N4 - O'5 0.0 0.0 (0.0) 0.0 (0.0) C'1 - N4 - C3 - H*10 0.0 0.0 (0.0) 0.0 (0.0)
Table III.A. Comparison of Formamide Frequencies Obtained by Ab Initio Calculation (HF/6-31G*) and the HDFF and QMFF Force Fields.
Frequencies
HF/6-31G* HDFF QMFF
Mode
1788.7 2055.6 (266.9) 1806.0 (17.2) NH2 bend 1563.0 1498.2 (-64.8) 1529.0 (-34.0) C'H rock 1378.6 1168.6 (-210.0) 1333.7 (-44.8) C'N stretch
1182.5 1276.8 (94.3) 1230.5 (48.0) C'O' oop deformation 1159.9 933.4 (-226.5) 1116.8 (-43.1) NH2 rock
[image:40.612.83.538.72.188.2] [image:40.612.73.535.282.712.2]673.4 679.5 (6.1) 697.2 (23.8) C'N torsion 617.6 578.3 (-39.3) 627.3 (9.7) C'O' rock 109.1 353.6 (244.5) 402.5 (293.3) amino inversion
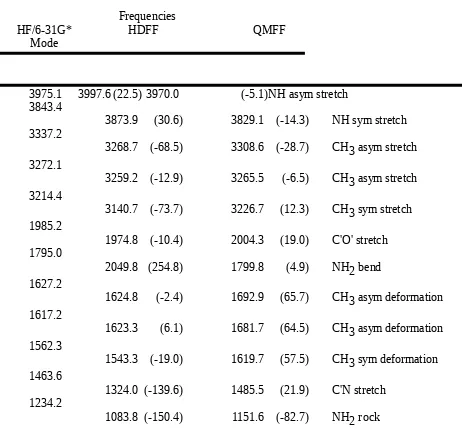
Table III.B. Comparison of Planar Acetamide Frequencies Obtained by Ab Initio Calculation (HF/6-31G*) and the HDFF and QMFF Force Fields.
Frequencies
HF/6-31G* HDFF QMFF
Mode
3975.1 3997.6 (22.5) 3970.0 (-5.1)NH asym stretch 3843.4
3873.9 (30.6) 3829.1 (-14.3) NH sym stretch 3337.2
3268.7 (-68.5) 3308.6 (-28.7) CH3 asym stretch 3272.1
3259.2 (-12.9) 3265.5 (-6.5) CH3 asym stretch 3214.4
3140.7 (-73.7) 3226.7 (12.3) CH3 sym stretch 1985.2
1974.8 (-10.4) 2004.3 (19.0) C'O' stretch 1795.0
2049.8 (254.8) 1799.8 (4.9) NH2 bend 1627.2
1624.8 (-2.4) 1692.9 (65.7) CH3 asym deformation 1617.2
1623.3 (6.1) 1681.7 (64.5) CH3 asym deformation 1562.3
1543.3 (-19.0) 1619.7 (57.5) CH3 sym deformation 1463.6
1324.0 (-139.6) 1485.5 (21.9) C'N stretch 1234.2
1168.9
1042.5 (-126.4) 1143.8 (-25.1) CH3 rock 1076.6
1001.5 (-75.1) 1060.5 (-16.1) CH3 rock 913.0
733.5 (-179.5) 886.4 (-26.6) C'C stretch 696.4
783.9 (87.5) 715.0 (18.5) C'O' oop deformation 598.1
558.2 (-39.9) 605.7 (7.6) C'O' rock 562.6
641.6 (79.0) 614.1 (51.5) C'N torsion 446.6
356.5 (-90.1) 403.7 (-42.9) C'C rock 11.7
-6.5a,b (-18.2) 118.0 (106.3) CH3 torsion -144.5a
175.9 (320.4) 380.8 (525.3) amino inversion
[image:41.612.80.522.507.723.2]a The minus sign indicates that the frequency was calculated from the absolute value of a negative eigenvalue. bThis imaginary frequency arises from selecting the force field stationary state with the geometry closest to the corresponding ab initio structure for comparison. In this case the configuration computed with the force field corresponds to a transition state.
Table III.C. Comparison of Planar Trans N-Methylformamide Frequencies Obtained by Ab Initio Calculation (HF/6-31G*) and the HDFF and QMFF Force Fields.
Frequencies
HF/6-31G* HDFF QMFF
Mode
3900.8 3934.0 (33.2) 3867.0 (-38.3) NH stretch 3312.3 3259.5 (-52.8) 3268.3 (-44.0) CH3 asym stretch 3296.9 3262.8 (-34.1) 3255.2 (-41.7) CH3 asym stretch 3230.5 3142.0 (-88.6) 3207.0 (-23.5) CH3 sym stretch 3206.7 3149.2 (-57.5) 3235.7 (28.9) C'H stretch 1977.2 1994.6 (17.4) 1978.5 (1.3) C'O' Stretch 1694.9 1792.2 (97.3) 1699.2 (4.2) NH rock
1609.3 1586.9 (-22.4) 1601.2 (-8.2) CH3 sym deformation 1559.9 1487.6 (-72.3) 1529.4 (-30.5) C'H rock
1348.6 1113.1 (-235.4) 1304.3 (-44.2) C'N stretch 1283.7 1039.3 (-244.4) 1239.1 (-44.6) CH3 rock 1266.0 1034.3 (-231.7) 1269.5 (3.5) CH3 rock
1170.7 1283.5 (112.8) 1242.8 (72.1) C'O' oop deformation 1035.4 823.9 (-211.5) 1000.1 (-35.3) NC stretch
837.8 692.0 (-145.9) 812.8 (-25.0) C'O' rock 486.9 560.8 (73.9) 574.4 (87.5) C'N torsion 291.8 301.2 (9.4) 317.4 (25.6) NC rock
228.5 323.4 (94.9) 328.1 (99.6) amino inversion -60.6a -62.5a (-1.9) -136.5a (-75.9) CH3 torsion
a The minus sign indicates that the frequency was calculated from the absolute value of a negative eigenvalue.
Table III.D. Comparison of Cis N-Methylformamide Frequencies Obtained by Ab Initio Calculation (HF/6-31G*) and the HDFF and QMFF Force Fields.
Frequencies
HF/6-31G* HDFF QMFF
Mode
3855.6 3941.0 (85.4) 3810.5 (-45.1) NH stretch 3299.0 3259.9 (-39.0) 3278.2 (-20.8) CH3 asym stretch 3268.8 3261.4 (-7.4) 3252.0 (-16.8) CH3 asym stretch 3215.4 3141.7 (-73.6) 3206.4 (-8.9) CH3 sym stretch 3201.4 3149.8 (-51.6) 3205.7 (4.3) C'H stretch 1988.8 2012.1 (23.3) 1988.0 (-0.8) C'O' stretch
1681.4 1616.1 (-65.3) 1690.8 (9.4) CH3 asym deformation 1643.2 1748.2 (105.0) 1626.3 (-16.9) NH rock
1631.5 1602.2 (-29.3) 1641.2 (9.7) CH3 asym deformation 1623.1 1595.2 (-27.9) 1638.1 (15.0) CH3 sym deformation 1542.4 1515.9 (-26.5) 1562.4 (20.1) C'H rock
1412.8 1049.9 (-362.9) 1386.1 (-26.7) C'N stretch 1270.2 1105.5 (-164.7) 1241.8 (-28.3) CH3 rock 1258.9 1024.2 (-234.7) 1241.0 (-18.0) CH3 rock
1179.5 1254.5 (75.1) 1180.0 (0.5) C'O' oop deformation 1096.1 953.9 (-142.1) 1068.9 (-27.1) NC stretch
[image:42.612.77.530.396.720.2]193.0 179.5 (-13.5) 215.3 (22.3) CH3 torsion 66.3 -50.8a (-117.1) -74.9a (-141.2) amino inversion
a The minus sign indicates that the frequency was calculated from the absolute value of a negative eigenvalue.
Table III.E. Comparison of Trans N-Methylacetamide Frequencies Obtained by Ab Initio Calculation (HF/6-31G*) and the HDFF and QMFF Force Fields.
Frequencies
HF/6-31G* HDFF QMFF
Mode
3914.8 3934.0 (19.2) 3886.2 (-28.6) NH stretch
3307.6 3259.5 (-48.1) 3269.1 (-38.4) N-CH3 asym stretch 3301.9 3261.6 (-40.3) 3267.1 (-34.9) C'-CH3 asym stretch 3296.4 3265.0 (-31.4) 3280.5 (-15.9) C'-CH3 asym stretch 3295.9 3262.9 (-33.0) 3256.2 (-39.7) N-CH3 asym stretch 3229.0 3142.0 (-87.0) 3207.7 (-21.3) N-CH3 sym stretch 3223.7 3140.9 (-82.8) 3222.0 (-1.6) C'-CH3 sym stretch 1956.5 1974.9 (18.4) 1973.3 (16.7) C'O' stretch
1711.0 1792.0 (81.0) 1707.0 (-4.0) NH rock
1658.9 1617.7 (-41.2) 1675.2 (16.2) N-CH3 asym deformation 1630.8 1627.0 (-3.8) 1650.4 (19.6) C'-CH3 asym deformation 1628.8 1605.9 (-22.9) 1639.8 (11.0) N-CH3 asym deformation 1617.3 1626.3 (9.0) 1693.2 (75.9) C'-CH3 asym deformation 1609.1 1586.3 (-22.8) 1603.4 (-5.7) N-CH3 sym deformation 1555.2 1542.0 (-13.2) 1594.6 (39.4) C'-CH3 sym deformation 1408.2 1289.9 (-118.3) 1416.8 (8.6) C'-N stretch
1311.3 1044.3 (-267.0) 1249.8 (-61.4) N-CH3 rock 1265.7 1030.5 (-235.2) 1268.7 (3.0) N-CH3 rock 1191.5 1007.8 (-183.7) 1116.5 (-75.0) N-C stretch 1172.2 1058.3 (-113.9) 1151.6 (-20.6) C'-CH3 rock 1094.9 1034.8 (-60.1) 1033.7 (-61.2) C'-CH3 rock 950.6 731.2 (-219.4) 932.8 (-17.8) C'-C stretch
686.0 793.6 (107.6) 714.4 (28.4) C'O' oop deformation 673.6 605.8 (-67.8) 653.6 (-20.0) C'O' rock
[image:43.612.77.530.277.706.2]365.9 373.4 (7.5) 506.5 (140.6) C'N torsion 284.0 266.6 (-17.4) 290.8 (6.8) NC rock
168.1 206.6 (38.5) 190.2 (22.1) amino inversion 19.8 -55.5a,b (-75.3) -110.4a,b(-130.2) C'-CH3 torsion -41.4a -78.5a (-37.1) -127.0a (-85.6) N-CH3 torsion
[image:44.612.78.530.409.706.2]a The minus sign indicates that the frequency was calculated from the absolute value of a negative eigenvalue. bThis imaginary frequency arises from selecting the force field stationary state with the geometry closest to the corresponding ab initio structure for comparison. In this case the configuration computed with the force field corresponds to a transition state.
Table III.F. Comparison of Cis Planar N-Methylacetamide Frequencies Obtained byAb Initio Calculation (HF/6-31G*) and the HDFF and QMFF Force Fields.
Frequencies
HF/6-31G* HDFF QMFF
Mode
3872.7 3940.6 (67.9) 3831.3 (-41.4) NH stretch
3337.8 3268.6 (-69.2) 3305.8 (-32.0) C'-CH3 asym stretch 3306.7 3258.8 (-47.9) 3263.7 (-43.1) N-CH3 asym stretch 3276.5 3258.6 (-17.9) 3267.2 (-9.3) C'-CH3 asym stretch 3260.8 3262.6 (1.8) 3259.1 (-1.8) N-CH3 asym stretch 3218.9 3140.4 (-78.5) 3227.3 (8.4) C'-CH3 sym stretch 3208.6 3141.8 (-66.8) 3208.0 (-0.6) N-CH3 sym stretch 1969.9 1985.1 (15.2) 1999.5 (29.6) C'O' stretch
1670.8 1620.2 (-50.6) 1694.5 (23.7) N-CH3 asym deformation 1644.2 1608.4 (-35.8) 1643.0 (-1.2) N-CH3 asym deformation 1633.9 1583.5 (-50.4) 1634.7 (0.8) N-CH3 sym deformation 1626.7 1742.4 (115.7) 1589.0 (-37.7) NH rock
1625.2 1627.7 (2.5) 1701.9 (76.7) C'-CH3 asym deformation 1611.2 1628.6 (17.4) 1686.0 (74.8) C'-CH3 asym deformation 1562.4 1543.6 (-18.8) 1617.5 (55.1) C'-CH3 deformation 1469.5 1314.4 (-155.1) 1492.6 (23.1) C'-N stretch
1193.1 1029.5 (-163.6) 1131.4 (-61.7) NC stretch 1166.8 1042.9 (-123.9) 1137.5 (-29.3) C'-CH3 rock 1093.9 1003.4 (-90.5) 1077.4 (-16.4) C'-CH3 rock 867.3 708.5 (-158.8) 838.7 (-28.6) C'C stretch
671.0 771.7 (100.7) 688.5 (17.4) C'O' oop deformation 611.9 457.7 (-154.2) 598.0 (-14.0) C'O' rock
529.0 507.0 (-22.0) 495.1 (-33.9) C'C rock 508.3 594.3 (86.0) 477.9 (-30.4) C'N torsion 297.8 268.3 (-29.5) 295.0 (-2.7) NC rock 183.4 142.1 (-41.3) 195.1 (11.7) N-CH3 torsion 115.2 98.5 (-16.7) 166.1 (50.9) C'-CH3 torsion -58.6a -66.9a (-8.3) -135.5a (-77.0) amino inversion
a The minus sign indicates that the frequency was calculated from the absolute value of a negative eigenvalue.
Table III.G. Comparison of N,N-Dimethylformamide Frequencies Obtained by Ab Initio Calculation (HF/6-31G*) and the HDFF and QMFF Force Fields.
Frequencies
HF/6-31G* HDFF QMFF
Mode
3361.4 3261.1 (-100.3) 3293.8 (-67.7) CH3 asym stretch 3300.4 3264.4 (-36.0) 3290.6 (-9.8) CH3 asym stretch 3249.6 3259.4 (9.8) 3233.1 (-16.5) CH3 asym stretch 3248.6 3259.0 (10.4) 3234.2 (-14.4) CH3 asym stretch 3214.5 3148.5 (-66.0) 3208.6 (-5.8) C'H stretch 3201.1 3142.7 (-58.4) 3194.5 (-6.6) CH3 sym stretch 3196.2 3143.5 (-52.7) 3193.1 (-3.1) CH3 sym stretch 1961.1 2001.1 (40.0) 1974.2 (13.1) C'O' stretch
[image:45.612.76.529.452.717.2]1557.1 1525.0 (-32.1) 1553.1 (-4.0) C'H rock
1404.6 1231.3 (-173.3) 1406.0 (1.4) NC asym stretch 1287.8 1042.6 (-245.2) 1273.9 (-13.9) CH3 rock 1239.0 1001.6 (-237.4) 1225.2 (-13.8) CH3 rock 1199.3 988.7 (-210.6) 1160.2 (-39.2) CH3 rock 1185.5 1028.8 (-156.7) 1180.7 (-4.8) CH3 rock
1165.1 1262.6 (97.5) 1187.1 (22.0) C'O' oop deformation 941.5 821.0 (-120.5) 909.7 (-31.8) NC sym stretch 711.9 586.9 (-125.0) 695.7 (- 16.2) C'O' rock 426.3 419.6 (-6.7) 402.1 (-24.2) CNC bend 351.2 307.3 (-43.9) 356.3 (5.1) CNC rock 346.1 394.9 (48.8) 352.0 (6.0) C'N torsion 242.7 258.8 (16.1) 243.1 (0.4) amino inversion 161.7 143.3 (-18.4) 83.4 (-78.2) CH3 sym torsion 83.9 87.8 (3.9) -127.2a,b (-211.1) CH3 asym torsion
[image:46.612.79.537.68.302.2]a The minus sign indicates that the frequency was calculated from the absolute value of a negative eigenvalue. bThis imaginary frequency arises from selecting the force field stationary state with the geometry closest to the corresponding ab initio structure for comparison. In this case the configuration computed with the force field corresponds to a transition state.
Table III.H. Comparison of Planar N,N-Dimethylacetamide Frequencies Obtained by Ab Initio Calculation (HF/6-31G*) and the HDFF and QMFF Force Fields.
Frequencies
HF/6-31G* HDFF QMFF
Mode
1935.2 1968.4 (33.3) 1974.6 (39.4) C'O' stretch
1690.9 1592.8 (-98.1) 1675.7 (-15.3) N-CH3 asym deformation 1657.5 1605.0 (-52.5) 1679.5 (22.0) N-CH3 asym deformation 1655.2 1619.7 (-35.5) 1647.8 (-7.4) N-CH3 asym deformation 1645.4 1633.5 (-11.9) 1640.0 (-5.5) N-CH3 asym deformation 1637.8 1669.2 (31.4) 1710.1 (72.3) N-CH3 sym deformation 1627.7 1610.9 (-16.8) 1703.5 (75.9) C'-CH3 asym deformation 1616.3 1633.6 (17.3) 1703.3 (87.0) C'-CH3 asym deformation 1593.0 1371.7 (-221.3) 1621.9 (28.9) N-CH3 sym deformation 1581.6 1675.5 (93.9) 1690.5 (108.9) C'-N stretch
1538.5 1549.1 (10.6) 1569.1 (30.6) C'-CH3 sym deformation 1408.1 1242.1 (-166.0) 1423.2 (15.1) NC asym stretch
1313.6 1156.7 (-156.9) 1302.0 (-11.6) N-CH3 rock 1286.7 1034.1 (-252.6) 1269.5 (-17.1) N-CH3 rock 1237.5 997.6 (-239.8) 1219.5 (-18.0) N-CH3 rock 1182.2 1040.0 (-142.2) 1184.7 (2.5) N-CH3 rock 1164.0 1053.9 (-110.1) 1129.3 (-34.7) C'-CH3 rock 1126.8 1028.2 (-98.6) 1109.2 (-17.7) C'-CH3 rock 1053.6 966.5 (-87.1) 1021.5 (-32.2) NC sym stretch 789.1 678.5 (-110.7) 767.4 (-21.7) C'C stretch
651.2 782.8 (131.6) 679.2 (28.0) C'O' oop deformation 639.1 564.8 (-74.2) 635.7 (-3.4) C'O' rock
500.7 390.0 (-110.8) 470.9 (-29.8) C'C rock 435.6 439.9 (4.3) 435.7 (0.2) CNC bend 348.3 321.6 (-26.7) 361.0 (12.8) CNC rock 274.7 322.3 (47.6) 284.7 (10.0) amino inversion 212.5 208.1 (-4.4) 230.0 (17.5) C'-CH3 torsion 143.1 145.8 (2.7) 128.7 (-14.5) C'N torsion 78.3 85.0 (6.7) -84.1a,b (-162.5) N-CH3 torsion -62.8a 10.9 (73.7) -171.2a (-108.4) N-CH3 torsion
[image:47.612.90.535.75.547.2]a The minus sign indicates that the frequency was calculated from the absolute value of a negative eigenvalue. bThis imaginary frequency arises from selecting the force field stationary state with the geometry closest to the corresponding ab initio structure for comparison. In this case the configuration computed with the force field corresponds to a transition state.
Frequencies
HF/6-31G* HDFF QMFF
Mode
3926.2 3989.0 (62.8) 3927.2 (1.0) NH asym stretch 3925.9 3996.5 (70.6) 3932.8 (6.9) NH asym stretch 3814.9 3884.8 (69.9) 3813.5 (-1.4) NH sym stretch 3810.8 3870.7 (59.9) 3801.9 (-8.9) NH sym stretch 2000.3 2037.0 (36.7) 1996.5 (-3.7) C'O' stretch 1811.9 1929.0 (117.1) 1726.5 (-85.4) NH2 asym bend 1805.5 1953.0 (147.5) 1755.0 (-50.5) NH2 sym bend 1550.2 1404.7 (-145.5) 1552.1 (1.8) C'N asym stretch 1303.6 1118.7 (-184.9) 1168.4 (-135.2) NH2 sym rock 1166.1 990.0 (-176.1) 1064.0 (-102.1) NH2 asym rock 1034.8 640.8 (-394.0) 947.2 (-87.6) C'N sym stretch 882.1 919.3 (37.2) 970.8 (88.7) C'O' oop deformation 648.8 770.6 (121.8) 720.7 (71.9) asym amino inversion 629.9 541.3 (-88.6) 666.3 (36.4) C'O' rock
598.8 675.5 (76.7) 607.6 (8.8) sym amino inversion 514.2 498.3 (-15.9) 567.5 (53.3) NC'N bend
[image:48.612.73.537.148.409.2]483.2 415.1 (-68.1) 435.5 (-47.6) asym C'N torsion 417.2 416.5 (-0.7) 452.3 (35.1) sym C'N torsion
Table III.J. Comparison of N-Formylformamide Frequencies Obtained by Ab Initio Calculation (HF/6-31G*) and the HDFF and QMFF Force Fields.
Frequencies
HF/6-31G* HDFF QMFF
Mode
3827.6 3944.4 (116.8) 3896.6 (69.0) NH stretch 3256.1 3152.0 (-104.1) 3228.4 (-27.7) C'H sym stretch 3232.7 3148.7 (-84.0) 3224.4 (-8.2) C'H asym stretch 2046.2 1993.8 (-52.4) 2010.4 (-35.8) C'O' sym stretch 2007.6 2017.5 (9.9) 1996.8 (-10.8) C'O asym stretch 1627.9 1534.7 (-93.2) 1570.2 (-57.7) NH rock
1584.9 1518.6 (-66.3) 1575.8 (-9.1) C'H sym rock 1488.2 1493.6 (5.4) 1483.4 (-4.9) C'H asym rock 1370.2 1133.2 (-237.0) 1311.5 (-58.7) C'N sym stretch 1294.4 992.4 (-302.0) 1305.6 (11.2) C'N asym stretch
777.1 856.5 (79.4) 803.6 (26.5) C'N asym torsion 687.2 680.5 (-6.7) 721.7 (34.5) C'O' asym rock 580.1 430.3 (-149.8) 570.7 (-9.4) C'O' sym rock 303.1 306.7 (3.6) 299.2 (-3.9) C'N sym torsion 275.7 301.4 (25.7) 325.9 (50.2) C'NC' bend 137.3 145.6 (8.3) 142.7 (5.4) amino inversion
Table III.K. Comparison of Butyrolactam Frequencies Obtained by Ab Initio Calculation (HF/6-31G*) and the HDFF and QMFF Force Fields.
Frequencies
HF/6-31G* HDFF QMFF
Mode
3878.3 3937.5 (59.2) 3893.3 (15.0) NH stretch 3324.1 3267.3 (-56.8) 3318.1 (-5.9) CH asym stretch 3282.5 3276.7 (-5.8) 3295.6 (13.1) CH asym stretch 3268.7 3187.2 (-81.5) 3278.1 (9.4) CH sym stretch 3238.4 3175.4 (-62.9) 3247.6 (9.3) CH sym stretch 2057.2 2013.5 (-43.7) 2072.5 (15.3) C'O' stretch 1684.3 1705.0 (20.7) 1709.9 (25.6) CH2 scissor 1609.6 1647.6 (37.9) 1685.9 (76.2) CH2 scissor 1548.7 1203.2 (-345.5) 1449.2 (-99.5) NH rock 1457.9 1345.6 (-112.2) 1431.7 (-26.2) CH2 wag 1353.7 1136.7 (-217.0) 1186.9 (-166.8) CH2 wag 1310.9 1027.4 (-283.5) 1357.4 (46.5) CH2 twist 1304.7 955.5 (-349.1) 1291.6 (-13.0) C'N stretch 1264.2 1088.0 (-176.2) 1270.6 (6.4) CH2 twist 1170.5 1142.2 (-28.3) 1121.1 (-49.4) CH2 rock 1099.4 1783.6 (684.3) 1057.9 (-41.4) NC stretch 1056.5 862.4 (-194.1) 1095.9 (39.4) C'C stretch 965.5 774.6 (-190.9) 913.0 (-52.5) CC stretch
896.7 891.3 (-5.4) 906.5 (9.8) CH2 rock
817.1 519.6 (-297.4) 674.4 (-142.6) ring deformation 611.2 725.9 (114.7) 653.2 (42.0) C'O' oop deformation 518.5 554.4 (35.9) 530.4 (11.9) C'O' rock
407.6 -498.6a (-906.2) 556.5 (148.9) ring puckering 75.3 366.1 (290.7) 199.7 (124.4) amino inversion